Haplotype Blocks An Overview A Polanski Department of

Haplotype Blocks An Overview A. Polanski Department of Statistics Rice University

Key Papers 1. 2. 3. N. Patil et al. , (2001), Blocks of Limited Haplotype Diversity Revealed by High-Resolution Scanning of Human Chromosome 21, Science, vol. 294, pp. 17191723 N. Wang et al. , (2002), Distribution of Recombination Crossovers and the Origin of Haplotype Blocks: The Interplay of Population History, Recombination and Mutation, Am. J. Hum. Genet. , vol. 71, pp. 1227 -1234. K. Zhang et al. , (2002), A Dynamic Programming Algorithm for Haplotype Block Partitioning, PNAS, vol. 99, pp. 7335 -7339

Supplementary Papers 1. 2. 3. R. Hudson, N. Kaplan, (1985), Statistical Properties of the Number of Recombination Events in The History of a Sample of DNA sequences, Genetics, vol. 111, pp. 147 -164 R. Hudson, 2002, Generating Samples under a Wright. Fisher Neutral Model of Genetic Variation, Bioinformatics, vol. 18, pp. 337 -338 D. Reich et al. , (2001), Linkage Disequilibrium in the Human Genome, Nature, vol. 411, pp. 199 -204

What are Haplotype Blocks ? Haplotype block = a sequence of contiguous markers on DNA, homogeneous according to some criterion Markers = Single Nucleotide Polymorphisms (SNPs)

Data (Patil et al. 2001) Chromosome 21 Physically separated the two copies of chromosome 21 using a rodent-human somatic cell hybrid technique Sample of 20 copies of chromosome 21 (32397439 bases) Found: 35989 SNPs
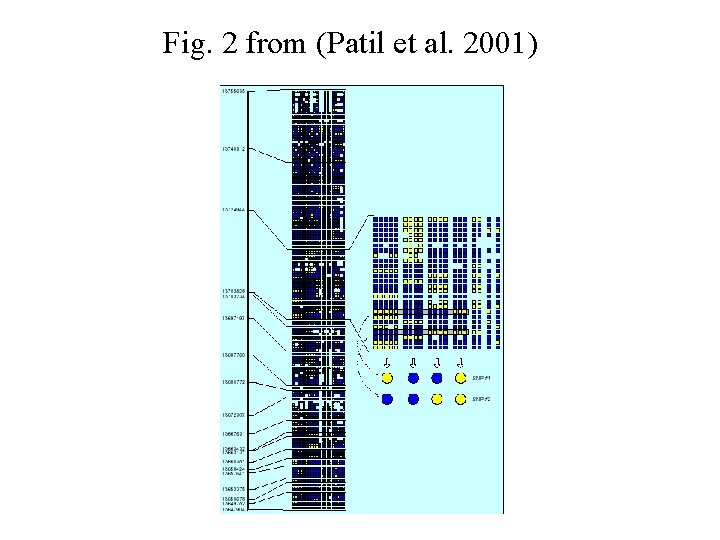
Fig. 2 from (Patil et al. 2001)
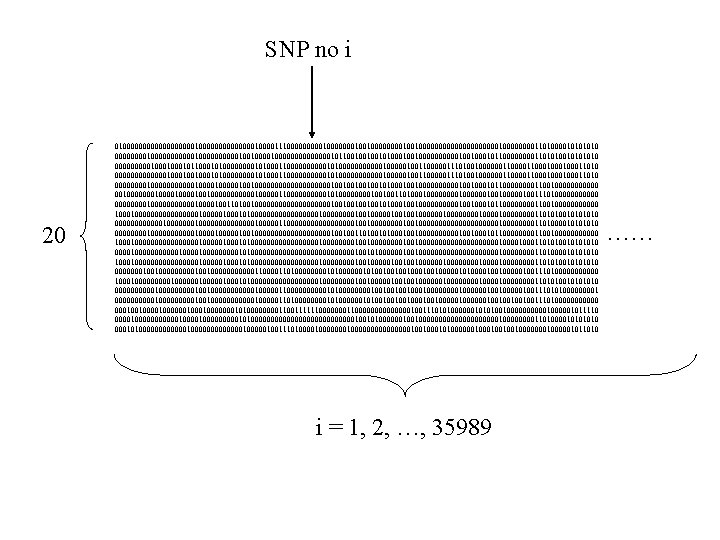
SNP no i 20 010000000001000000010000111000001000000010010000000000010000110100001010 0000000010000000000100100000000101100100100101001000001001011000011010101010 00000100010110001010000101000110000010100000010000010011000001110100100000011000100011010 000000010001010000101000110000010100000010000010011000001110100100000011000100011010 000000001000010000010010000000000100100101001000001001011000011001000000010000100100000010000011000001010000100100110100000000100100000100111010000000010000100110100100000000001001001010010000010010110000110010000000000100000101000000000100001001000001001001000000001000011010101010 0000001000000000000001100000000010010000000000000010000110100001010 000000001000010000010010000000000100100110100010010000010010110000110010000000000100000101000000000100000000100100000000001000011010101010 00001000000100000000010100000000000001001010000001001000000000010000110100001010 10000000000100000101000000000100001001000001001001000000001000011010101010 00000001001000000011000011010000101000000101001000100100000101000010010000010011101000000000100000101000000000100001001000001001001000000001000011010101010 000001000001001000000100000110000010100001001000100001000000100111010100001 00000100000100100000010000011010000101000000101001000100100000010010011101000000 0001001000000100000001010000110011111100000000000000100111010100000010101001000001000001011110 000010000001000000000101000000000000010010100000010010000000000100001101000010100000000000001000001001110100000001000000001001010000001001001000001011010 i = 1, 2, …, 35989 ……

Problems

How do we determine boundaries between blocks ? 1. Average value of standarized coefficient of linkage disequilibrium is greater than some threshold (Wang et al. 2002, Reich et al. 2001) 2. Infer sites in the sample of DNA sequences where recombination events happened in the past history (Wang et al. 2002, Hudson, 2002) 3. Chromosome coverage – minimum number of SNPs to account for majority of haplotypes (Patil et al. 2001, Zhang et al. 2002)

What evolutionary forces are responsible for haplotype blocks formation ? • • Mutation Genetic drift Recombination hot spots

Methods

Method 1 (Wang et al. 2002) Infer sites in the sample of DNA sequences where recombination events happened in the past history
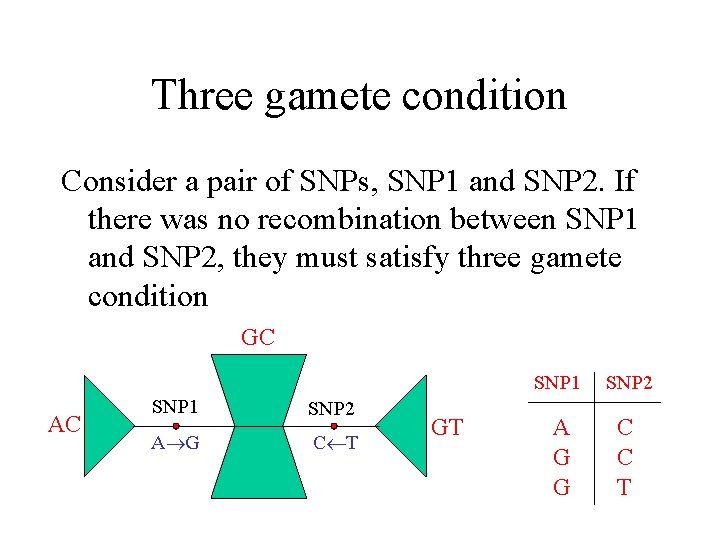
Three gamete condition Consider a pair of SNPs, SNP 1 and SNP 2. If there was no recombination between SNP 1 and SNP 2, they must satisfy three gamete condition GC AC SNP 1 SNP 2 A G C T GT SNP 1 SNP 2 A G G C C T

Four gamete test (Hudson and Kaplan, 1985) If we see all four gametes at SNP 1 and SNP 2 SNP 1 SNP 2 A G G A C C T T 4 GT Then there must have been a recombination event between these sites in their past history

Array of pairwise 4 GT test results Hudson and Kaplan, 1985 0, if there are less then 4 gametes D, dij= 1, if there are 4 gametes What is the minimal number of recombinations that could explain observed data ? Statistics FR (Hudson and Kaplan, 1985)
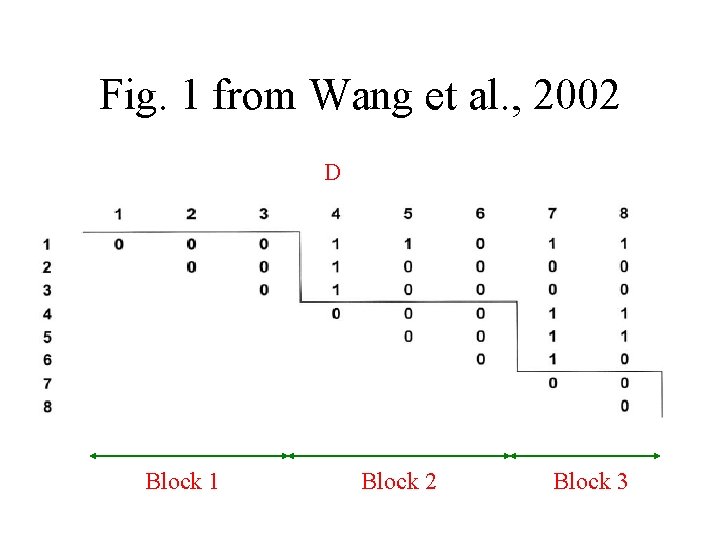
Fig. 1 from Wang et al. , 2002 D Block 1 Block 2 Block 3

Wang et al. , 2002 - Study • R. Hudson’s program for simulating genealogies with mutation, drift and recombination under various demographic scenarios • Study of dependence of average lengths of blocks on different factors • Comparison of simulation results to data from Patil et al. , 2002
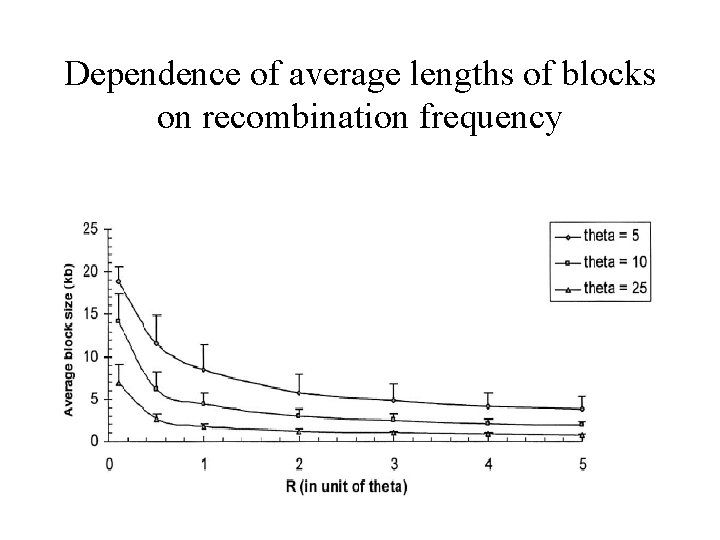
Dependence of average lengths of blocks on recombination frequency
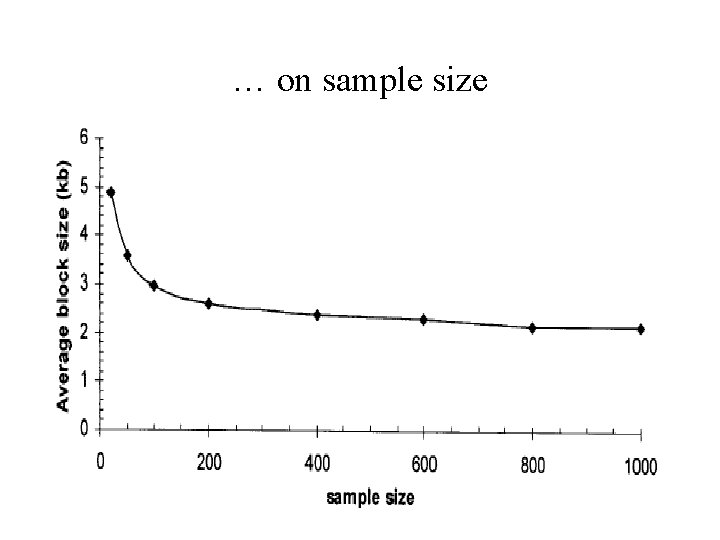
… on sample size
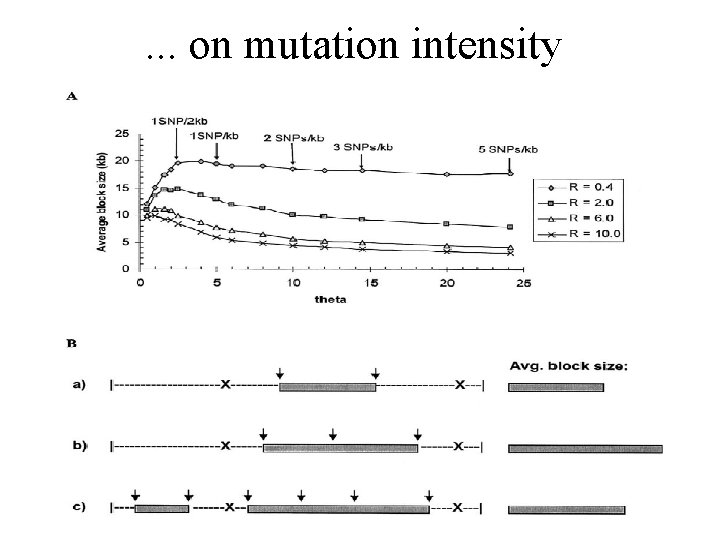
. . . on mutation intensity

Comparison to data from Patil et al. 2001 • Compute distribution of haplotype block lengths in the data from Patil et al. 2001 • Try to tune parameters and R to obtain similar distribution in the simulations
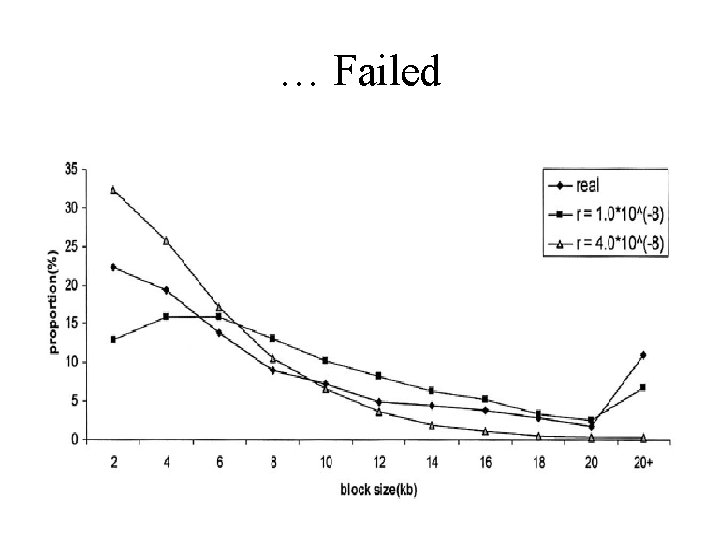
… Failed
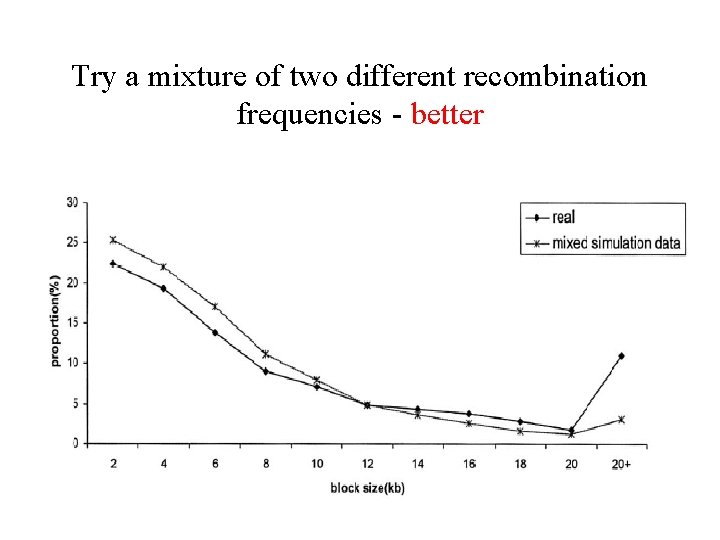
Try a mixture of two different recombination frequencies - better

Method 2 (Patil, 2001) Chromosome coverage – minimum number of SNPs to account for majority of haplotypes
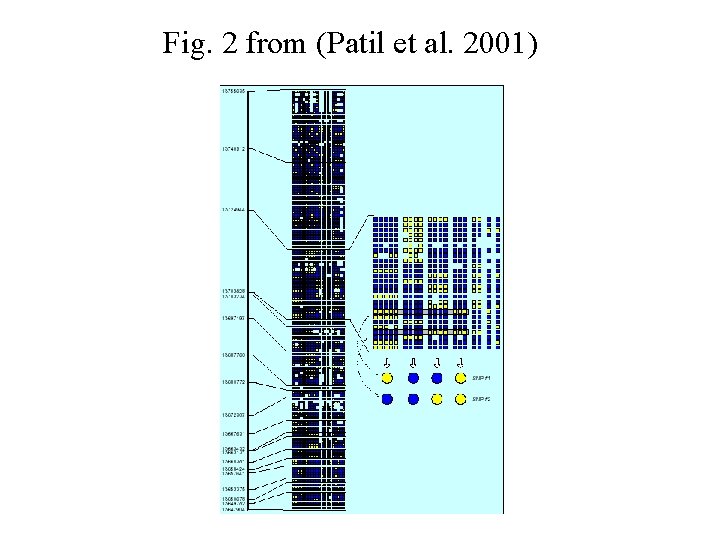
Fig. 2 from (Patil et al. 2001)

Problem formulation Define block boundaries to minimize the number of SNPs that distinguish at least percent of the haplotypes in each block

Common haplotypes Those represented more than one in the block

Condition Common haplotypes must constitute at least =80 percent of all haplotypes in the block Blocks that do not satisfy this are not allowed
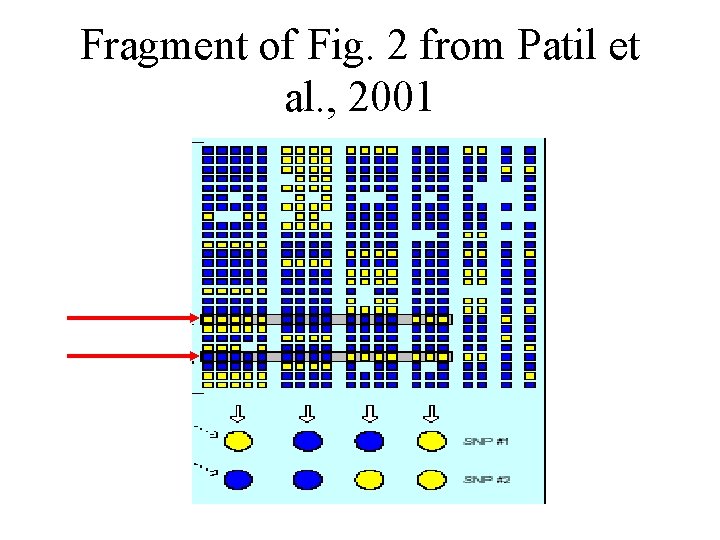
Fragment of Fig. 2 from Patil et al. , 2001

Notation • B – block defined as numbers of SNPs, e. g. , B = 45, 46, …. 50, or B = i, i+1, …, j • L(B) length of the block (number of SNPs) • f(B) – minimum number of SNP’s required to distinguish common haplotypes

Greedy Solution 010000000001000000010000111000001000000010010000000000010000110100001010 0000000010000000000100100000000101100100100101001000001001011000011010101010 00000100010110001010000101000110000010100000010000010011000001110100100000011000100011010 000000010001010000101000110000010100000010000010011000001110100100000011000100011010 000000001000010000010010000000000100100101001000001001011000011001000000010000100100000010000011000001010000100100110100000000100100000100111010000000010000100110100100000000001001001010010000010010110000110010000000000100000101000000000100001001000001001001000000001000011010101010 0000001000000000000001100000000010010000000000000010000110100001010 000000001000010000010010000000000100100110100010010000010010110000110010000000000100000101000000000100000000100100000000001000011010101010 00001000000100000000010100000000000001001010000001001000000000010000110100001010 10000000000100000101000000000100001001000001001001000000001000011010101010 00000001001000000011000011010000101000000101001000100100000101000010010000010011101000000000100000101000000000100001001000001001001000000001000011010101010 000001000001001000000100000110000010100001001000100001000000100111010100001 00000100000100100000010000011010000101000000101001000100100000010010011101000000 0001001000000100000001010000110011111100000000000000100111010100000010101001000001000001011110 000010000001000000000101000000000000010010100000010010000000000100001101000010100000000000001000001001110100000001000000001001010000001001001000001011010 Start End 0. Fix Start =End 1. Increment end 2. Compute ratio L(B)/f(B) 4. Go to 0 3. Stop at max …….

Results • 4563 representative SNPs (13%) • 4135 blocks

Method 3 (Zhang et al. 2002) Solves the same problem of 80% chromosome coverage, but using the better method of dynamic programming

Dynamic programming solution B 1(i) B 2(i) B 3(i) i 010000000001000000010000111000001000000010010000000000010000110100001010 0000000010000000000100100000000101100100100101001000001001011000011010101010 00000100010110001010000101000110000010100000010000010011000001110100100000011000100011010 000000010001010000101000110000010100000010000010011000001110100100000011000100011010 000000001000010000010010000000000100100101001000001001011000011001000000010000100100000010000011000001010000100100110100000000100100000100111010000000010000100110100100000000001001001010010000010010110000110010000000000100000101000000000100001001000001001001000000001000011010101010 0000001000000000000001100000000010010000000000000010000110100001010 000000001000010000010010000000000100100110100010010000010010110000110010000000000100000101000000000100000000100100000000001000011010101010 00001000000100000000010100000000000001001010000001001000000000010000110100001010 10000000000100000101000000000100001001000001001001000000001000011010101010 00000001001000000011000011010000101000000101001000100100000101000010010000010011101000000000100000101000000000100001001000001001001000000001000011010101010 000001000001001000000100000110000010100001001000100001000000100111010100001 00000100000100100000010000011010000101000000101001000100100000010010011101000000 0001001000000100000001010000110011111100000000000000100111010100000010101001000001000001011110 000010000001000000000101000000000000010010100000010010000000000100001101000010100000000000001000001001110100000001000000001001010000001001001000001011010 …… Optimal partition of SNPs 1, 2, … i Assume that for all i=1, 2, …, j-1 we know optimal block partition, B 1(i), B 2(i), …, Bk(i) that minimizes:

Bellman’s equation

Results • 3582 representative SNPs (compared to 4563 from greedy algorithm) • 2575 blocks (compared to 4135 blocks from greedy algorithm)

Conclusions • Studying haplotype block partitions is very important to 1. Constructing haplotype maps for genetic traits 2. Understanding recombination in human genome

To expect • A lot of papers in this area appearing in scientific journals
- Slides: 38