GWA promising but challenging Jianfeng Xu M D

GWA ─ promising but challenging Jianfeng Xu, M. D. , Dr. PH Professor of Public Health and Cancer Biology Director, Program for Genetic and Molecular Epidemiology of Cancer Associate Director, Center for Human Genomics Wake Forest University School of Medicine

Outline n The need for genome-wide association studies n The reality of genome-wide association studies n Important issues in genome-wide association studies n Genome coverage n Strategies for pre-association analysis n Strategies for association analysis n Sample size and false positives (Type I and II errors) n Confirmation in independent study populations n Increase the magnitude of effects of a specific gene

The need for GWA n Current understanding of disease etiology is limited n Therefore, candidate genes or pathways are insufficient n Current understanding of functional variants is limited n Therefore, the focusing on nonsynonymous changes is not sufficient n Results from linkage studies are often inconsistent and broad n Therefore, the utility of identified linkage regions is limited n GWA studies offer an effective and objective approach n Better chance to identify disease associated variants n Improve understanding of disease etiology n Improve ability to test gene-gene interaction and predict disease risk

GWA is promising n Many diseases and traits are influenced by genetic factors n i. e. , they are caused by sequence variants in the genome n Over 6 millions SNPs are known in the genome n i. e. , some SNPs will be directly or indirectly associated with causal variants n The cost of SNP Genotyping is reduced n i. e. , it is affordable to genotype a large number of SNPs in the genome n Large numbers of cases and controls are available n i. e. , there is statistical power to detect variants with modest effect n When the above conditions are met… n …associated SNPs will have different frequencies between cases and controls

GWA is challenging n Many diseases and traits are influenced by genetic factors n But probably due to multiple modest risk variants n They confer a stronger risk when they interact n True associated SNPs are not necessary highly significant n Too many SNPs are evaluated n False positives due to multiple tests n Single studies tend to be underpowered n False negatives n Considerable heterogeneity among studies n Phenotypic and genetic heterogeneity n False positives due to population stratification

Reality of GWA AMD, IBD, T 1 D, etc. Parkinson’s, nicotine dependence, T 2 D, etc. Prostate cancer, breast cancer, and other ongoing studies Heart diseases, lung diseases, psychiatric diseases, inflammatory diseases, cancers, and many other studies that are in planning stages

Important issues in genome-wide association studies n Genome coverage n Strategies for pre-association analysis n Strategies for association analysis n Sample size and false positives (Type I and II errors) n Confirmation in independent study populations n Increase the magnitude of effects of a specific gene

Genome coverage n Two major platforms for GWA n Illumina: Human. Hap 300, Human. Hap 550, and Human. Hap 1 M n Affymetrix: Gene. Chip 100 K, 500 K, and 1 M n Genome-wide coverage n The percentage of known SNPs in the genome that are in LD with the genotyped SNPs n Calculated based on Hap. Map n Calculated based on ENCODE

Genome coverage n Genome-wide coverage n Genome coverage of common SNPs (MAF ≥ 0. 05) n Genome coverage of rare SNPs n Genome coverage using multi-markers Pe’er, 2006
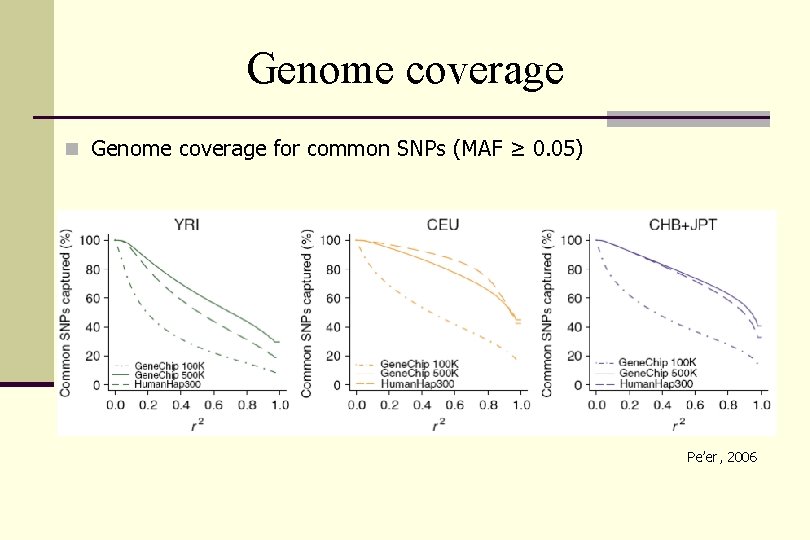
Genome coverage n Genome coverage for common SNPs (MAF ≥ 0. 05) Pe’er, 2006
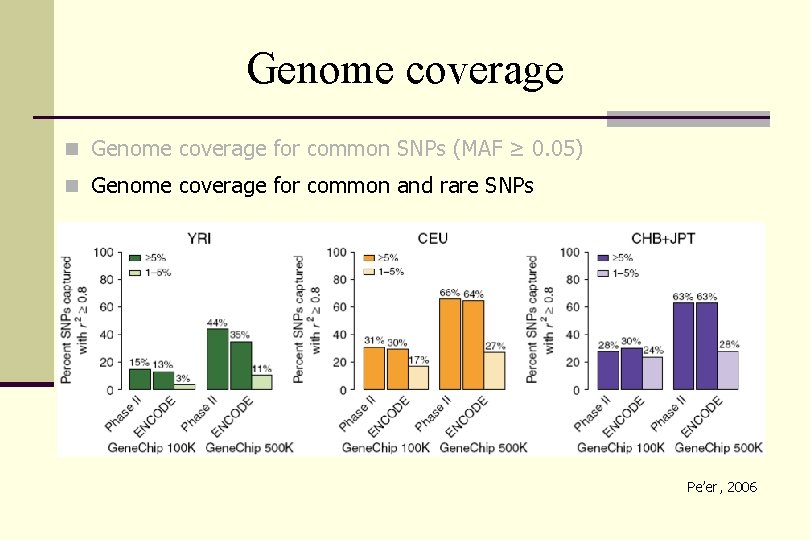
Genome coverage n Genome coverage for common SNPs (MAF ≥ 0. 05) n Genome coverage for common and rare SNPs Pe’er, 2006
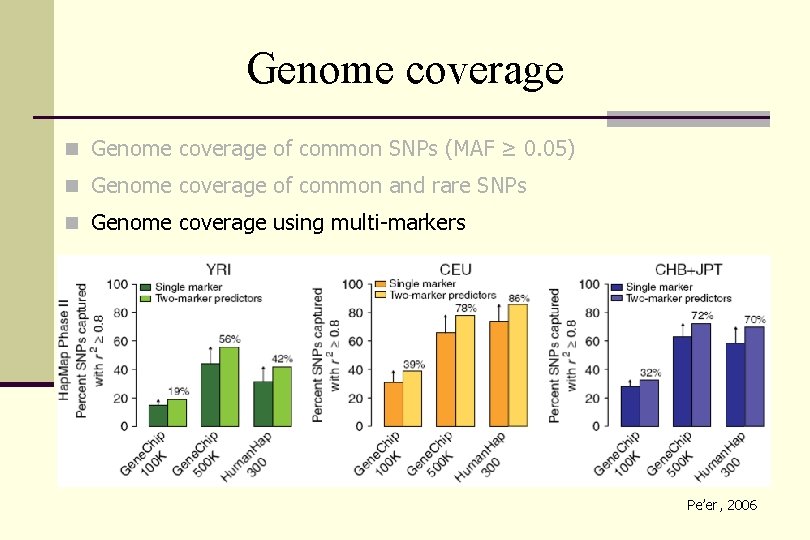
Genome coverage n Genome coverage of common SNPs (MAF ≥ 0. 05) n Genome coverage of common and rare SNPs n Genome coverage using multi-markers Pe’er, 2006

Important issues in genome-wide association studies n Genome coverage n Strategies for pre-association analysis n Strategies for association analysis n Sample size and false positives (Type I and II errors) n Confirmation in independent study populations n Increase the magnitude of effects of a specific gene

Strategies for pre-association analysis n Quality control n Filter SNPs by genotype call rates n Filter SNPs by minor allele frequencies n Filter SNPs by testing for Hardy-Weinberg Equilibrium

Strategies for pre-association analysis n Quality control n Quantile-quantile plot (Q-Q plot) n Evaluate whethere is an upward bias in association tests
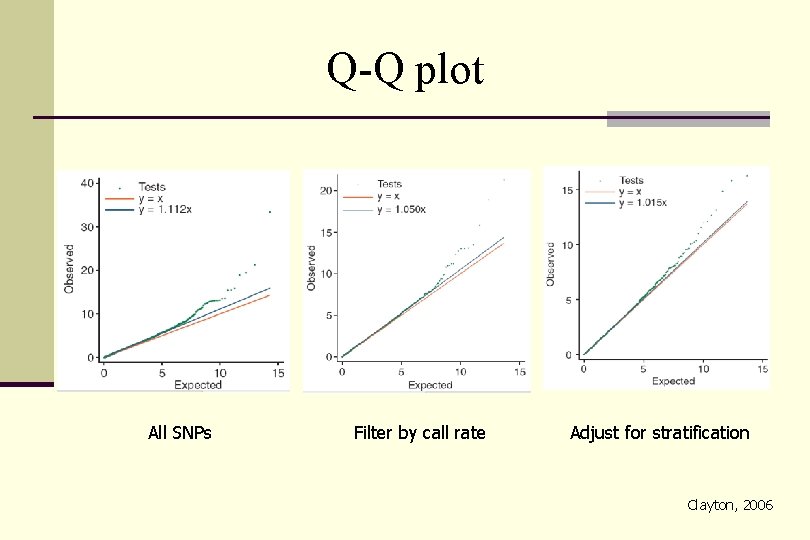
Q-Q plot All SNPs Filter by call rate Adjust for stratification Clayton, 2006

Strategies for pre-association analysis n Quality control n Quantile-quantile plot (Q-Q plot) n Population stratification n Genomic control n n Correct for stratification by adjusting association statistics at each SNP by a uniform overall inflation factor Is susceptible to over or under adjustment

Strategies for pre-association analysis n Quality control n Quantile-quantile plot (Q-Q plot) n Population stratification n Genomic control n Structure (STRUCTURE) n Used to assign the samples to discrete subpopulation clusters and then aggregate evidence of association within each cluster n Estimate individual proportion of ancestry and treat it as a covariate n Computationally intensive when there a large number of AIMs

Strategies for pre-association analysis n Quality control n Quantile-quantile plot (Q-Q plot) n Population stratification n Genomic control n Structure (STRUCTURE) n Principal component analysis (EIGENSTRAT) n Identify several eigenvectors (ancestries or geographic regions) n Adjust genotypes and phenotypes along each eigenvector n Compute association statistics using adjusted genotypes and phenotypes n No need for AIMs

Important issues in genome-wide association studies n Genome coverage n Strategies for pre-association analysis n Strategies for association analysis n Sample size and false positives (Type I and II errors) n Confirmation in independent study populations n Increase the magnitude of effects of a specific gene

Strategies for association analysis n Single SNP analysis using pre-specified genetic models n 2 x 3 table (2 -df) n Additive model (1 -df), and test for additivity n All possible genetic models

Strategies for association analysis n Single SNP analysis using pre-specified genetic models n Haplotype analysis n Two-marker and three-marker slide n Multi-marker n Within haplotype block n Between two recombination hot spots

Strategies for association analysis n Single SNP analysis using pre-specified genetic models n Haplotype analysis n Gene-gene and gene-environment interactions n Interaction with main effect n n Logistic regression Interaction without main effect: data mining n Classification and recursive tree (CART) n Multifactor Dimensionality Reduction (MDR)

Important issues in genome-wide association studies n Genome coverage n Strategies for pre-association analysis n Strategies for association analysis n Sample size and false positives (Type I and II errors) n Confirmation in independent study populations n Increase the magnitude of effects of a specific gene

Sample size and false positives n Estimate sample size n Sample size n OR n MAF n Type I error n Power n Quanto n Effective sample size

Sample size and false positives n Estimate sample size n False positives: too many dependent tests n Adjust for number of tests n Bonferroni correction § Nominal significance level = study-wide significance / number of tests § Nominal significance level = 0. 05/500, 000 = 10 -7 n Effective number of tests § Take LD into account n Permutation procedure § Permute case-control status § Mimic the actual analyses § Obtain empirical distribution of maximum test statistic under null hypothesis

Sample size and false positives n Estimate sample size n False positives: too many dependent tests n Adjust for number of tests n False discovery rate (FDR) n Expected proportion of false discoveries among all discoveries n Offers more power than Bonferroni n Holds under weak dependence of the tests

Sample size and false positives n Estimate sample size n False positives: too many dependent tests n Adjust for number of tests n False discovery rate (FDR) n Bayesian approach n Taking a priori into account, False-Positive Report Probability (FPRP)

Important issues in genome-wide association studies n Genome coverage n Strategies for pre-association analysis n Strategies for association analysis n Sample size and false positives (Type I and II errors) n Confirmation in independent study populations n Increase the magnitude of effects of a specific gene

Confirmation in independent study populations n The above approaches may limit the number of false positives n Confirmation is needed to dissect true from false positives n Replication, examine the results from the 2 nd stage only n Joint analysis, combining data from 1 st stage with 2 nd stage n Multiple stages
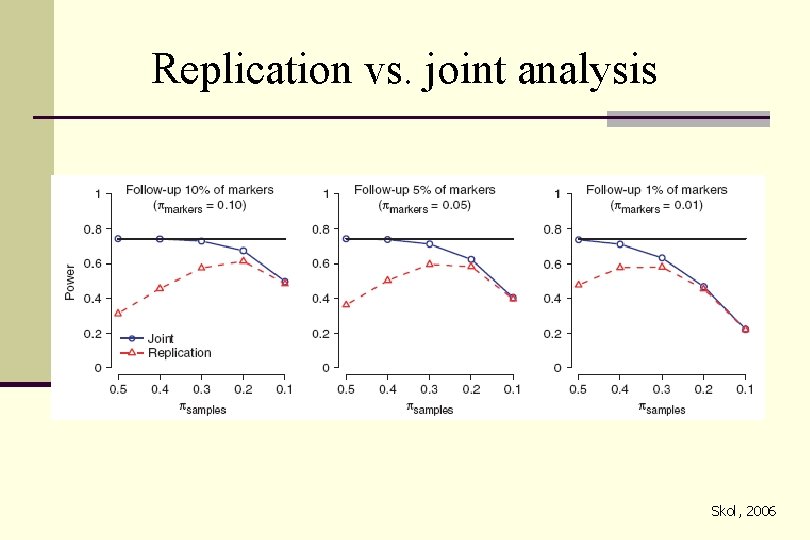
Replication vs. joint analysis Skol, 2006
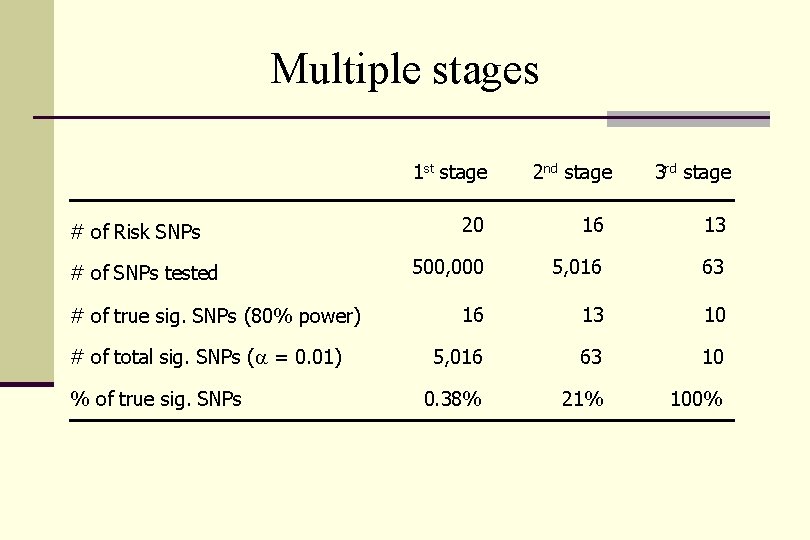
Multiple stages 1 st stage # of Risk SNPs # of SNPs tested # of true sig. SNPs (80% power) # of total sig. SNPs (a = 0. 01) % of true sig. SNPs 2 nd stage 3 rd stage 20 16 13 500, 000 5, 016 63 16 13 10 5, 016 63 10 0. 38% 21% 100%

Important issues in genome-wide association studies n Genome coverage n Strategies for pre-association analysis n Strategies for association analysis n Sample size and false positives (Type I and II errors) n Confirmation in independent study populations n Increase the magnitude of effects of a specific gene

Increase the magnitude of effects of a specific gene n Increase their effects by focusing on a subset of study subjects n Cases with a uniform phenotype, e. g. aggressive or early onset

Study aggressive cases

Increase the magnitude of effects of a specific gene n Increase their effects by focusing on a subset of study subjects n Cases with a uniform phenotypes, e. g. aggressive or early onset n Cases with family history

Study cases with family history Antoniou and Easton, 2003

Increase the magnitude of effects of a specific gene n Increase their effects by focusing on a subset of study subjects n Cases with a uniform phenotypes, e. g. aggressive or early onset n Cases with family history n Controls that are disease free

Disease free controls

Increase the magnitude of effects of a specific gene n Increase their effects by focusing on a subset of study subjects n Cases with a uniform phenotypes, e. g. aggressive or early onset n Cases with family history n Controls that are disease free n Increase their effects by studying a homogeneous population n Lower levels of genetic heterogeneity

Summary n GWA studies are promising but difficult n There are many important issues in GWA n The impact of these issues can be minimized by a well- designed study
- Slides: 41