Group 1 Lab 2 exercises and Assignment 2

Group 1 Lab 2 exercises and Assignment 2 Peter Fox Data Analytics – ITWS-4600/ITWS-6600/MATP-4450 Group 1, Lab 2, February 1, 2018 1

Labs • 2 a. Regression – New multivariate dataset • 2 b. k. NN – New Abalone dataset • 2 c. Kmeans – (Sort of) New Iris dataset • Do all three for Assignment 2 • And then general exercises 2

The Dataset(s) • http: //aquarius. tw. rpi. edu/html/DA • See slides: Data. Analytics 2018_Assignment_2. pptx on LMS or https: //tw. rpi. edu/web/Courses/Data. Analytics/ 2018 (under week 3) • Code fragments, i. e. they will not run as-is, on the following slides as group 1/lab 2_knn 1. R, etc. 3

Remember a few useful cmds head(<object>) tail(<object>) summary(<object>) 4

Regression Exercises • Using the EPI (under /EPI on web) dataset find the single most important factor in increasing the EPI in a given region • Examine distributions down to the leaf nodes and build up an EPI “model” 5
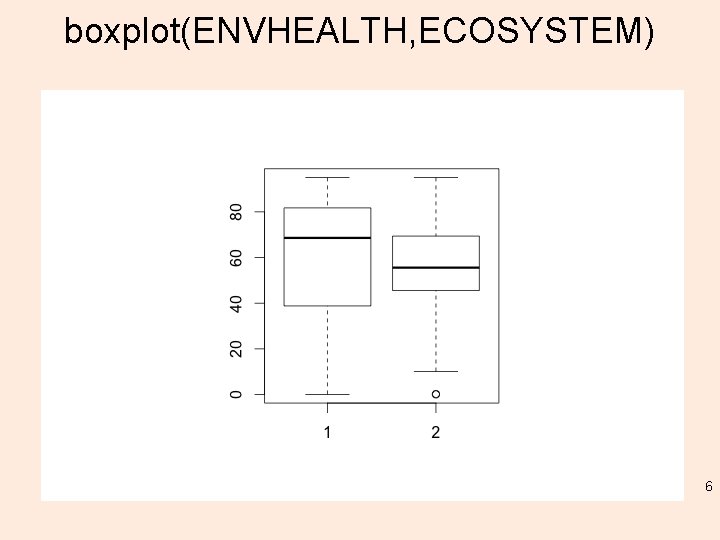
boxplot(ENVHEALTH, ECOSYSTEM) 6
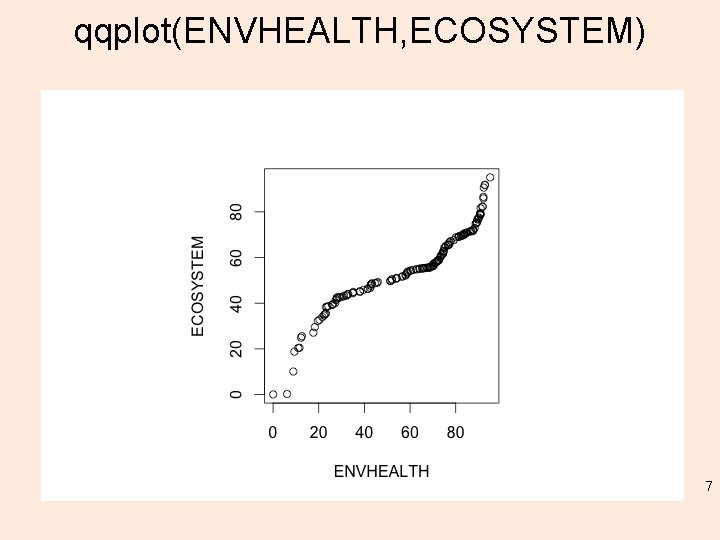
qqplot(ENVHEALTH, ECOSYSTEM) 7

ENVHEALTH/ ECOSYSTEM > shapiro. test(ENVHEALTH) Shapiro-Wilk normality test data: ENVHEALTH W = 0. 9161, p-value = 1. 083 e-08 ------- Reject. > shapiro. test(ECOSYSTEM) Shapiro-Wilk normality test data: ECOSYSTEM W = 0. 9813, p-value = 0. 02654 ----- ~reject 8

Kolmogorov- Smirnov - KS test > ks. test(ENVHEALTH, ECOSYSTEM) Two-sample Kolmogorov-Smirnov test data: ENVHEALTH and ECOSYSTEM D = 0. 2965, p-value = 5. 413 e-07 alternative hypothesis: two-sided Warning message: In ks. test(ENVHEALTH, ECOSYSTEM) : p-value will be approximate in the presence of ties 9

Linear and least-squares > EPI_data <- read. csv(”EPI_data. csv") > attach(EPI_data); > boxplot(ENVHEALTH, DALY, AIR_H, WATER_H) > lm. ENVH<lm(ENVHEALTH~DALY+AIR_H+WATER_H) > lm. ENVH … (what should you get? ) > summary(lm. ENVH) … > c. ENVH<-coef(lm. ENVH) 10

Predict > DALYNEW<-c(seq(5, 95, 5)) > AIR_HNEW<-c(seq(5, 95, 5)) > WATER_HNEW<-c(seq(5, 95, 5)) > NEW<data. frame(DALYNEW, AIR_HNEW, WATER_H NEW) > p. ENV<predict(lm. ENVH, NEW, interval=“prediction”) > c. ENV<predict(lm. ENVH, NEW, interval=“confidence”) 11

Repeat for AIR_E CLIMATE 12

Classification Exercises (group 1/lab 2_knn 1. R) > nyt 1<-read. csv(“nyt 1. csv") > nyt 1<-nyt 1[which(nyt 1$Impressions>0 & nyt 1$Clicks>0 & nyt 1$Age>0), ] > nnyt 1<-dim(nyt 1)[1] # shrink it down! > sampling. rate=0. 9 > num. test. set. labels=nnyt 1*(1. -sampling. rate) > training <-sample(1: nnyt 1, sampling. rate*nnyt 1, replace=FALSE) > train<-subset(nyt 1[training, ], select=c(Age, Impressions)) > testing<-setdiff(1: nnyt 1, training) > test<-subset(nyt 1[testing, ], select=c(Age, Impressions)) > cg<-nyt 1$Gender[training] > true. labels<-nyt 1$Gender[testing] > classif<-knn(train, test, cg, k=5) # > classif > attributes(. Last. value) # interpretation to come! 13

Classification Exercises (group 1/lab 2_knn 2. R) 2 examples in the script 14

Clustering Exercises • group 1/lab 2_kmeans 1. R • group 1/lab 2_kmeans 2. R – plotting up results from the iris clustering 15
- Slides: 15