Harvey Lodish Arnold Berk Paul Matsudaira Chris A

Harvey Lodish • Arnold Berk • Paul Matsudaira • Chris A. Kaiser • Monty Krieger • Matthew P. Scott • Lawrence Zipursky • James Darnell Molecular Cell Biology Fifth Edition Chapter 8: Cellular Energetics Copyright © 2004 by W. H. Freeman & Company

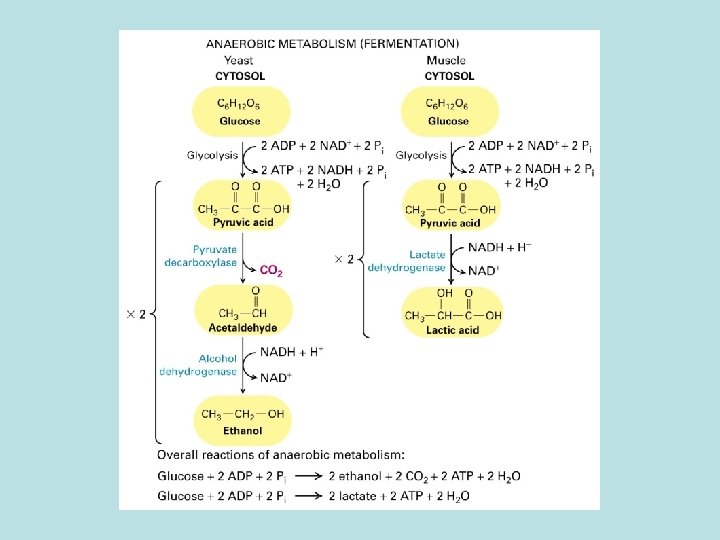
How cell generate ATP? ATP : 1. synthesis of protein and nucleic acid 2. transport molecules against concentration gradient 3. movement of cilia
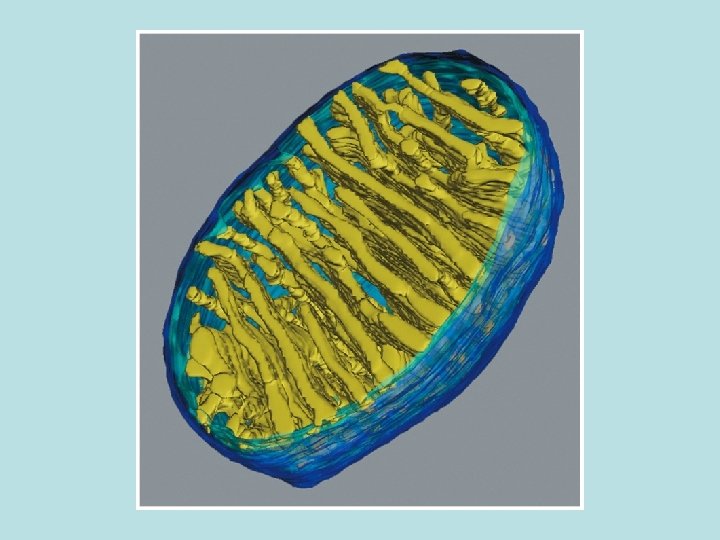

Chemiosmosis ATP generation for bacteria, mitochondria, and chloroplast ·Occur only in sealed membrane ·Stepwise movement of electrons for higher energy state to low energy state through electron carrier
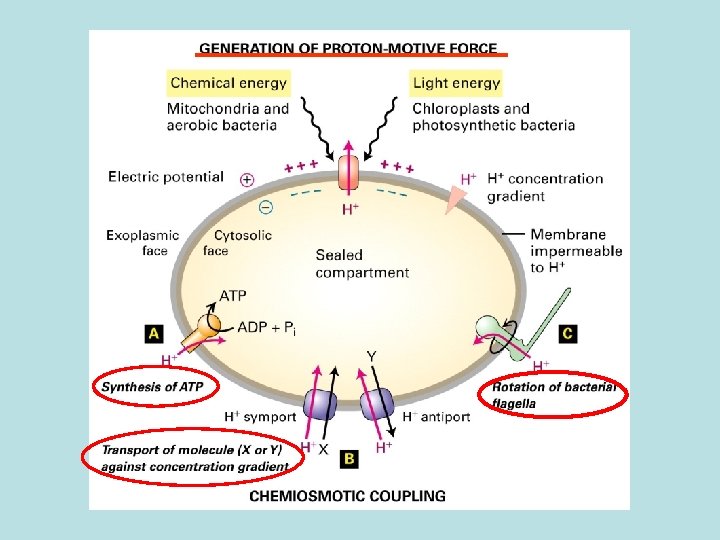

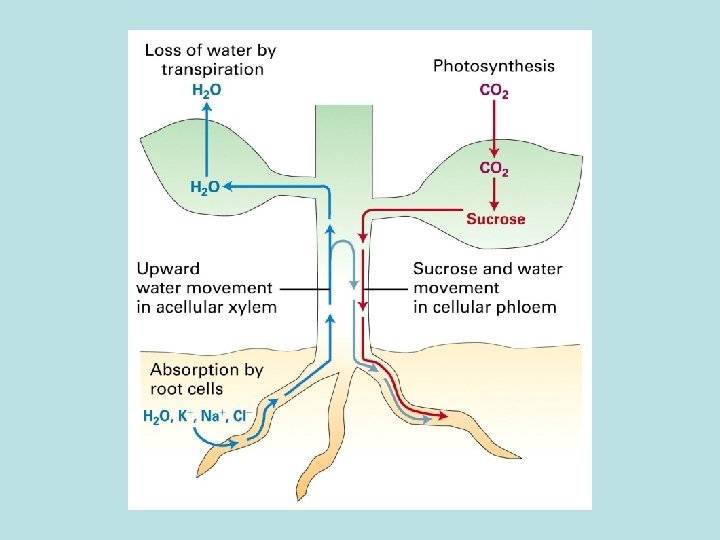
Proton motive force: Supply energy for transporting small molecule across membrane and against its concentration gradient
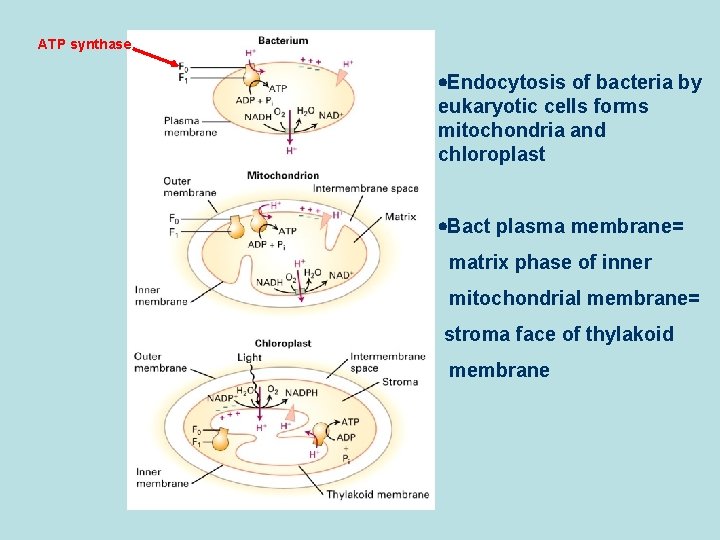
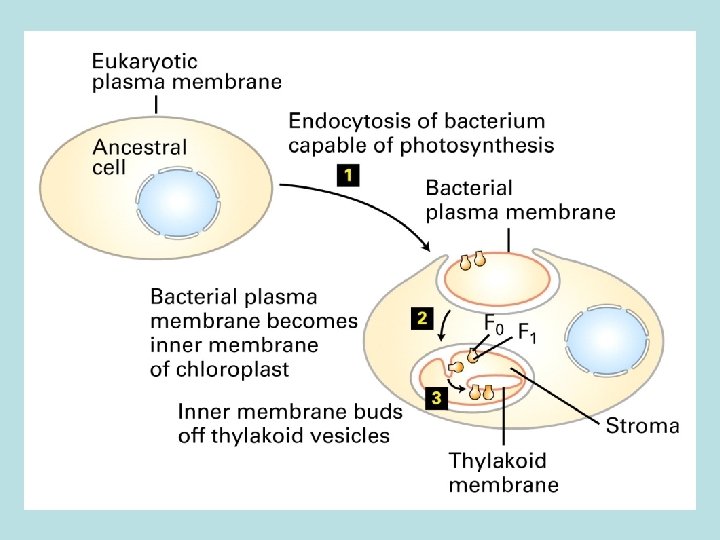
ATP synthase Endocytosis of bacteria by eukaryotic cells forms mitochondria and chloroplast Bact plasma membrane= matrix phase of inner mitochondrial membrane= stroma face of thylakoid membrane
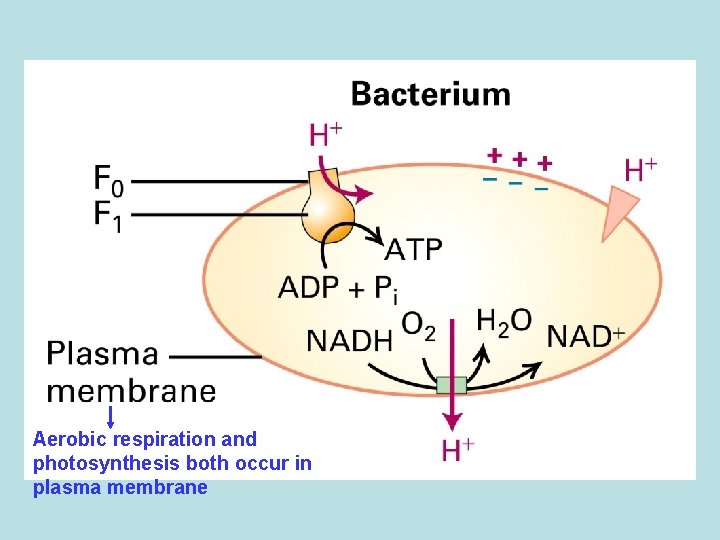
Aerobic respiration and photosynthesis both occur in plasma membrane
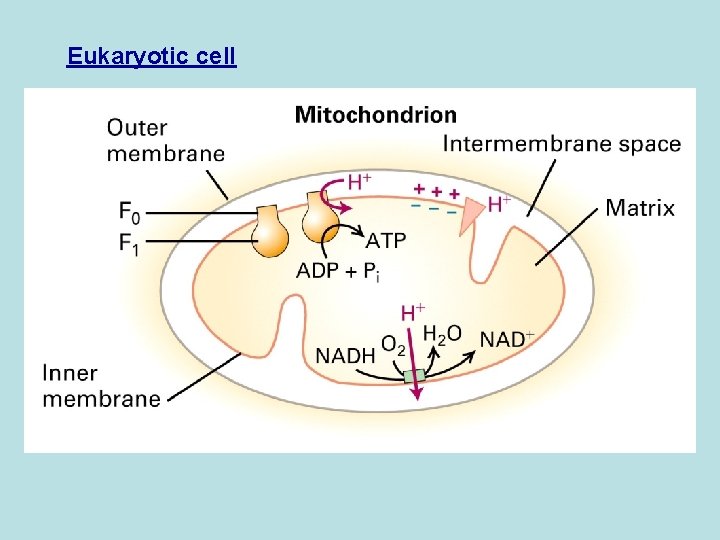
Eukaryotic cell
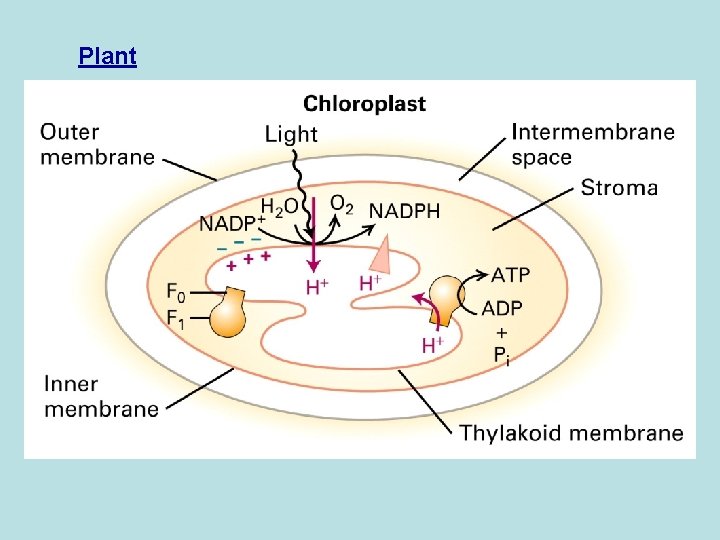
Plant
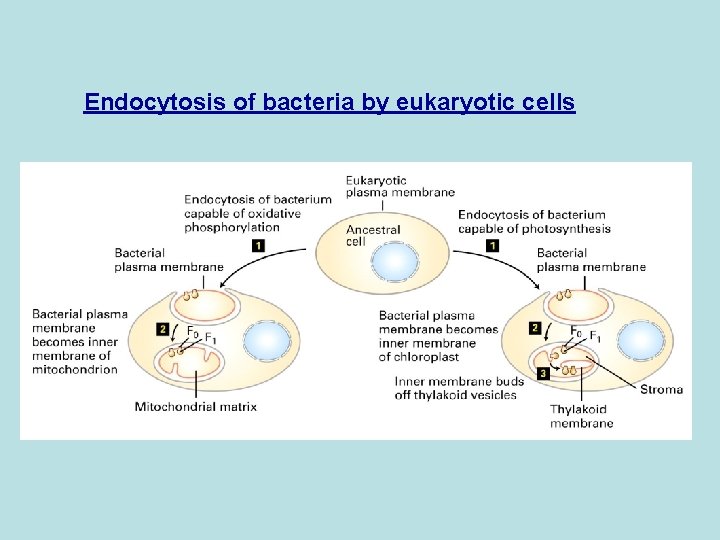
Endocytosis of bacteria by eukaryotic cells
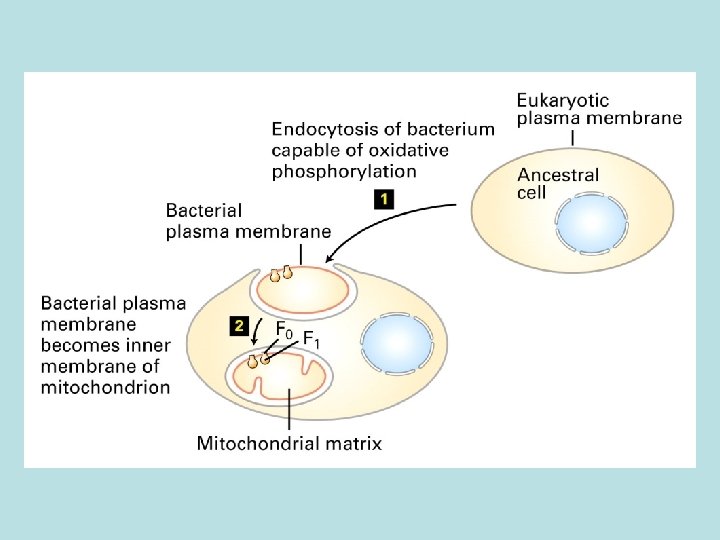
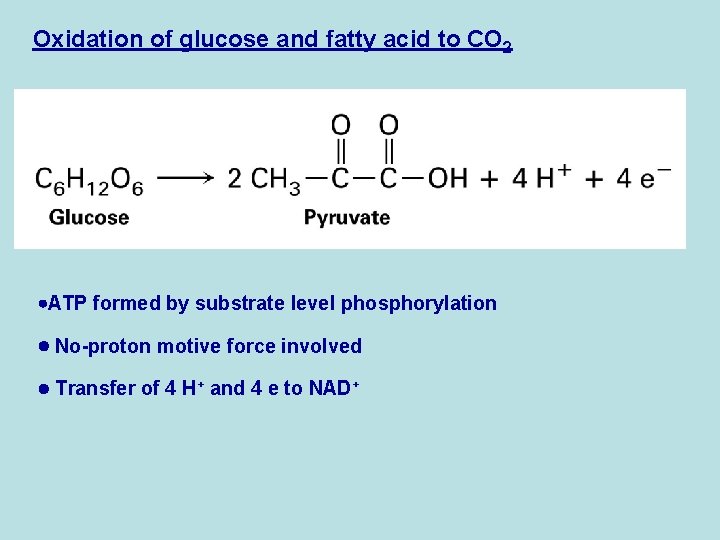
Oxidation of glucose and fatty acid to CO 2 ATP formed by substrate level phosphorylation No-proton motive force involved Transfer of 4 H+ and 4 e to NAD+
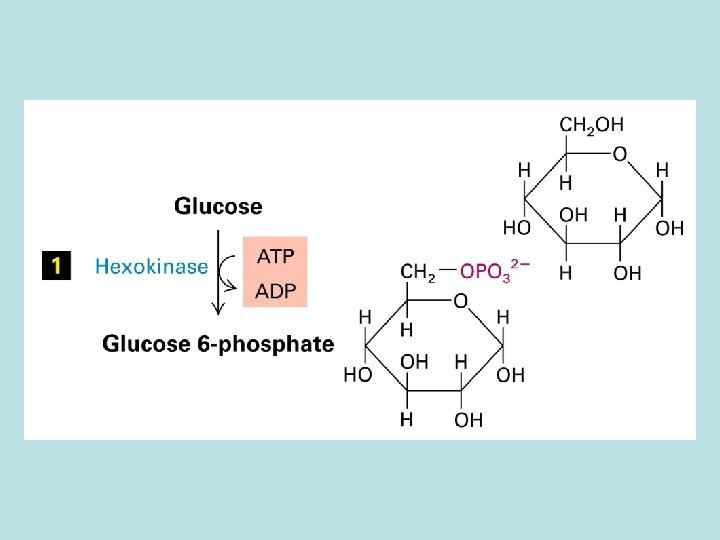
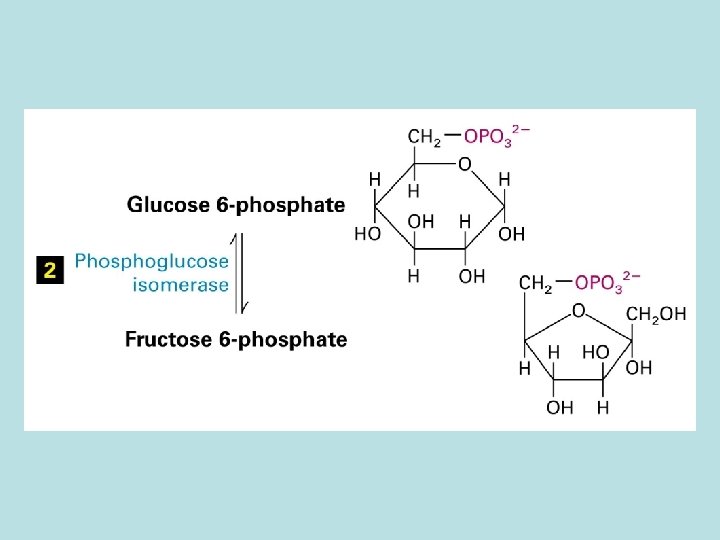
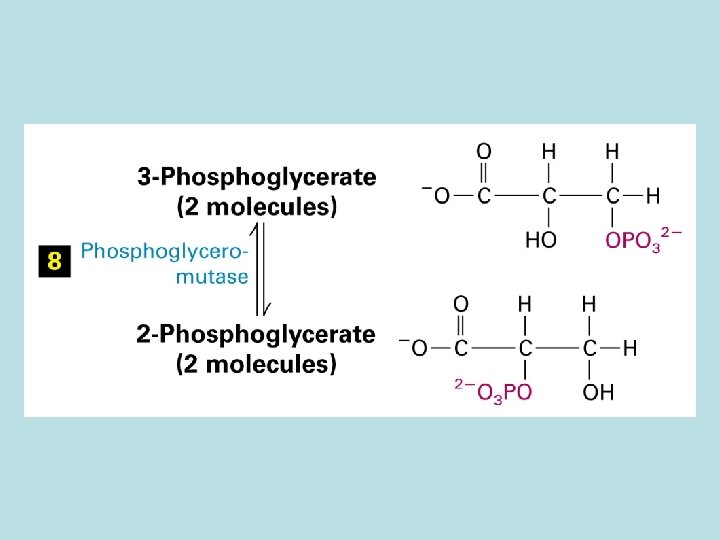
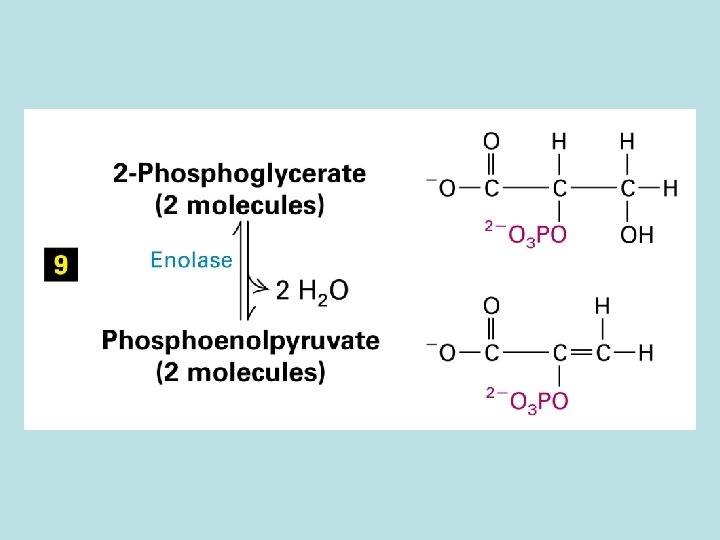
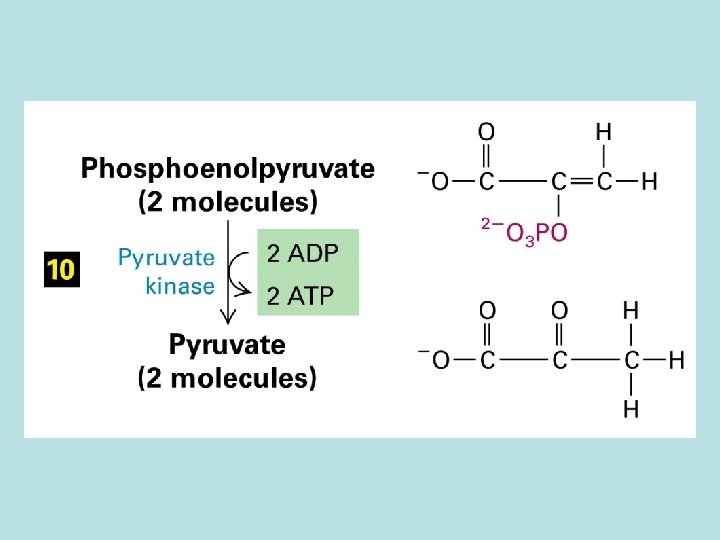
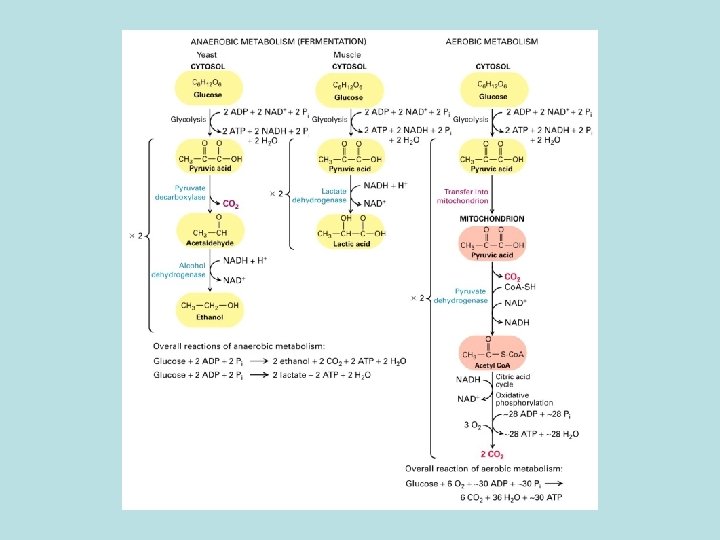
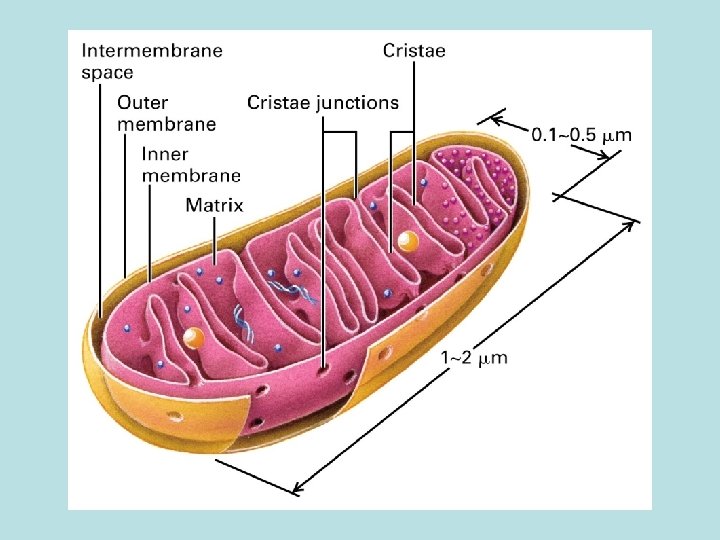
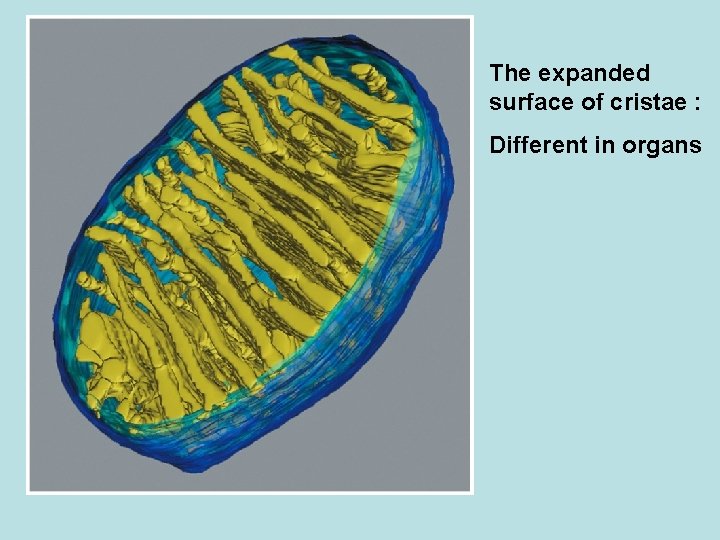
The expanded surface of cristae : Different in organs

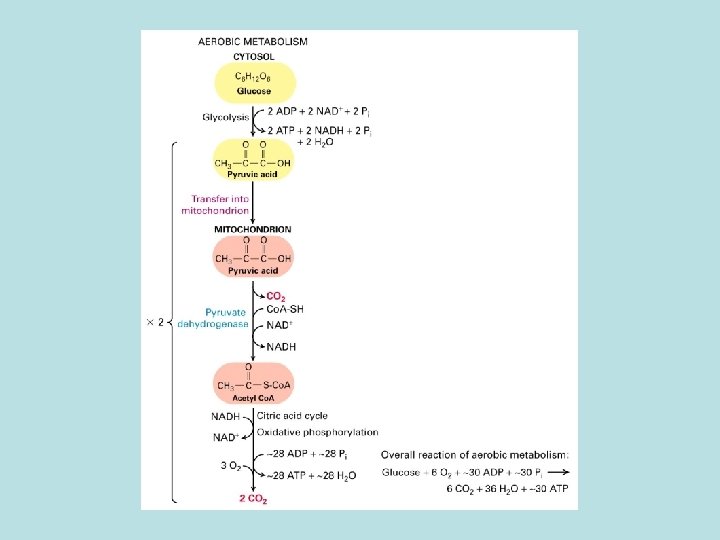
Aerobic oxidation of fatty acid and pyruvate in mitochondria Oxidation of pyruvate and fatty acid to CO 2 and coupled reduction of NAD+ to NADH and FADH 2 ·Electron transfer from NADH and FADH 2 to O 2 ·Harness energy stored in electrochemical gradient for ATP synthesis by F 0 F 1 complex
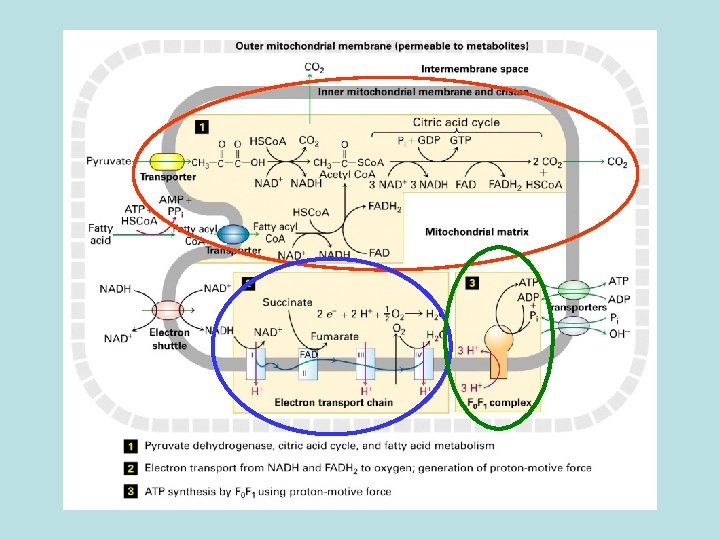
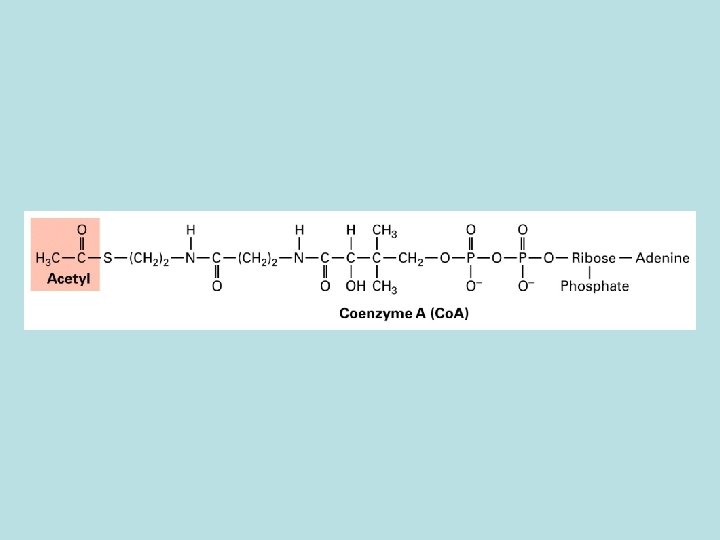

TCA cycle Oxidation of of acetyl co. A No O 2 involved in the oxidation energy stored in the reduced form of NADH and FADH 2
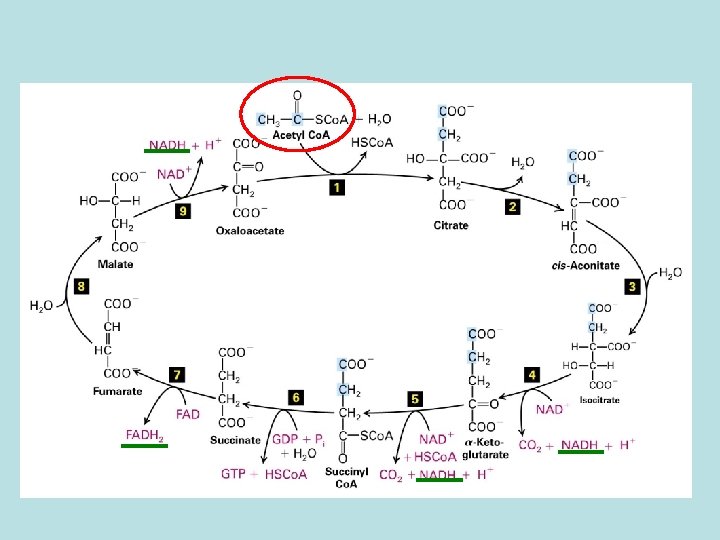
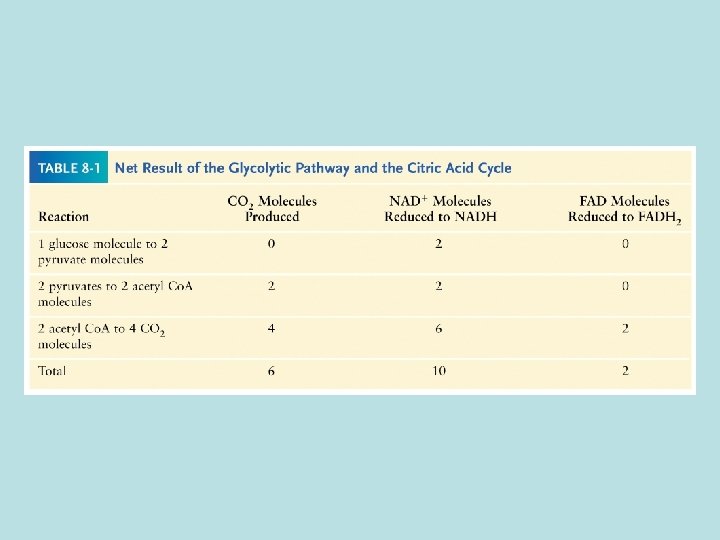

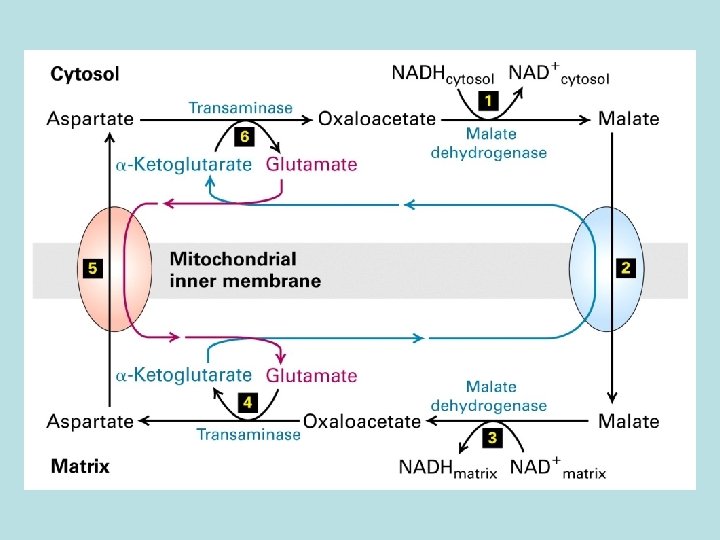
Mitochondria membrane is impermeable to NADH How does NADH enter mitochondria ? ----- malate aspartate shuttle 1. malate/ -ketoglutarate antiport 2. glutamate/aspartate antiport NADH cytosol + NAD+ cytosol+ NADHmatrix
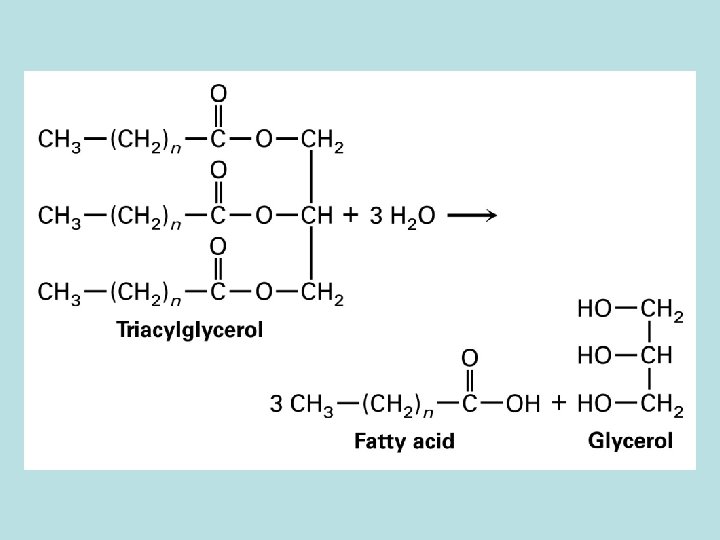
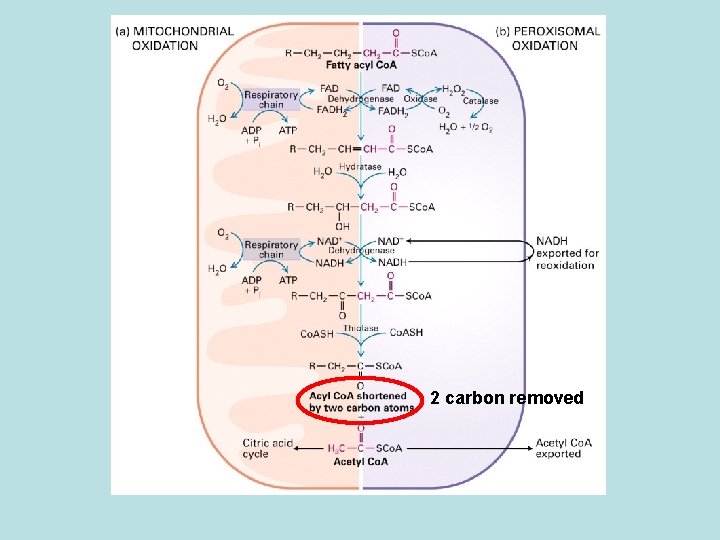
2 carbon removed
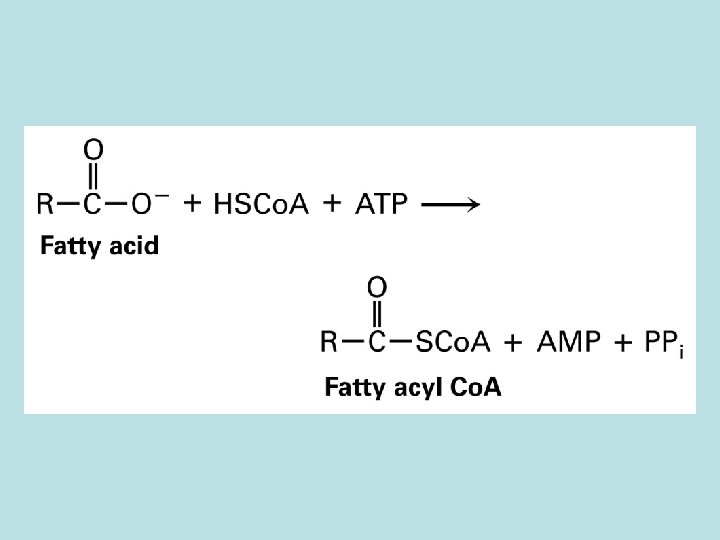

Peroxisome single membrane oxidize long chain fatty acid ·generate no ATP · electron from FADH 2 produced during ·oxidation of FA was transferred O 2 and generate H 2 O 2 ·NADH is exporeted to cytosol and reoxidize ·No citric acid cycle---acetyl co. A genetated was send to cytosol for cholesterol synthesis

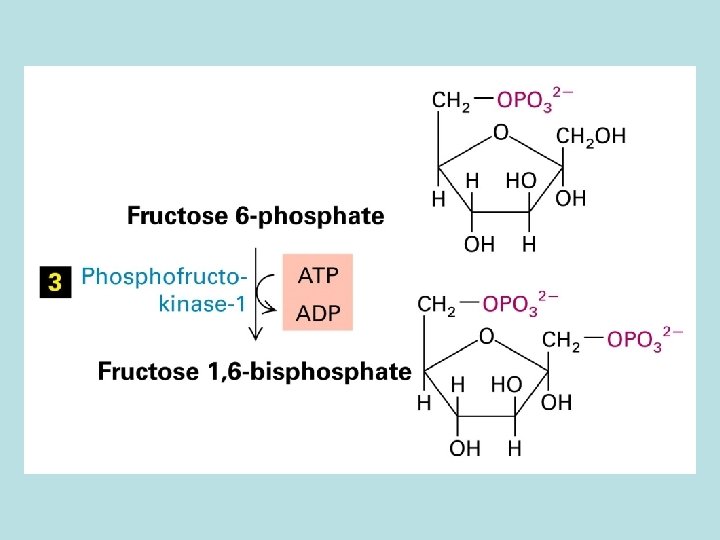
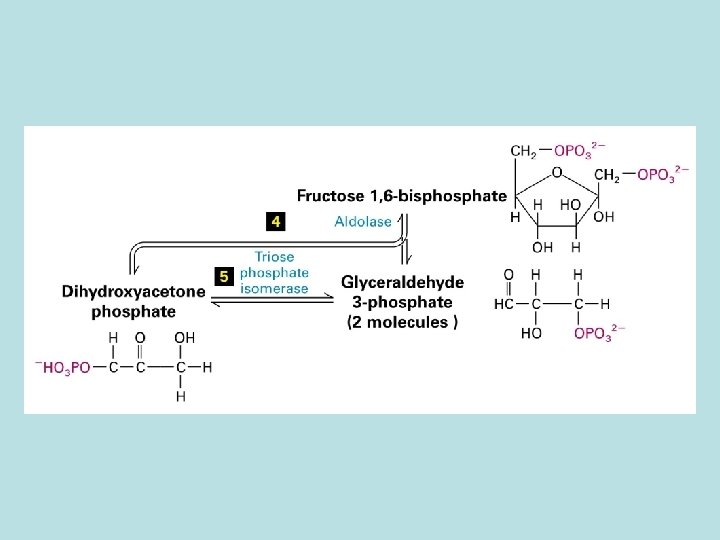
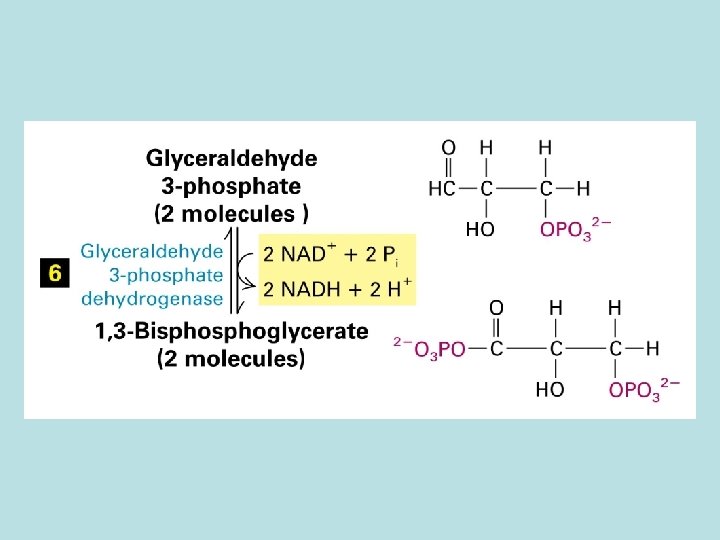
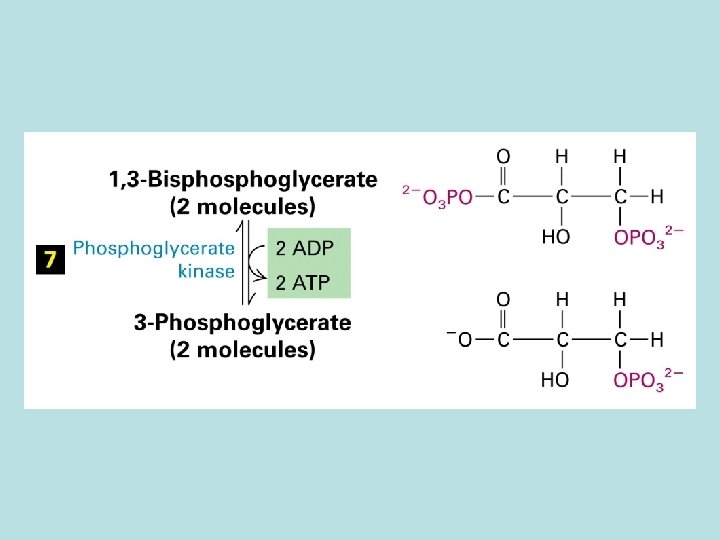
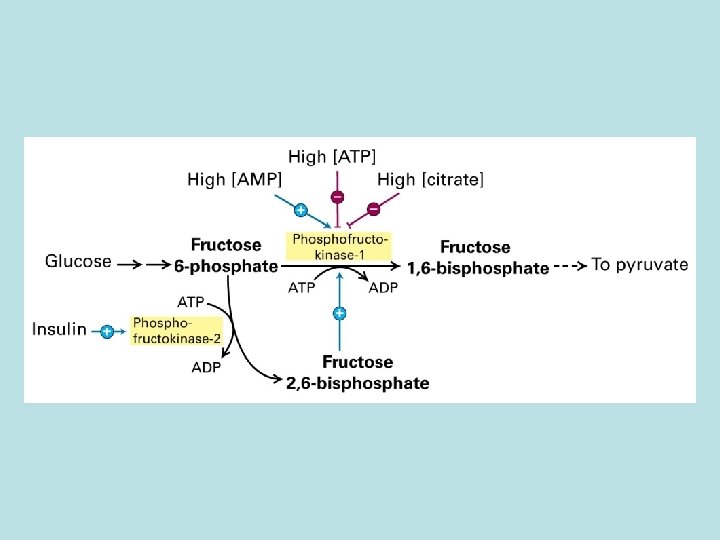
Allosteric control of glucose metabolism 1. Hexoinase: inhibited by glucose 6 -phosphate 2. Pyruvate kinase: inhibited by ATP 3. Phosphofructokinase : 4. rate limiting enzyme of glycolic pathway 5. inhibited by ATP and citrate 6. stimulated by insulin 7. activated by fructose 2. 6. biphosphate( feed 8. forward control)
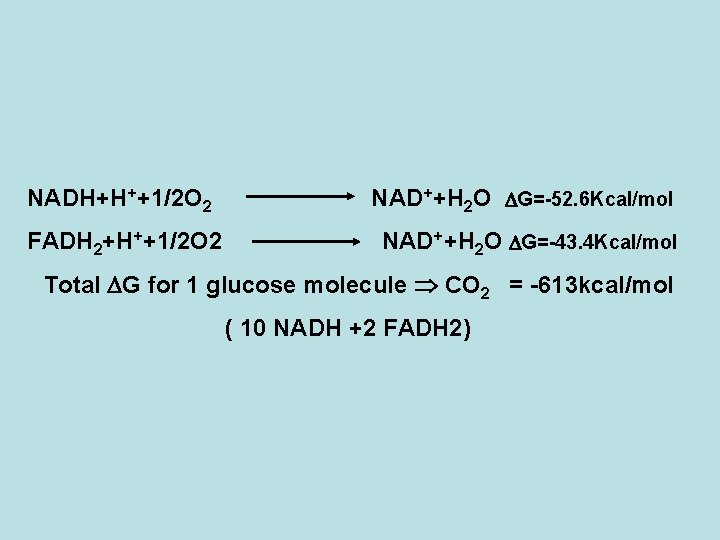
NADH+H++1/2 O 2 FADH 2+H++1/2 O 2 NAD++H 2 O G=-52. 6 Kcal/mol NAD++H 2 O G=-43. 4 Kcal/mol Total G for 1 glucose molecule CO 2 = -613 kcal/mol ( 10 NADH +2 FADH 2)
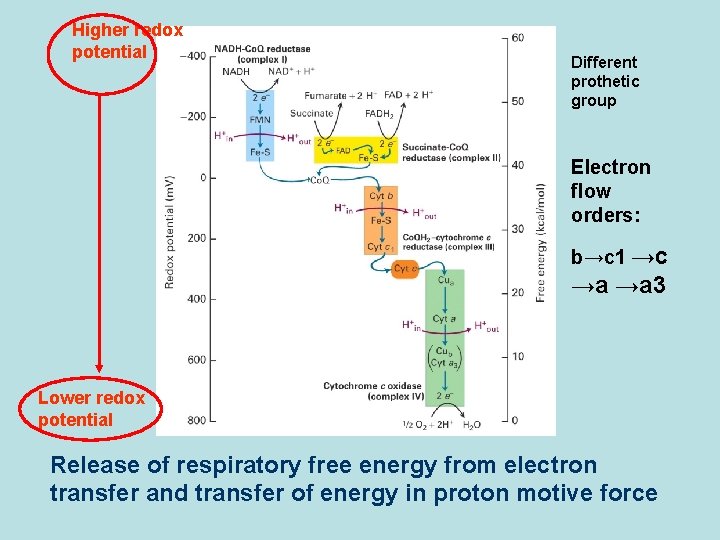
Higher redox potential Different prothetic group Electron flow orders: b→c 1 →c →a →a 3 Lower redox potential Release of respiratory free energy from electron transfer and transfer of energy in proton motive force

Electron transfer by prothetic protein in electron transport chain Fe 3++e- Fe 2+
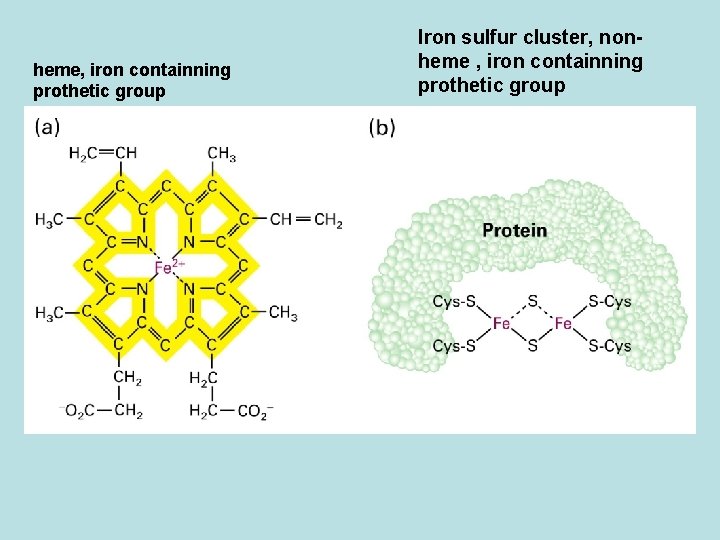
heme, iron containning prothetic group Iron sulfur cluster, nonheme , iron containning prothetic group
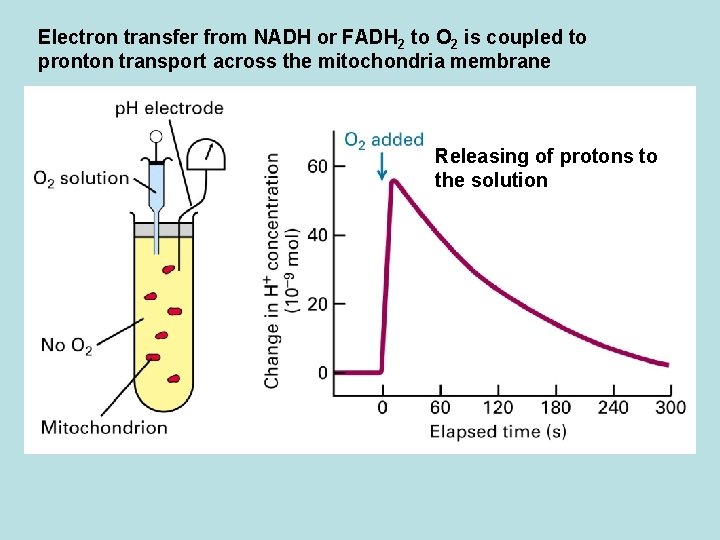
Electron transfer from NADH or FADH 2 to O 2 is coupled to pronton transport across the mitochondria membrane Releasing of protons to the solution
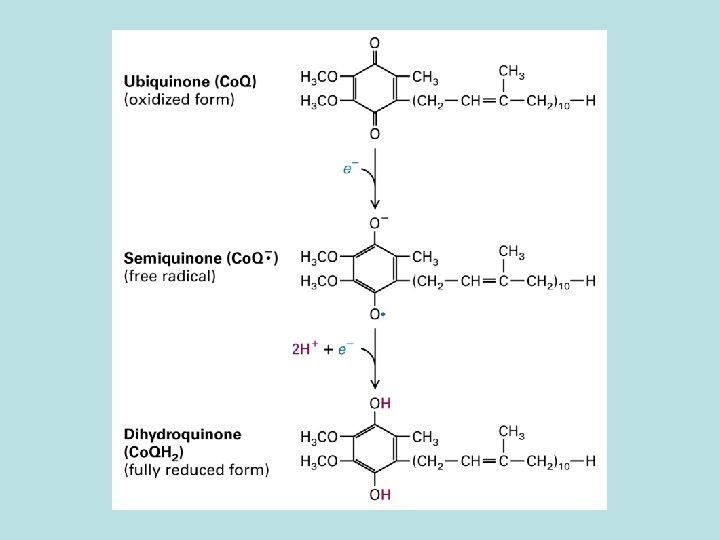
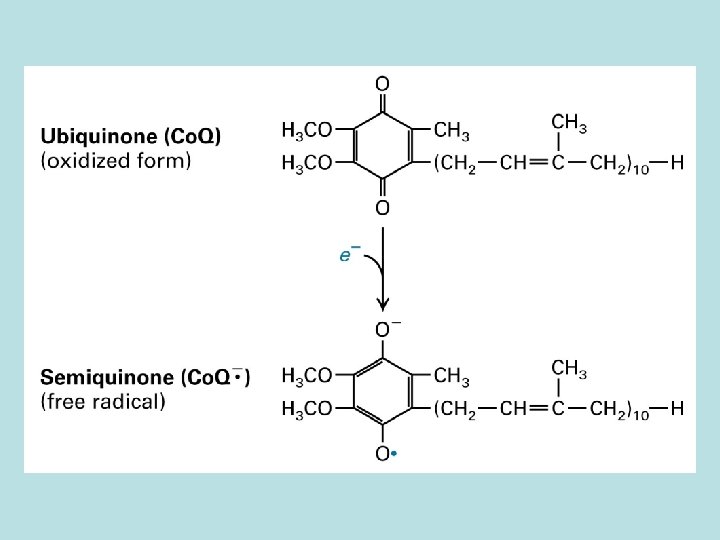
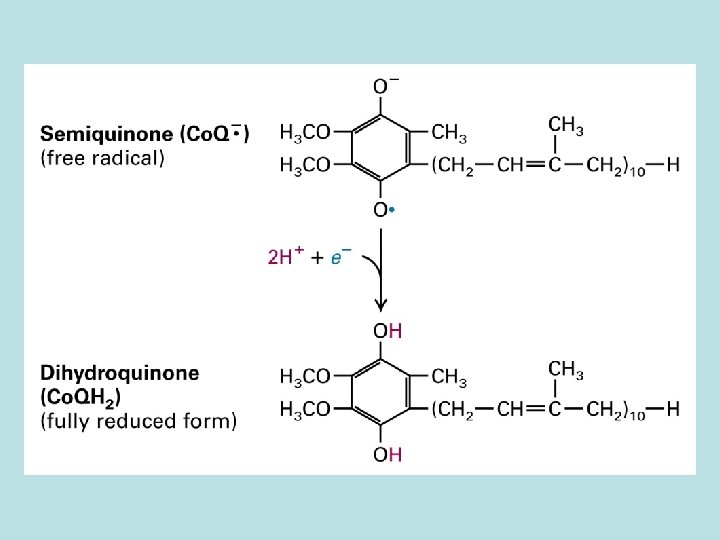
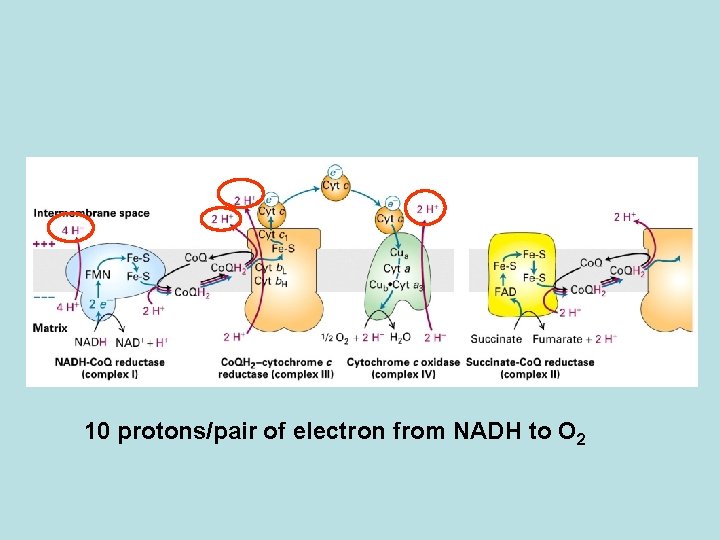
10 protons/pair of electron from NADH to O 2
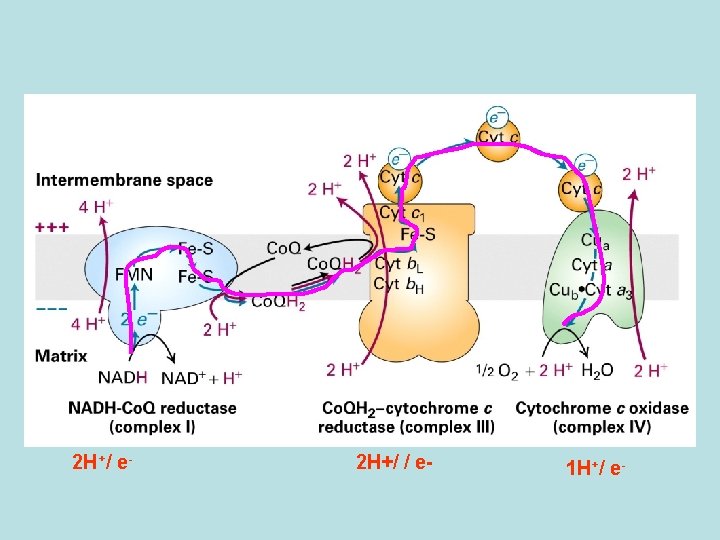
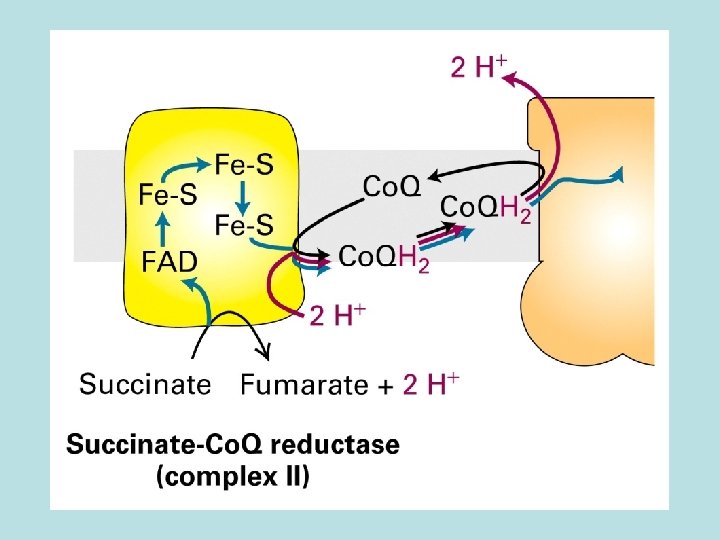
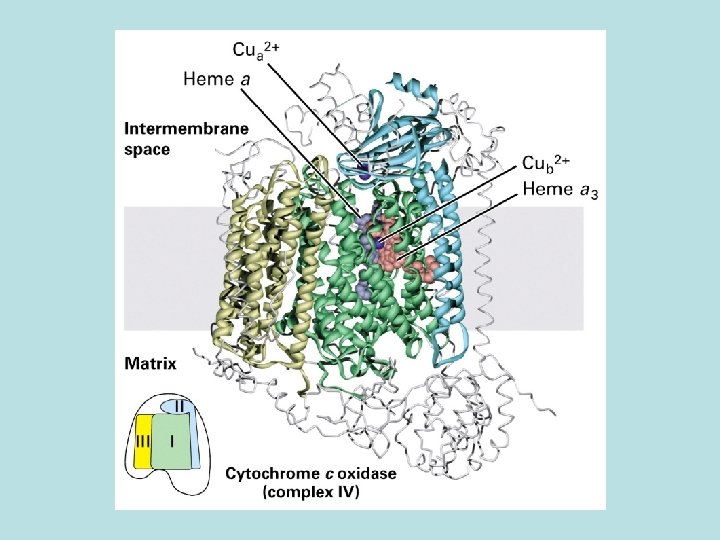
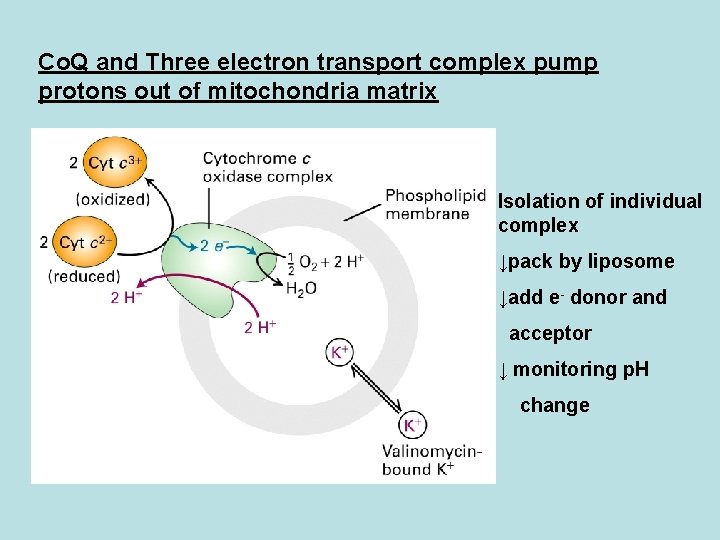
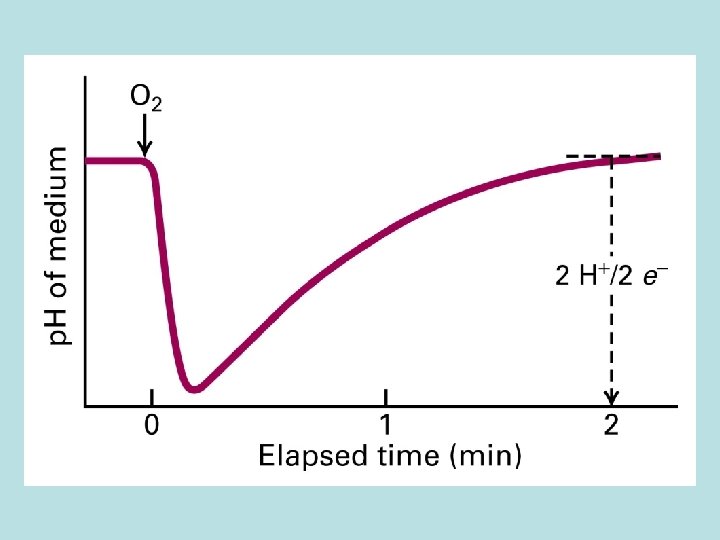
Co. Q and Three electron transport complex pump protons out of mitochondria matrix Isolation of individual complex ↓pack by liposome ↓add e- donor and acceptor ↓ monitoring p. H change
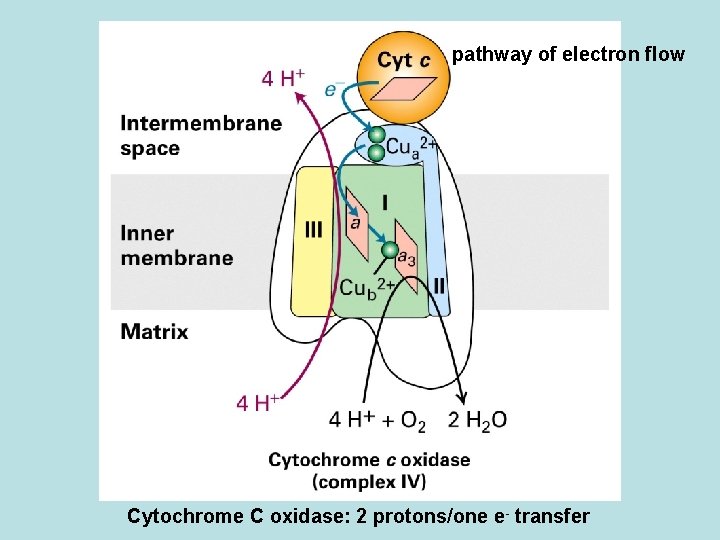
pathway of electron flow Cytochrome C oxidase: 2 protons/one e- transfer
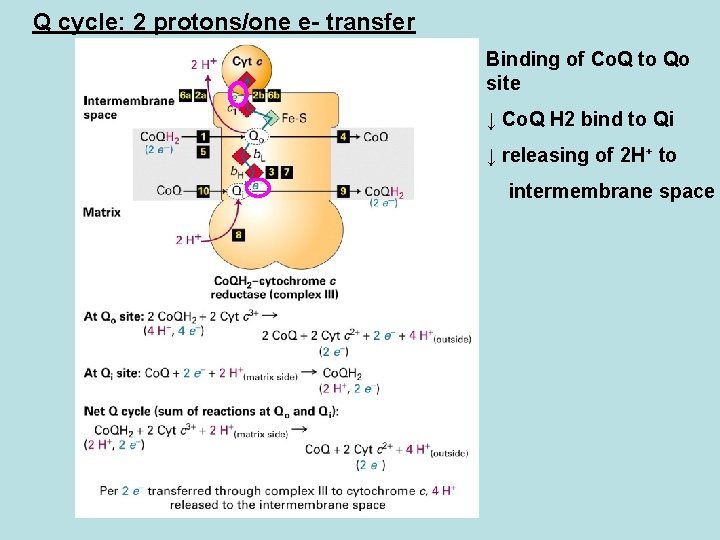
Q cycle: 2 protons/one e- transfer Binding of Co. Q to Qo site ↓ Co. Q H 2 bind to Qi ↓ releasing of 2 H+ to intermembrane space
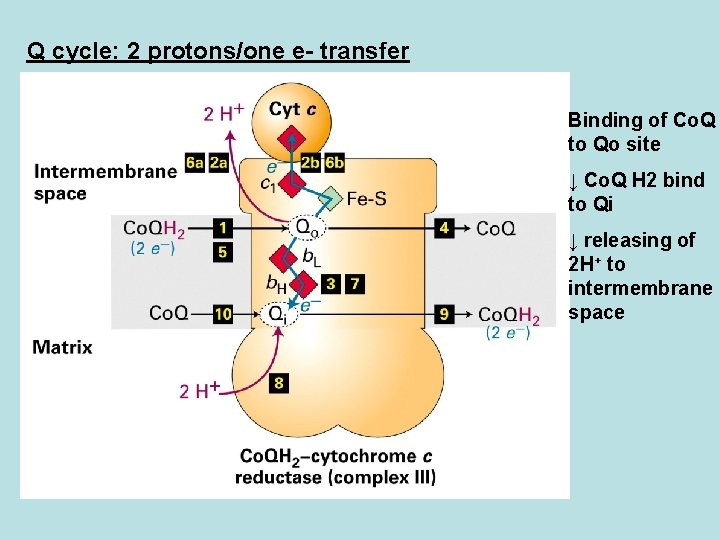
Q cycle: 2 protons/one e- transfer Binding of Co. Q to Qo site ↓ Co. Q H 2 bind to Qi ↓ releasing of 2 H+ to intermembrane space

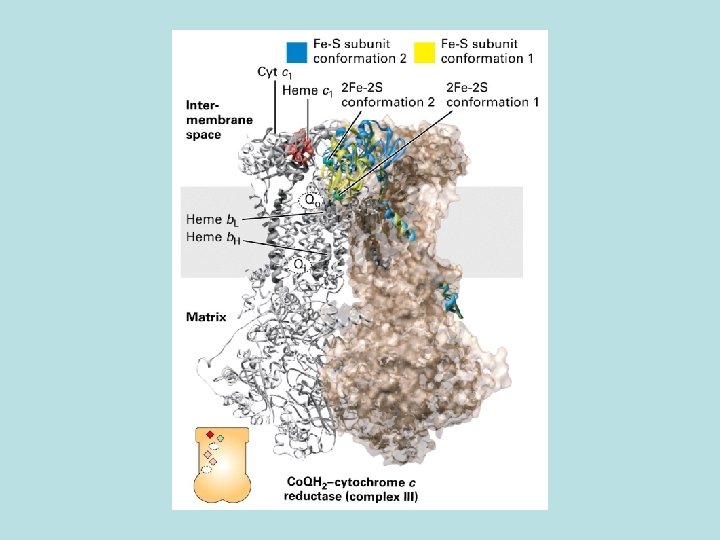
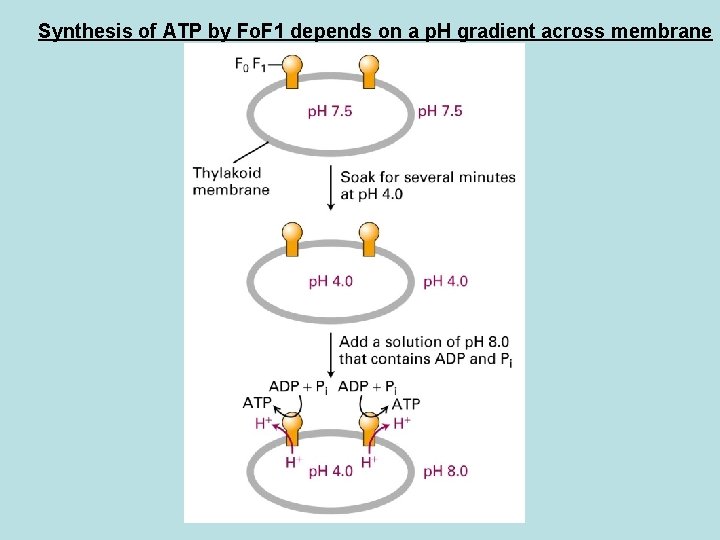
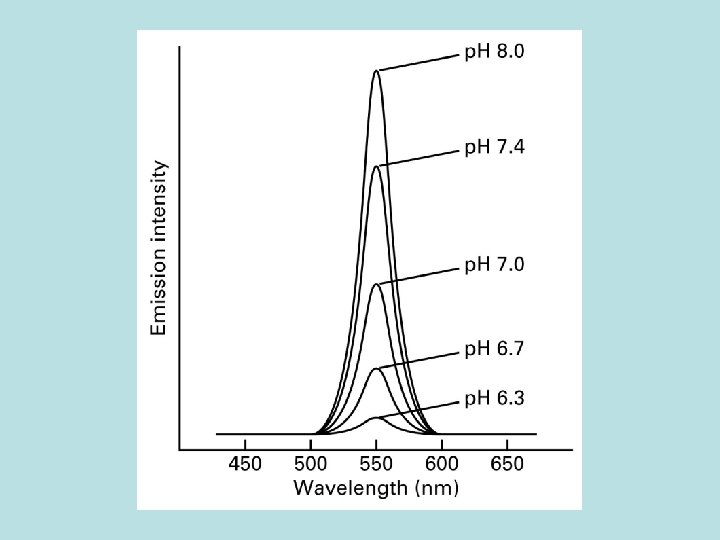
Synthesis of ATP by Fo. F 1 depends on a p. H gradient across membrane
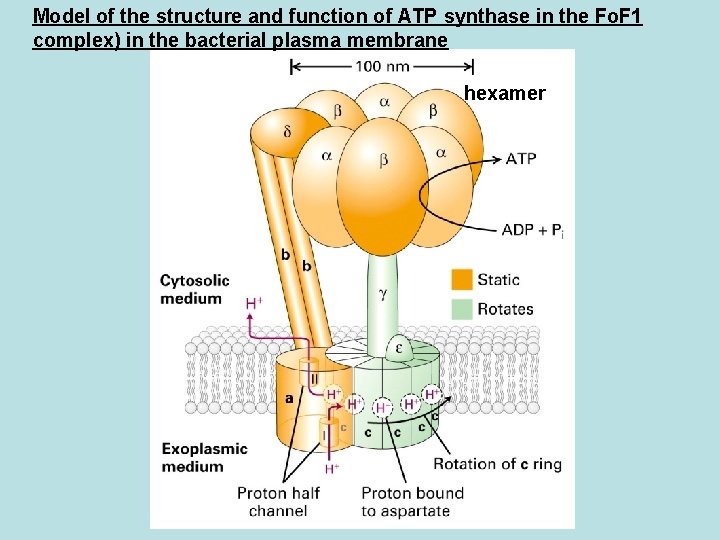
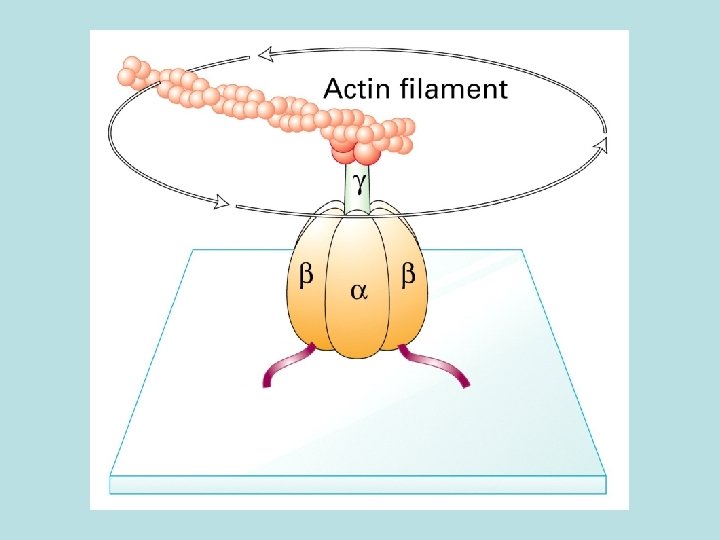
Model of the structure and function of ATP synthase in the Fo. F 1 complex) in the bacterial plasma membrane hexamer
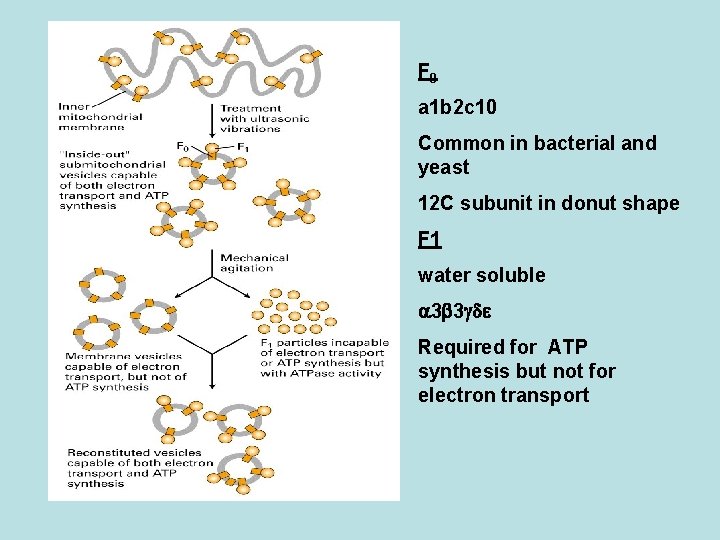
F 0 a 1 b 2 c 10 Common in bacterial and yeast 12 C subunit in donut shape F 1 water soluble 3 3 Required for ATP synthesis but not for electron transport
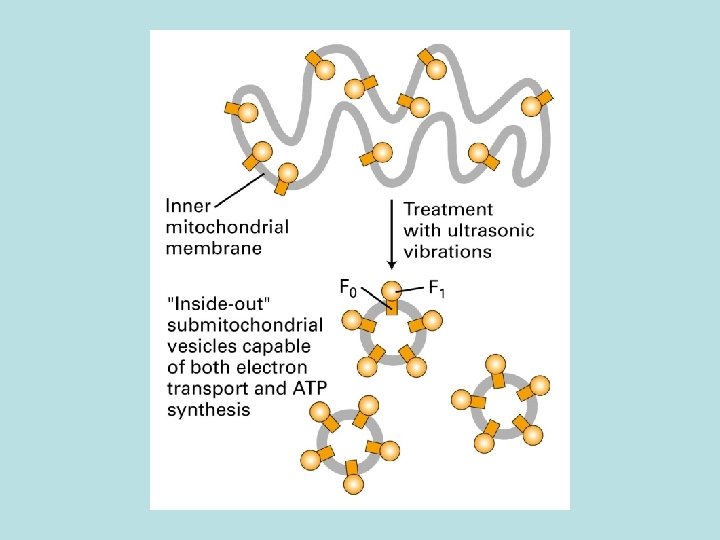
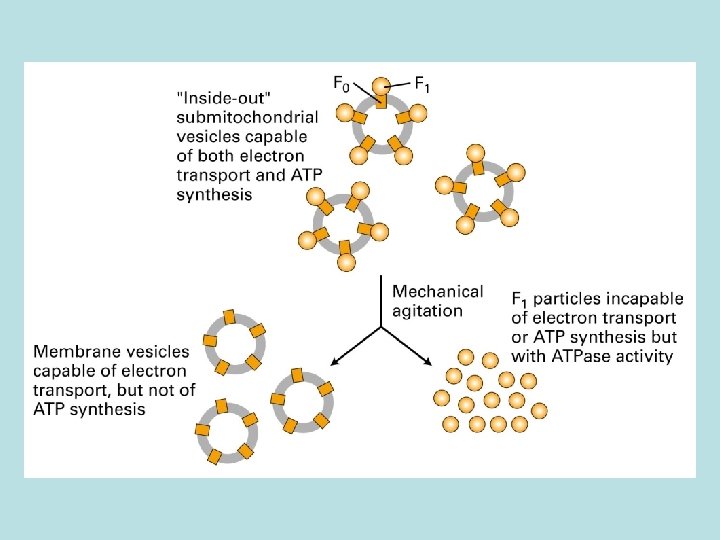
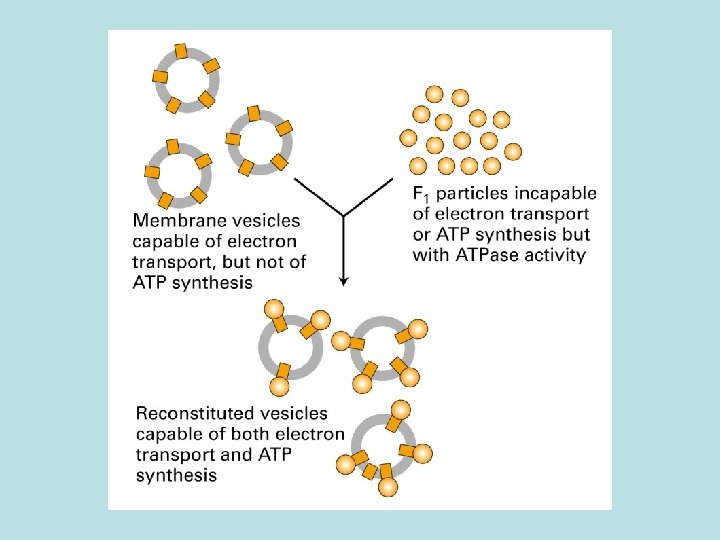
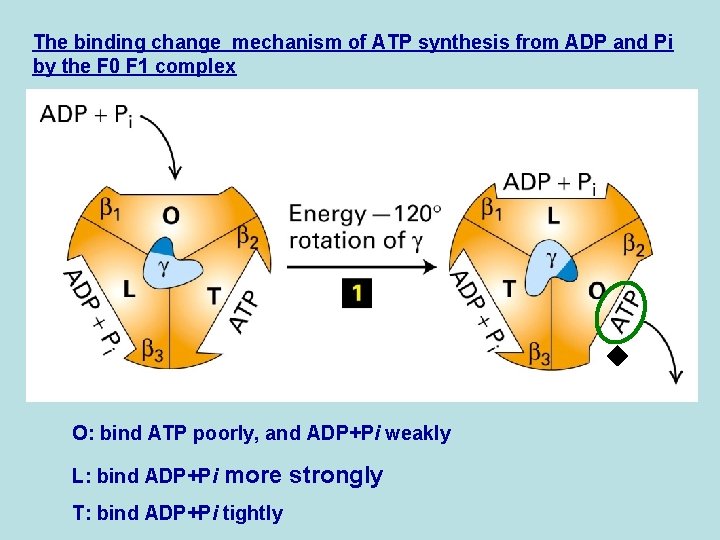
The binding change mechanism of ATP synthesis from ADP and Pi by the F 0 F 1 complex O: bind ATP poorly, and ADP+Pi weakly L: bind ADP+Pi more strongly T: bind ADP+Pi tightly
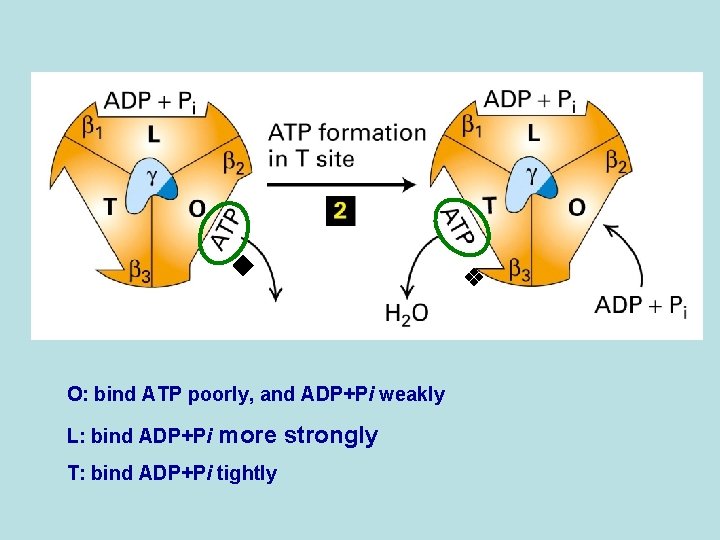
O: bind ATP poorly, and ADP+Pi weakly L: bind ADP+Pi more strongly T: bind ADP+Pi tightly
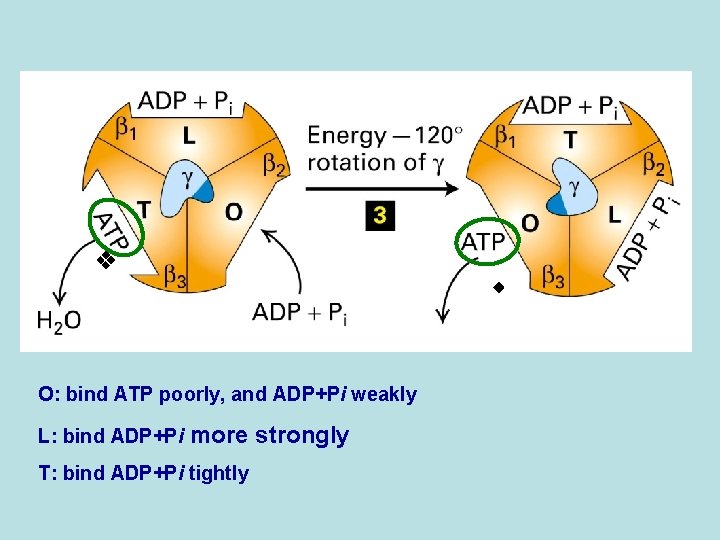
O: bind ATP poorly, and ADP+Pi weakly L: bind ADP+Pi more strongly T: bind ADP+Pi tightly
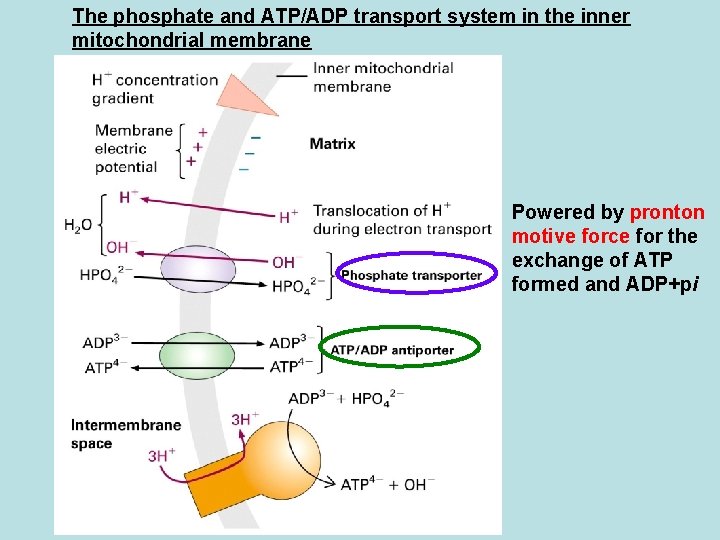
The phosphate and ATP/ADP transport system in the inner mitochondrial membrane Powered by pronton motive force for the exchange of ATP formed and ADP+pi

Respiratory control · Rate of Motochondria oxidation normally depends on ATP level Oxidation of FADH 2 and NADH occur ed only there is a source of ADP and Pi ·Coupling of NADH and FADH 2 oxidation and proton transport across inner mitochondrial membrane and ATP synthesis is important in maintaining the membrane electro potential DNP: uncoupler of mitochondria membrane potential shuttle proton from intermembrane space to matrix and abolish ATP synthesis

Thermogenin A natural uncoupler in brown fat mitochondrial membrane Oxidize NADH and convert the energy to heat Slow in proton transport a. a. sequence similar to ATP/ADP anti-port
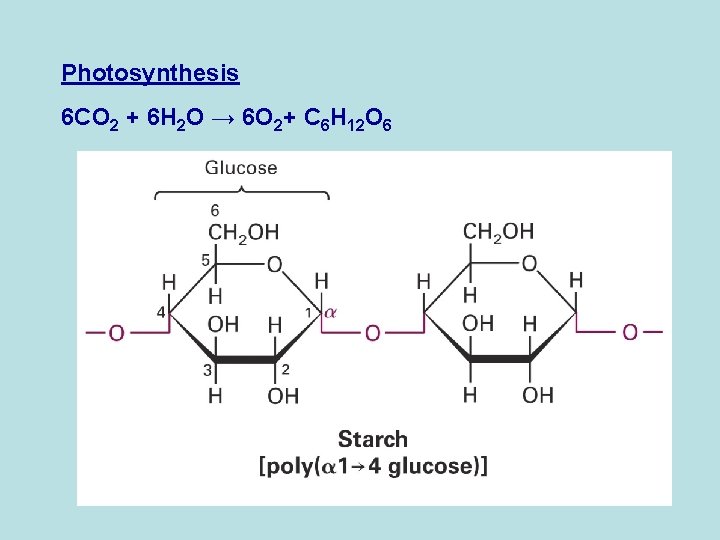
Photosynthesis 6 CO 2 + 6 H 2 O → 6 O 2+ C 6 H 12 O 6
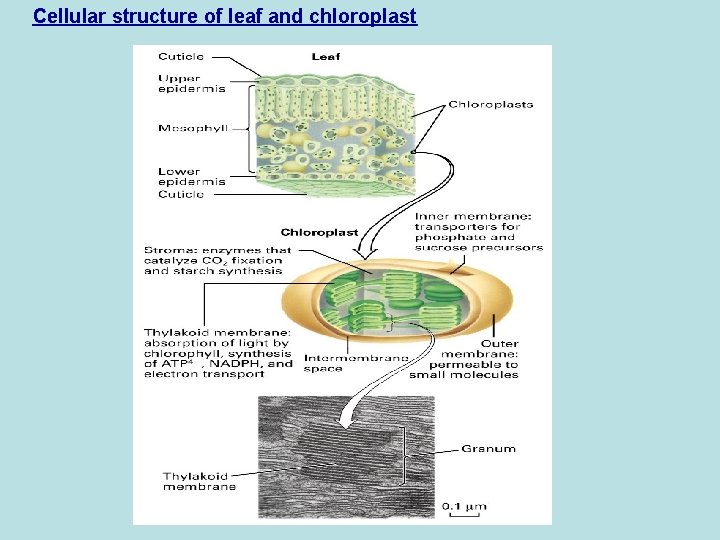
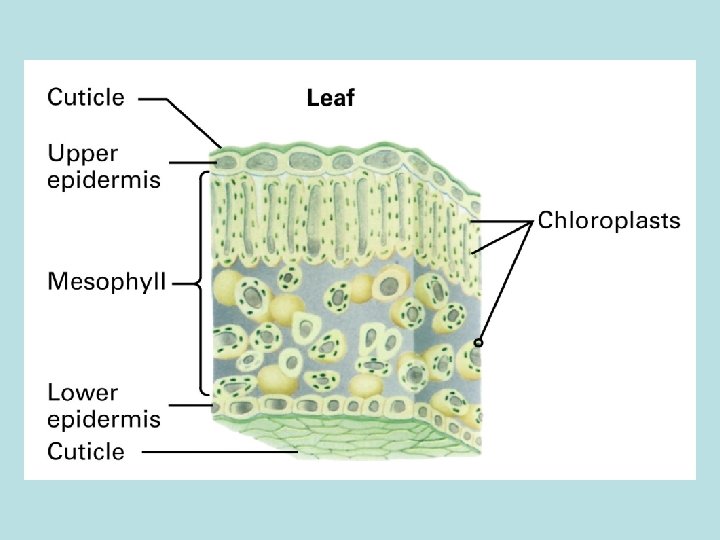
Cellular structure of leaf and chloroplast
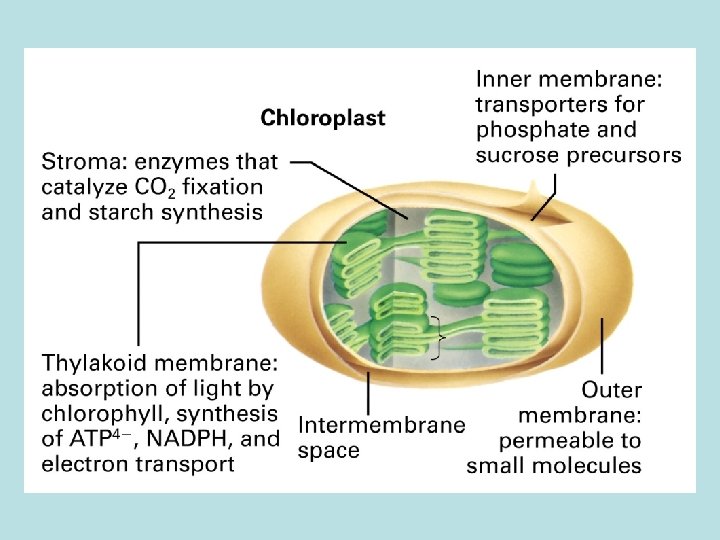
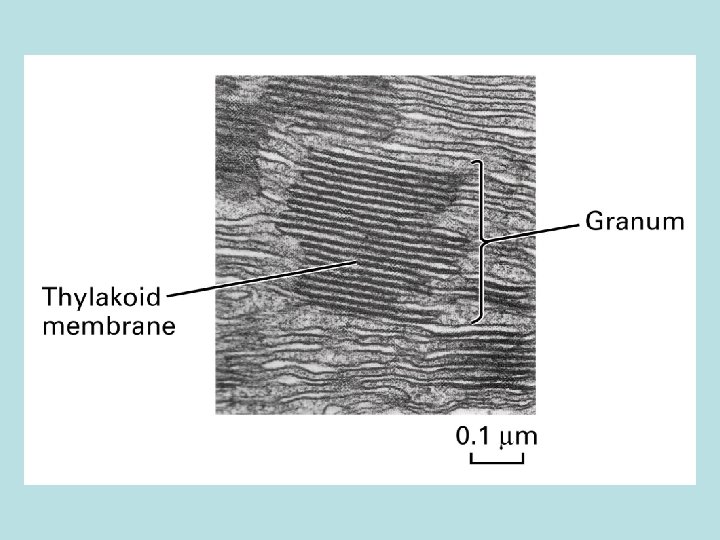

Four stages in photosynthesis 1. Absorption of light 2. light absorpthin by chlorophyll on thylakoid membrane 2 H 2 O → O 2 + 4 H++ 4 e- transfer of e- to quinone Q 2. Electron transport and generation of a proton motive force 2 H 2 O+ 2 NADP+ light 2 H+ + 2 NADPH + O 2 3. Synthesis of ATP 4. Carbon fixation 5. 6. synthesis of polymer of 6 -C sugars from CO 2 and H 2 O

Phtosystem 1. Light reaction center 2. chlorophyll a : function both as light reaction and harvesting 3. chlorophyll b : seen in vascular plant 4. carotenoid : seen in plant and bacteria 2. Light harvesting complex( LHCs)
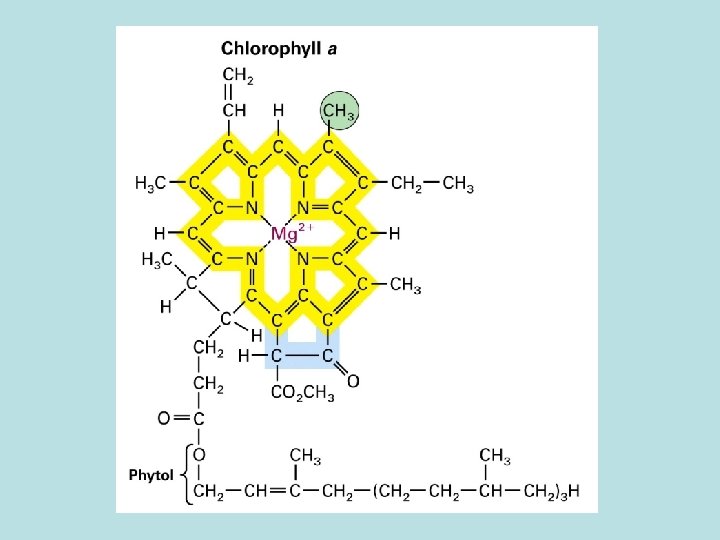
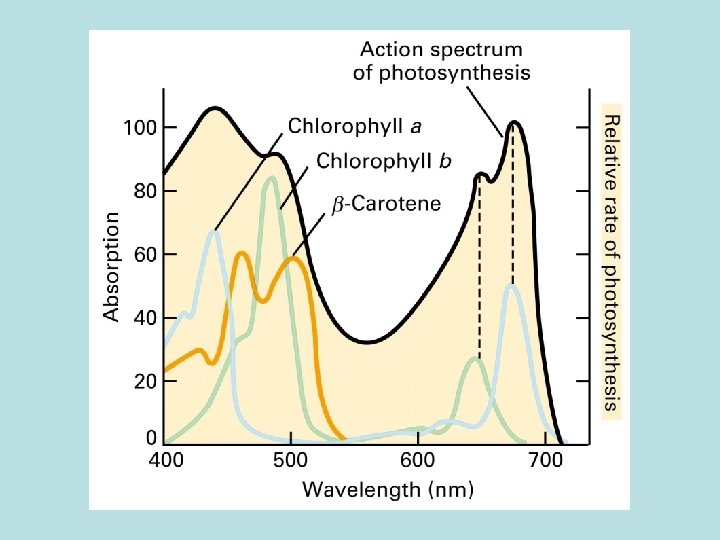
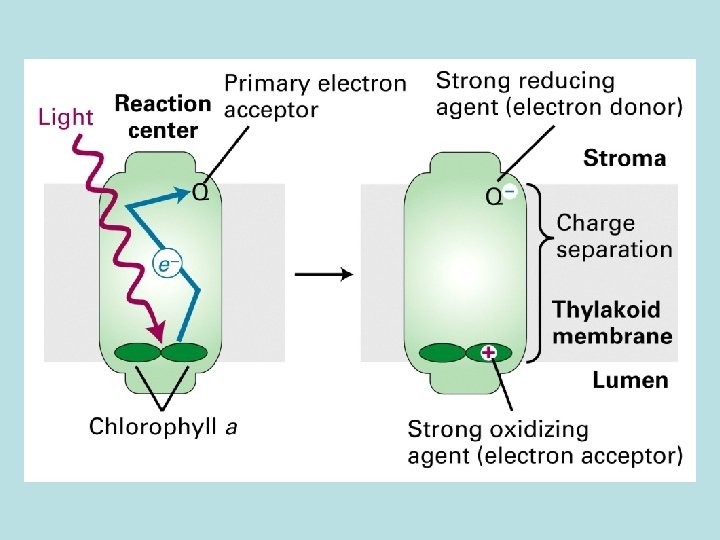
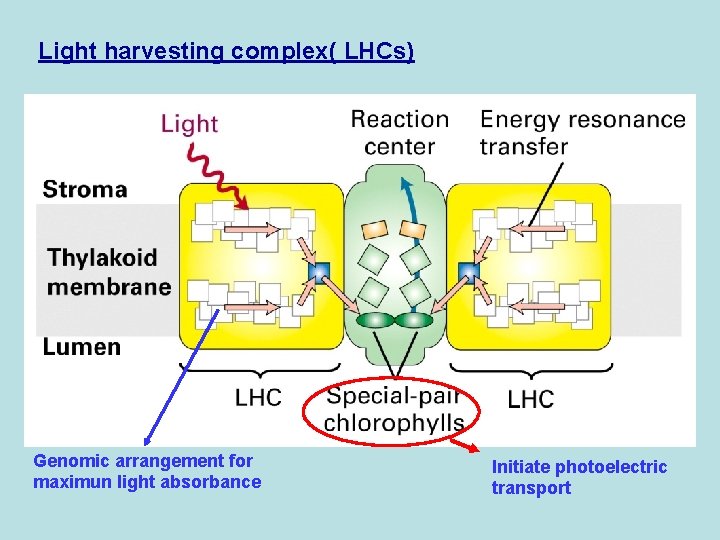
Light harvesting complex( LHCs) Genomic arrangement for maximun light absorbance Initiate photoelectric transport
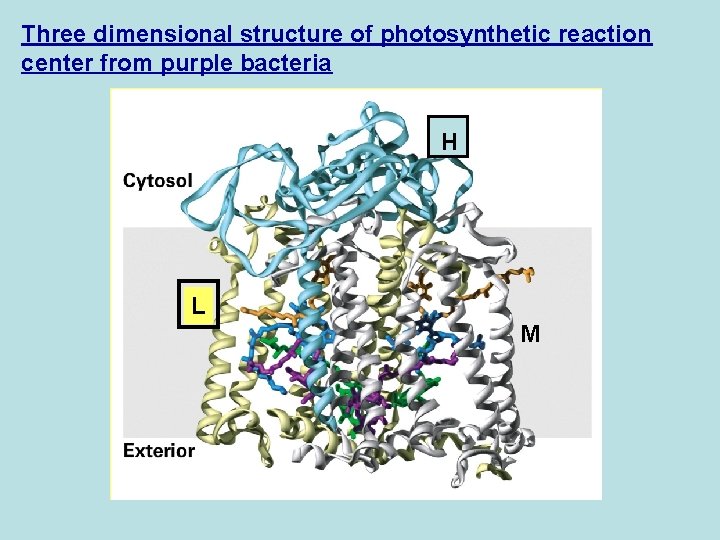
Three dimensional structure of photosynthetic reaction center from purple bacteria H L M
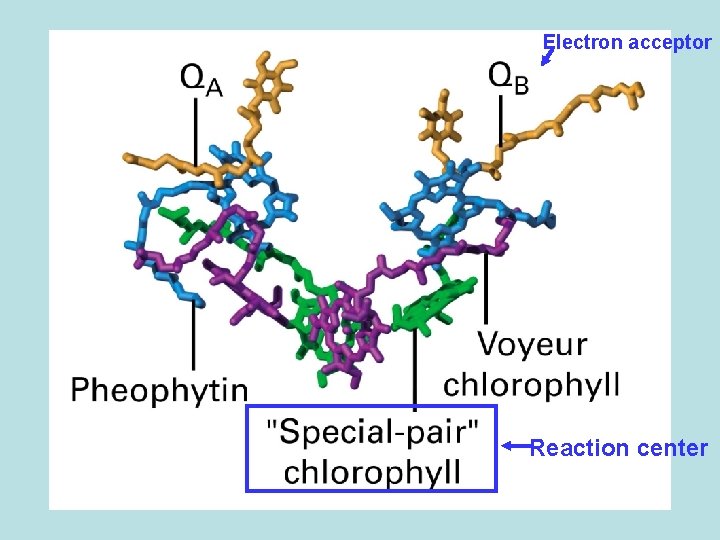
Electron acceptor Reaction center
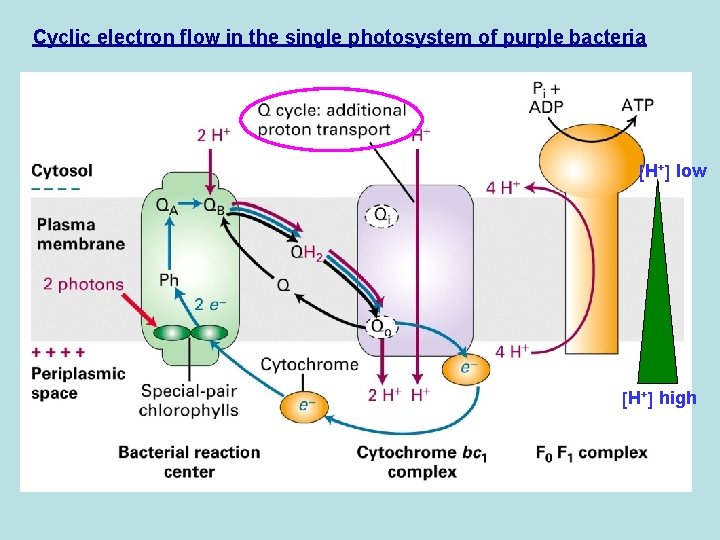
Cyclic electron flow in the single photosystem of purple bacteria H+ low H+ high
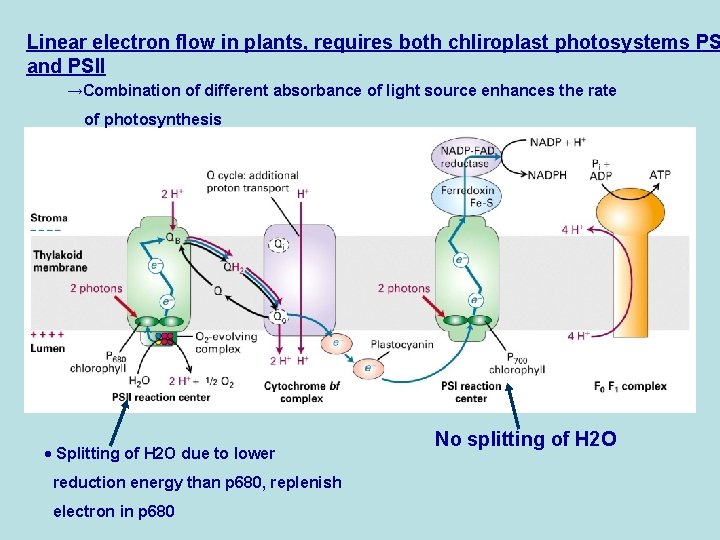
Linear electron flow in plants, requires both chliroplast photosystems PS and PSII →Combination of different absorbance of light source enhances the rate of photosynthesis · Splitting of H 2 O due to lower reduction energy than p 680, replenish electron in p 680 No splitting of H 2 O
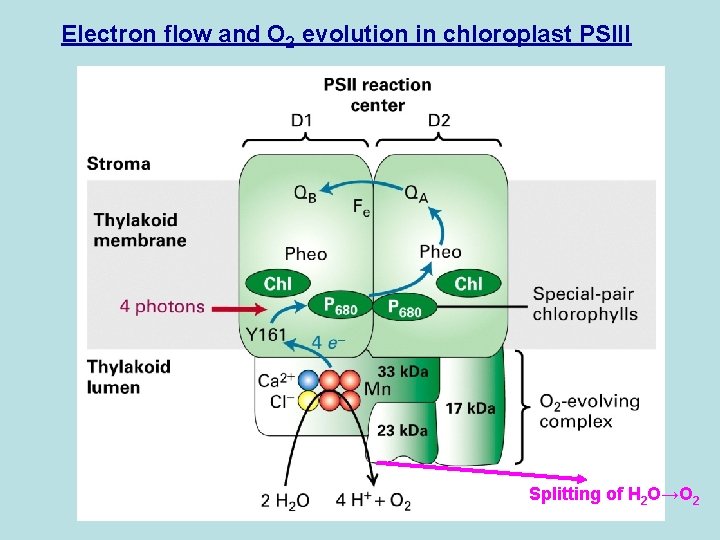
Electron flow and O 2 evolution in chloroplast PSIII Splitting of H 2 O→O 2
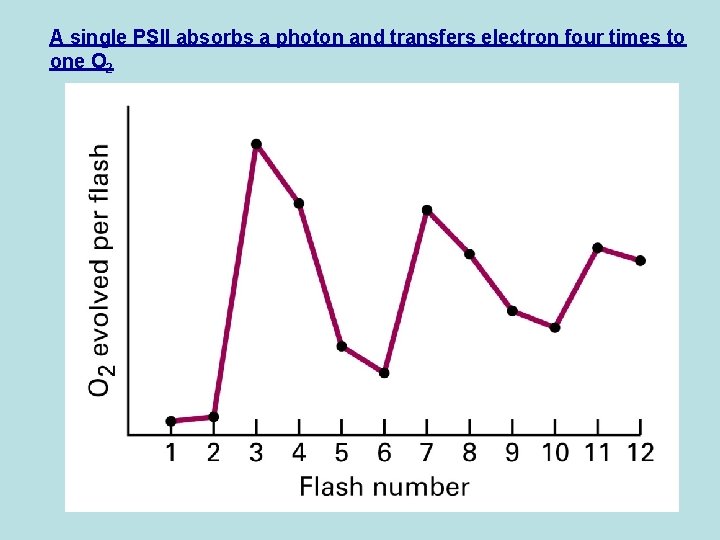
A single PSII absorbs a photon and transfers electron four times to one O 2
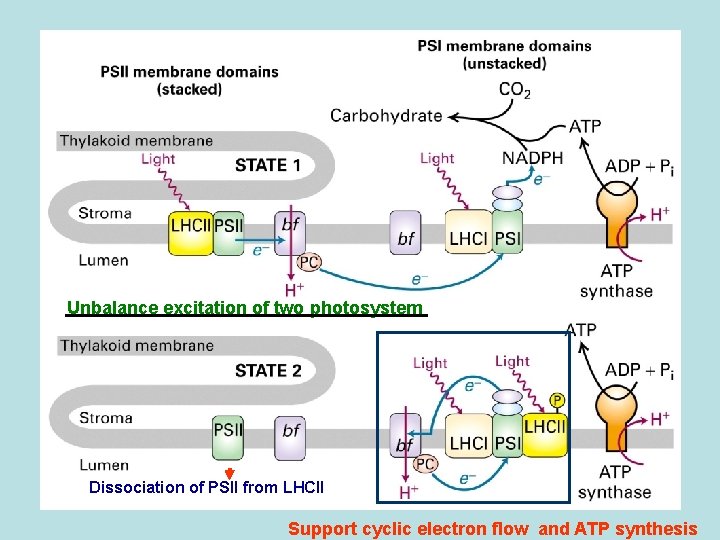
Unbalance excitation of two photosystem Dissociation of PSII from LHCII Support cyclic electron flow and ATP synthesis
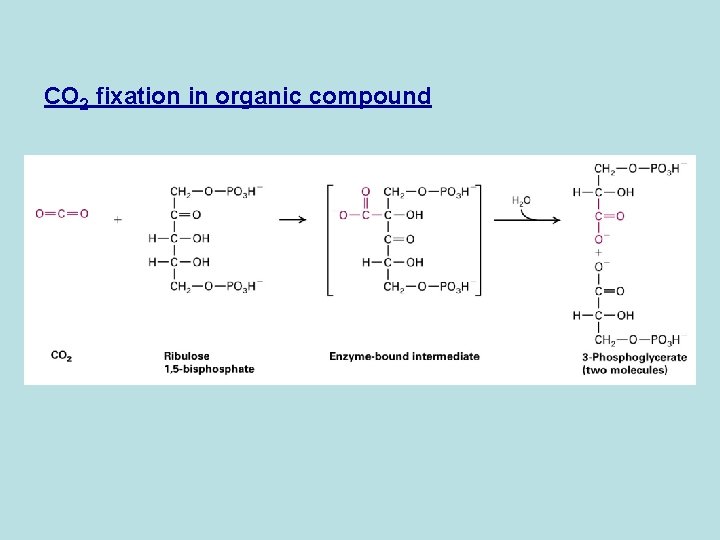
CO 2 fixation in organic compound
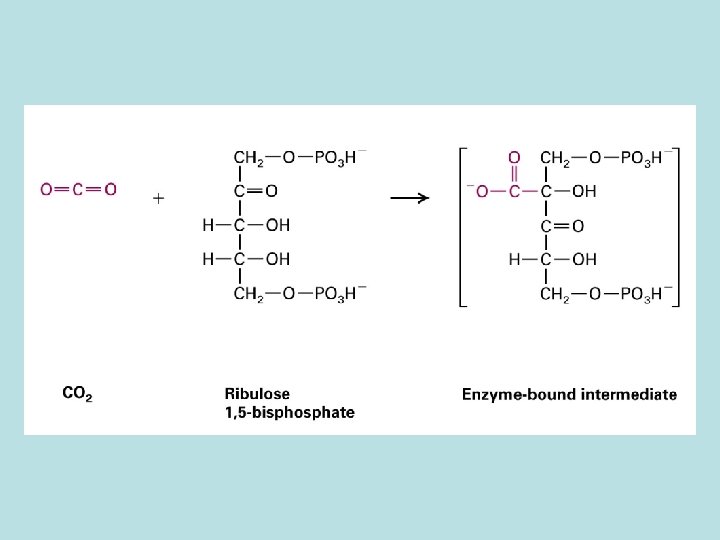
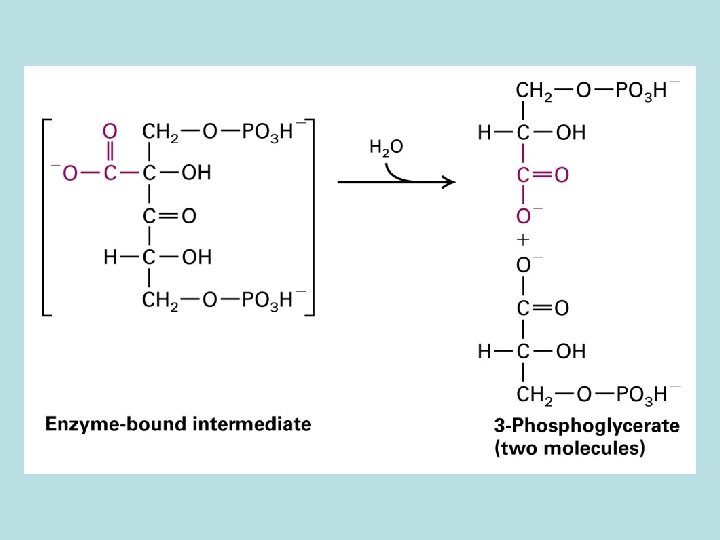
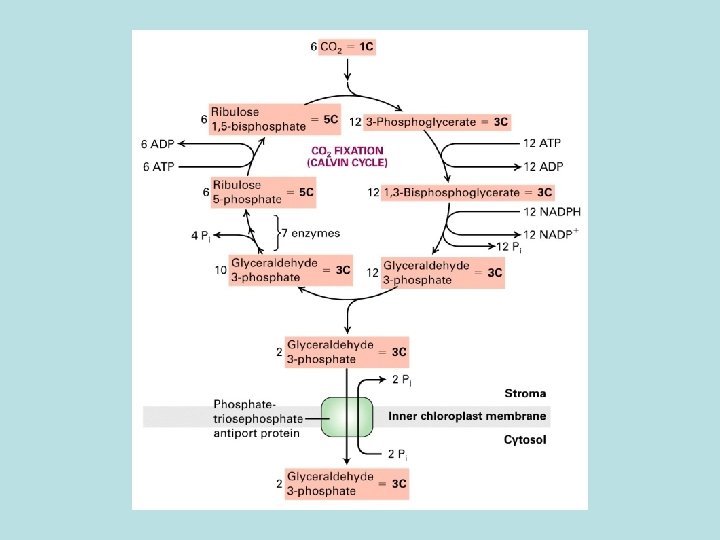
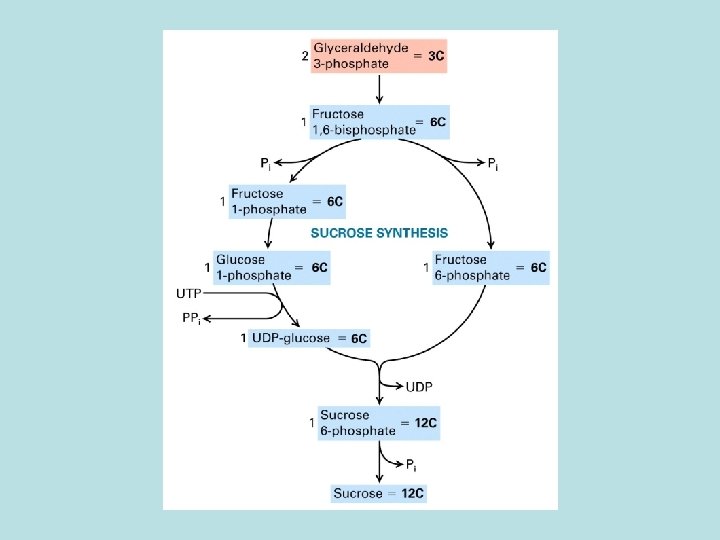
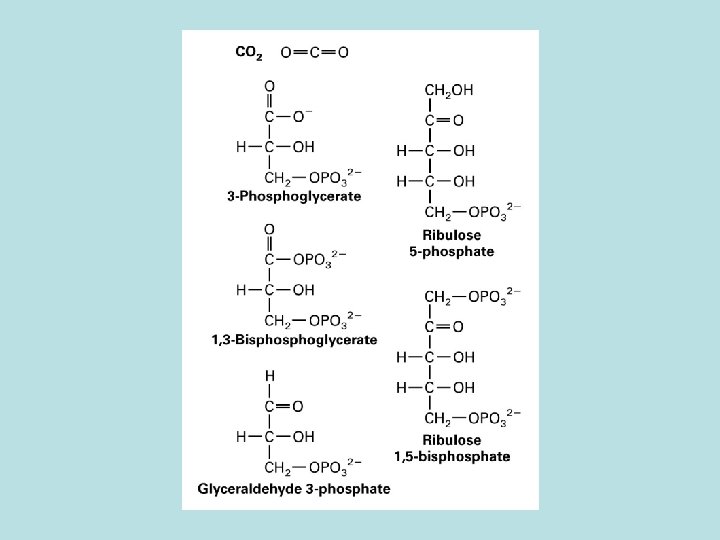
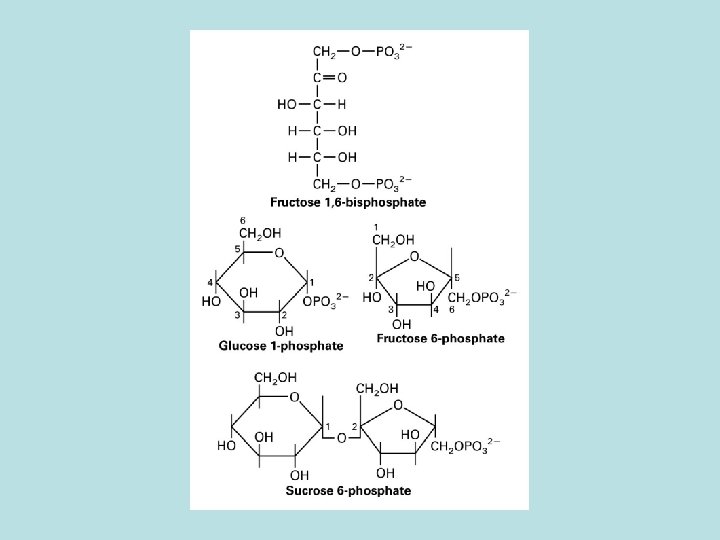
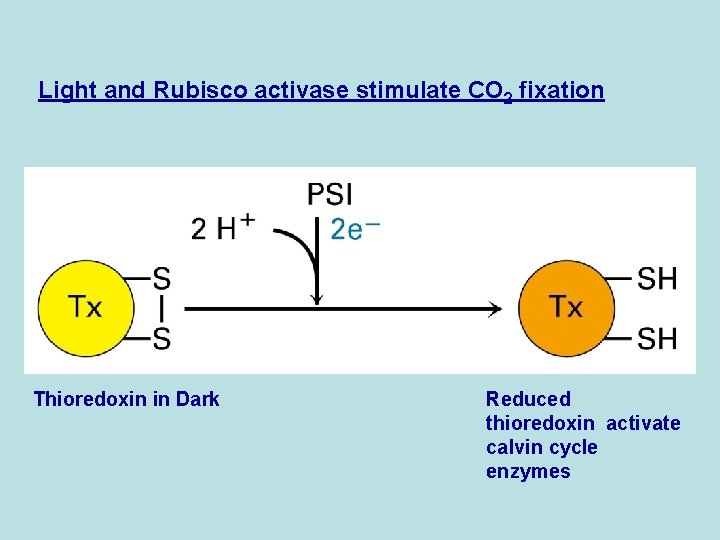
Light and Rubisco activase stimulate CO 2 fixation Thioredoxin in Dark Reduced thioredoxin activate calvin cycle enzymes
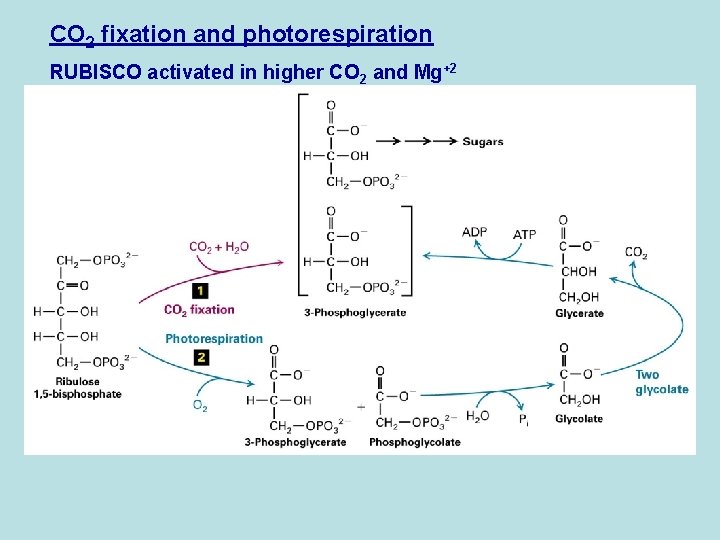
CO 2 fixation and photorespiration RUBISCO activated in higher CO 2 and Mg+2
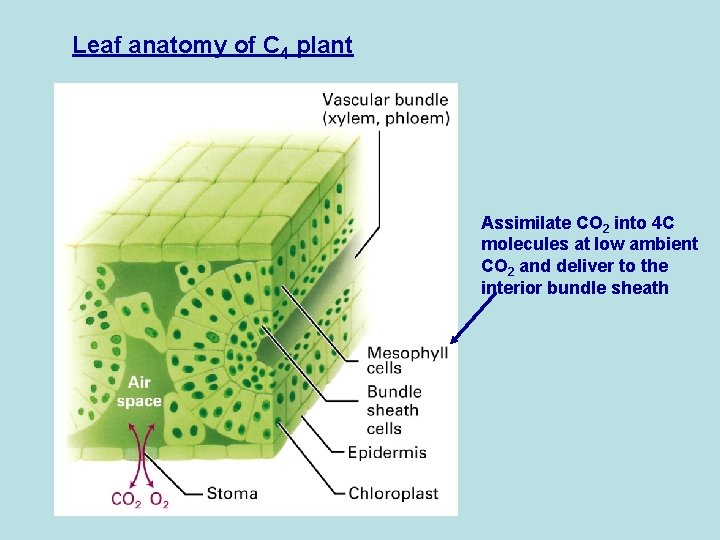
Leaf anatomy of C 4 plant Assimilate CO 2 into 4 C molecules at low ambient CO 2 and deliver to the interior bundle sheath
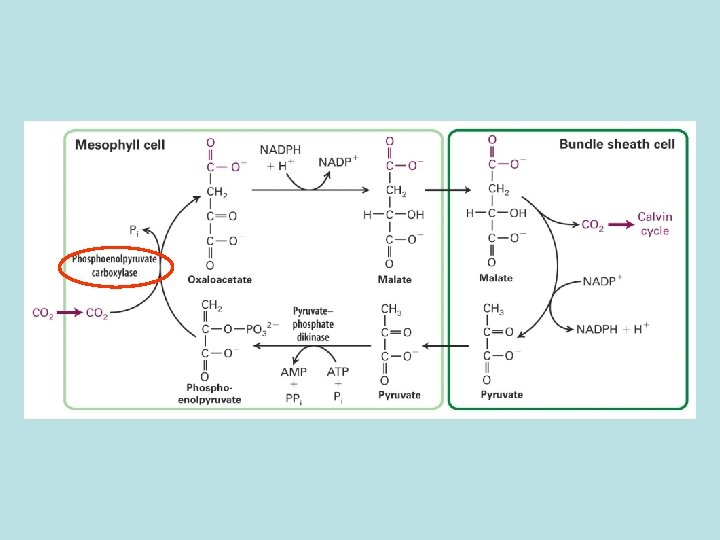
- Slides: 106