Grids and Biology Professor Carole Goble University of

Grids and Biology Professor Carole Goble University of Manchester, UK BBSRC Bioinformatics and e. Science Grant Holders Workshop, Warwick, UK 28 th October 2002

Grids and Biology A take on the Grid Issues in Bioinformatics for Grid Various Bio. Grids Applicability of Grid to Biology Reality check
![What is the Grid? “ Grid computing [is] distinguished from conventional distributed computing by What is the Grid? “ Grid computing [is] distinguished from conventional distributed computing by](http://slidetodoc.com/presentation_image_h/bcbc97690df9773cf8ab14a692883d2b/image-3.jpg)
What is the Grid? “ Grid computing [is] distinguished from conventional distributed computing by its focus on large-scale resource sharing, innovative applications, and, in some cases, high-performance orientation. . . we review the "Grid problem", which we define as flexible, secure, coordinated resource sharing among dynamic collections of individuals, institutions, and resources - what we refer to as virtual organizations. " From "The Anatomy of the Grid: Enabling Scalable Virtual Organizations" by Foster, Kesselman and Tuecke

What is the Grid? Resource sharing & coordinated problem solving in dynamic, multi-institutional virtual organizations On-demand, ubiquitous access to computing, data, and services New capabilities constructed dynamically and transparently from distributed services No central location, No central control, No existing trust relationships, Little predetermination Uniformity for Pooling Resources Virtual pools of resources: databases, clusters….
Biology as a Grid Application Informational Science Large Scale Distributed No one organisation owns it all
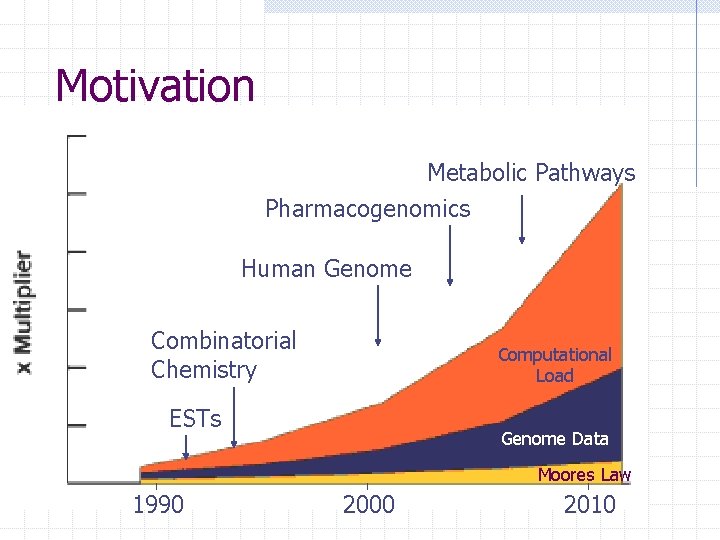
Motivation Metabolic Pathways Pharmacogenomics Human Genome Combinatorial Chemistry Computational Load ESTs Genome Data Moores Law 1990 2000 2010
![Bio. Medical Computation [Rick Stevens, Argonne Labs] Bio. Medical Computation [Rick Stevens, Argonne Labs]](http://slidetodoc.com/presentation_image_h/bcbc97690df9773cf8ab14a692883d2b/image-7.jpg)
Bio. Medical Computation [Rick Stevens, Argonne Labs]
![Biomedical Data: [Rick Stevens, Argonne Labs] High Complexity and Large Scale billions millions Hundred Biomedical Data: [Rick Stevens, Argonne Labs] High Complexity and Large Scale billions millions Hundred](http://slidetodoc.com/presentation_image_h/bcbc97690df9773cf8ab14a692883d2b/image-8.jpg)
Biomedical Data: [Rick Stevens, Argonne Labs] High Complexity and Large Scale billions millions Hundred thousands Protein-Protein Interactions metabolism pathways receptor-ligand 4º structure Proteins sequence 2º structure 3º structure MPMILGYWDIRGLAHAIRLLLEYTDSSYEEKKYT. . . billions DNA sequences alignments . . . atcgaattccaggcgtcacattctcaattcca. . . millions Physiology Cellular biology Biochemistry Neurobiology Polymorphism Endocrinology and Variants genetic variants etc. individual patients epidemiology Genetics and Maps Linkage Cytogenetic Clone-based millions ESTs Expression patterns Large-scale screens

Bio. Grid Projects EUROGRID Bio. GRID Asia Pacific Bio. GRID North Carolina Bio. Grid Bioinformatics Research Network Osaka University Bio. Grid Indiana University Bio. Archive Bio. Grid my. Grid Bio. Sim e-Protein Obi. Grid

Today’s Grid A Single System Image Transparent wide-area access to large data banks Transparent wide-area access to applications on heterogeneous platforms Transparent wide-area access to processing resources Security, certification, single sign-on authentication, AAA n Grid Security Infrastructure, Data access, Transfer & Replication n Grid. FTP, Giggle Computational resource discovery, allocation and process creation n GRAAM, Unicore, Condor-G

Immediate benefits Uniform file views of directories, regardless of platform Grid-based data transfer libraries for faster access to large files, reducing need for mirror-site servers. Replication to support mirroring Grid APIs provide a job manager with metadata about services to the user. Evaluate the quality of service providers based on factors that may include more than just server performance and availability. Grid-aware applications -- split sequence reference libraries among several servers, where BLAST comparisons can be conducted in parallel. Shielding from a variety of low-level computing problems would otherwise have to address themselves.
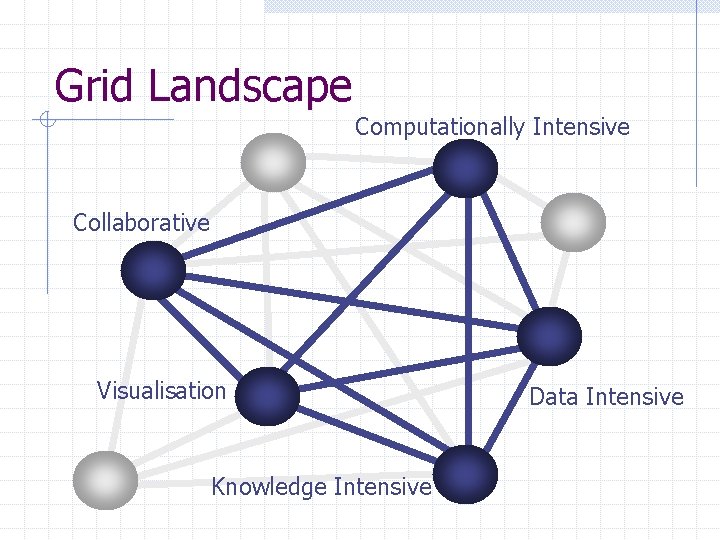
Grid Landscape Computationally Intensive Collaborative Visualisation Knowledge Intensive Data Intensive
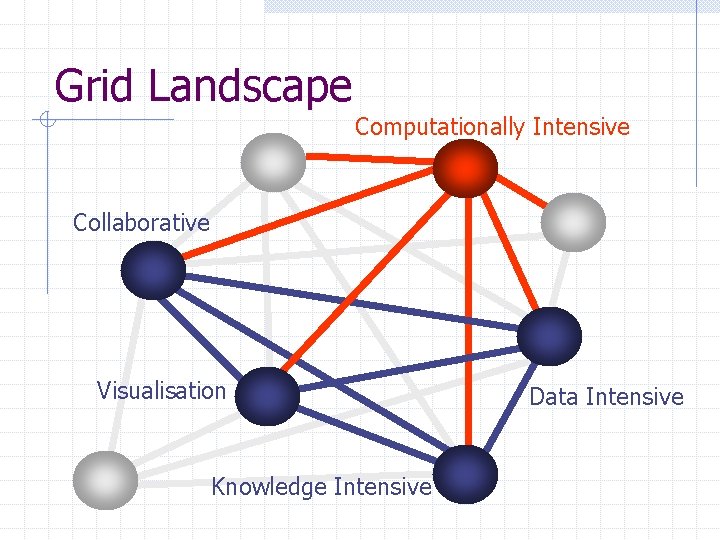
Grid Landscape Computationally Intensive Collaborative Visualisation Knowledge Intensive Data Intensive
Classical Grids emphasise sharing of physical resources. Existing Grid middleware (e. g. Globus, Condor, Unicore) allows resource discovery, resource allocation, data movement, certification …
![High Performance Bioinformatics Software [Jack da Silva, NCSC, Paracel] High Performance Bioinformatics Software [Jack da Silva, NCSC, Paracel]](http://slidetodoc.com/presentation_image_h/bcbc97690df9773cf8ab14a692883d2b/image-15.jpg)
High Performance Bioinformatics Software [Jack da Silva, NCSC, Paracel]
European Data. Grid

Managed access to specialist remote resources

Access portal for biomolecular modeling resources. Interfaces to enable chemists and biologists to be able to submit work to HPC facilities Visualization of electrostatic field generated by a molecule. dr Krzysztof Nowinski (ICM)
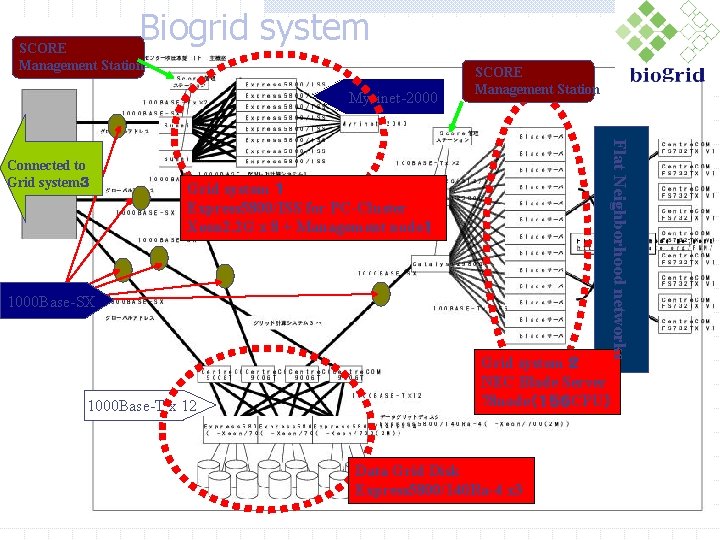
Biogrid system SCORE Management Station Myrinet-2000 Grid system 1 Express 5800/ISS for PC-Cluster Xeon 2. 2 G x 8 + Management node1 1000 Base-SX 1000 Base-T x 12 Flat Neighborhood networks Connected to Grid system3 SCORE Management Station Grid system 2 NEC Blade Server 78 node(156 CPU) Data Grid Disk Express 5800/140 Ra-4 x 3

Remote control of instruments Sharing of UHVEM(Ultra High Voltage Electron Microscopy) in Osaka University with NCMIR (National Center for Microscopy and Imaging Research) n 3 Million electron volts n the most powerful microscopy Osaka University Tokyo XP JGN UHVEM (Osaka, Japan) (Chicago) STAR TAP Trans. PAC APAN (UC San Diego) SDSC v. BNS NCMIR (San Diego)

Home Computers Evaluate AIDS Drugs Community = n n n 1000 s of home computer users Philanthropic computing vendor (Entropia) Research group (Scripps) Common goal= advance AIDS research From Steve Tuecke 12 Oct. 01

Matlab Geodise release in November 02 sjc@soton. ac. uk Matlab and toolboxes for mathematical computation, analysis, visualization, and algorithm development: MATLAB is an intuitive language and a technical computing environment. It provides core mathematics and advanced graphical tools for data analysis, visualization, and algorithm and application development. With more than 600 mathematical, statistical, and engineering functions, engineers and scientists rely on the MATLAB environment for their technical computing needs. ” (www. mathworks. com) CROSS PLATFORM/ OS
![Bio. Sim -- Molecular simulations as a tool for protein structure analysis [Sansom] synchrotron Bio. Sim -- Molecular simulations as a tool for protein structure analysis [Sansom] synchrotron](http://slidetodoc.com/presentation_image_h/bcbc97690df9773cf8ab14a692883d2b/image-23.jpg)
Bio. Sim -- Molecular simulations as a tool for protein structure analysis [Sansom] synchrotron compute GRID MD database novel biology… u Overall vision – simulation as an integral component of structural genomics u Needs both capacity (many systems) and capability (large systems - HPCx) u Molecular Dynamics database (distributed)
Grid Landscape Computationally Intensive Collaborative Visualisation Knowledge Intensive Data Intensive
![[Rick Stevens Argonne Labs] Visualization + Bioinformatics Visualization Environment Bioinformatic Analysis Tools Genome Visualization [Rick Stevens Argonne Labs] Visualization + Bioinformatics Visualization Environment Bioinformatic Analysis Tools Genome Visualization](http://slidetodoc.com/presentation_image_h/bcbc97690df9773cf8ab14a692883d2b/image-25.jpg)
[Rick Stevens Argonne Labs] Visualization + Bioinformatics Visualization Environment Bioinformatic Analysis Tools Genome Visualization Tools Function Assignment Whole Genome Analysis Metabolic Reconstruction Enzymatic Constants Metabolic *** Network Visualization Tools Microbiology & Biochemistry Stoichiometric Representation & Flux Analysis Interactive Stoichiometric Graphical Tools Whole Cell Visualizations Image/Spectra Augmentations Proteomics Dynamic Simulation Laboratory Verification
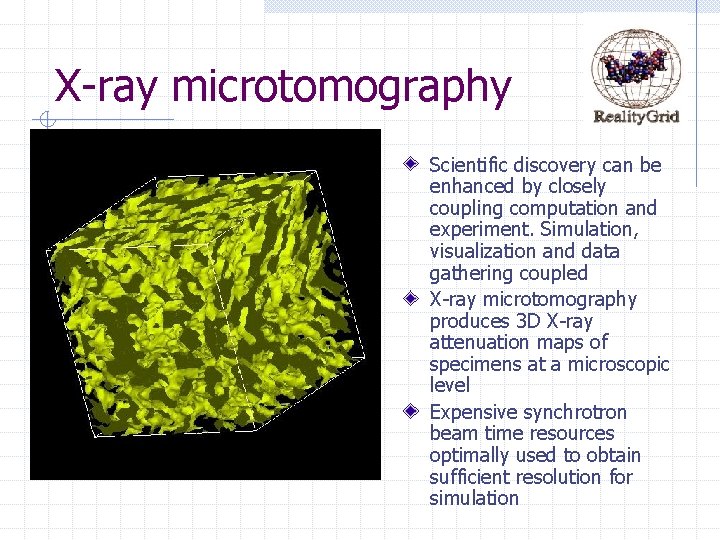
X-ray microtomography Scientific discovery can be enhanced by closely coupling computation and experiment. Simulation, visualization and data gathering coupled X-ray microtomography produces 3 D X-ray attenuation maps of specimens at a microscopic level Expensive synchrotron beam time resources optimally used to obtain sufficient resolution for simulation

Interactive Steering • User steers calculation from laptop • Controlled steering on supercomputers • Visualization and computation use large scale machines accessed via Grid. Enables controlled simulation using knowledge and skills of trained scientist.

Scalable molecular dynamics • Structure of a protein in a fluid medium • Calculation takes into account forces between protein and ambient medium (in this case water molecules) • Run on world largest academic computer, Le. Mieux at PSC (6 Tflops theoretical peak)
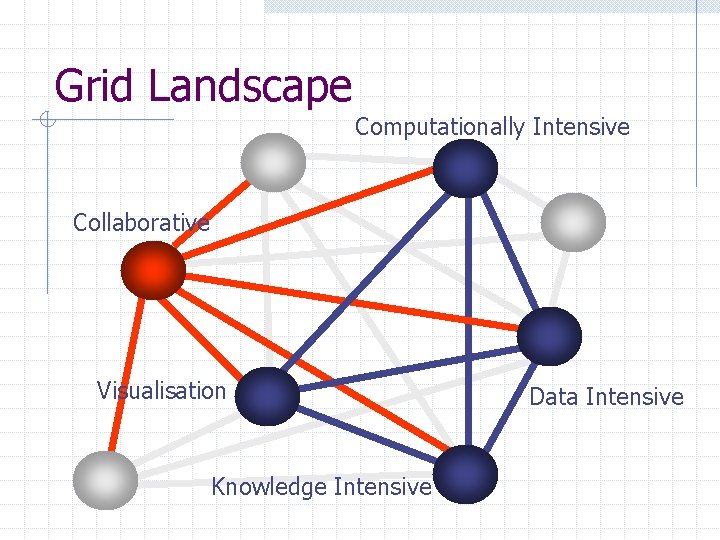
Grid Landscape Computationally Intensive Collaborative Visualisation Knowledge Intensive Data Intensive
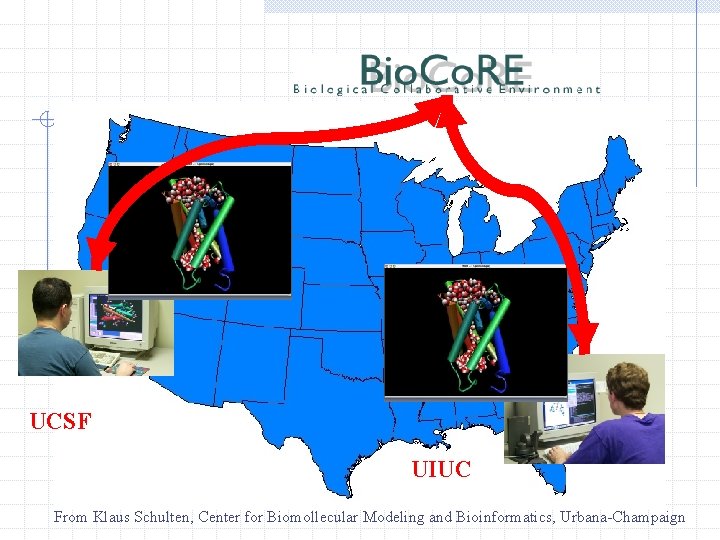
UCSF UIUC From Klaus Schulten, Center for Biomollecular Modeling and Bioinformatics, Urbana-Champaign

http: //www. ks. uiuc. edu/Research/biocore/
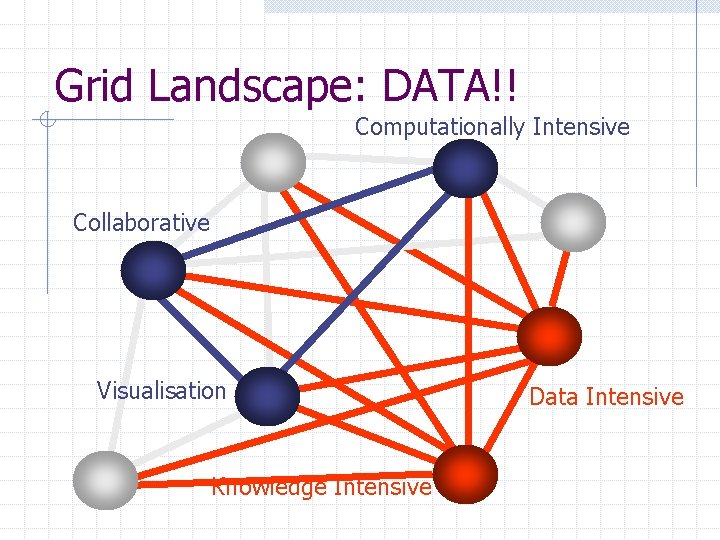
Grid Landscape: DATA!! Computationally Intensive Collaborative Visualisation Knowledge Intensive Data Intensive

Information Weaving and Question Answering Large amounts of different kinds of data & many applications. Highly heterogeneous. n Different types, algorithms, forms, implementations, communities, service providers High autonomy. Highly complex and interrelated, & volatile.
![[Mike Sternberg] proteome sequences SCOP CATH PDB NRPROT TM, CC, LC, SIG & MOTIFS [Mike Sternberg] proteome sequences SCOP CATH PDB NRPROT TM, CC, LC, SIG & MOTIFS](http://slidetodoc.com/presentation_image_h/bcbc97690df9773cf8ab14a692883d2b/image-34.jpg)
[Mike Sternberg] proteome sequences SCOP CATH PDB NRPROT TM, CC, LC, SIG & MOTIFS PSIBLAST & HHMs PDB hit 3 D modelling x 2 Annotation Pipeline INTERPRO no PDB hit fold recognition x 2 structure-based function prediction structural and functional annotation
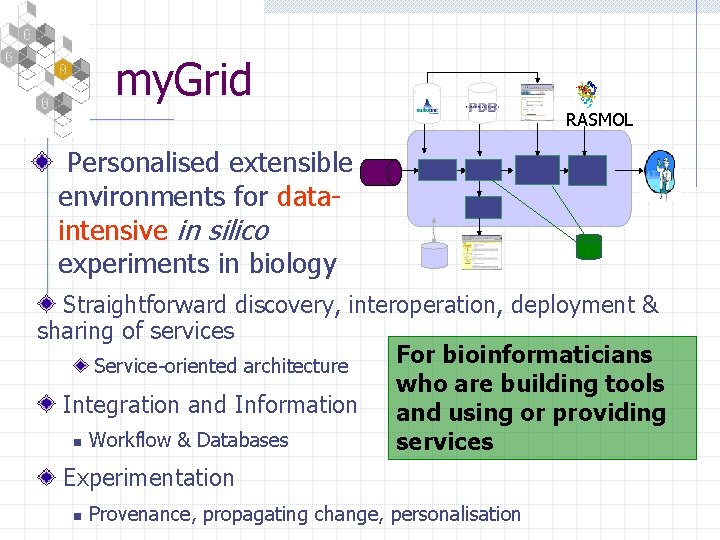
my. Grid RASMOL Personalised extensible environments for dataintensive in silico experiments in biology Straightforward discovery, interoperation, deployment & sharing of services For bioinformaticians Service-oriented architecture who are building tools Integration and Information and using or providing n Workflow & Databases services Experimentation n Provenance, propagating change, personalisation
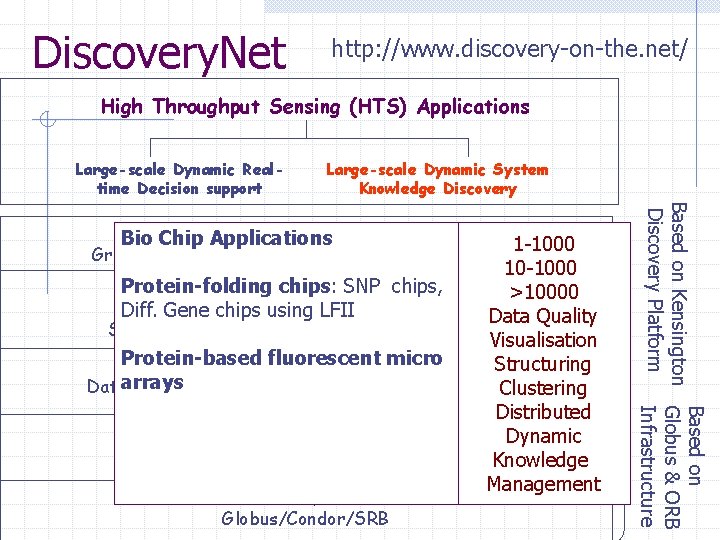
Discovery. Net http: //www. discovery-on-the. net/ High Throughput Sensing (HTS) Applications Large-scale Dynamic Realtime Decision support Large-scale Dynamic System Knowledge Discovery 1 -1000 10 -1000 Information Structuring Protein-folding chips: SNP chips, >10000 Information Integration Diff. Gene chips using LFII& Composition, Data Quality Semantics & Domain-based Ontologies, Sharing Visualisation Protein-based micro Distributedfluorescent Data Engineering Structuring Dataarrays Registration, Data Normalisation, Data Quality Clustering Distributed High Throughput Computing Services Dynamic Utilising Grid Infrastructure for HT Computing Knowledge Management Grid Basic Infrastructure Grid-based Data Mining, Collaborative Visualisation Globus/Condor/SRB Based on Kensington Globus & ORB Discovery Platform Infrastructure Knowledge Discovery Bio. Grid-based Chip Applications

Grid Evolution 1 st Generation Grid Computationally intensive, file access/transfer n Bag of various heterogeneous protocols & toolkits n Recognises internet, Ignores Web n Academic teams We are here! nd n 2 n n n Generation Grid Data intensive -> knowledge intensive Services-based architecture Recognises Web and Web services Global Grid Forum Industry participation
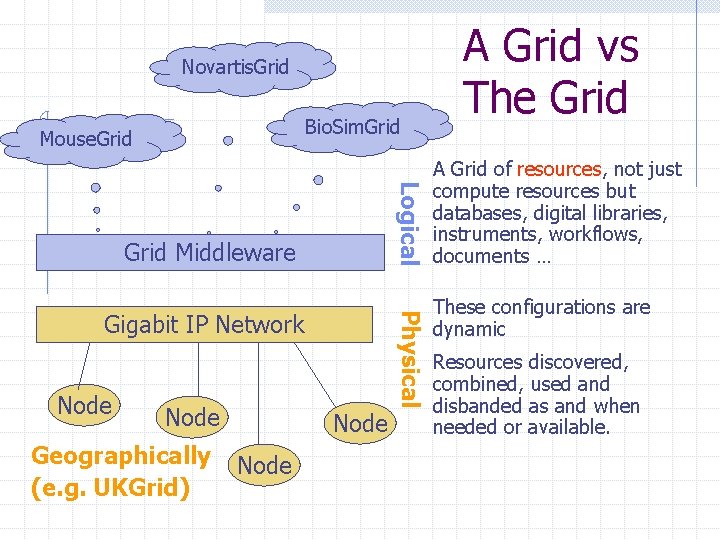
Novartis. Grid Bio. Sim. Grid Mouse. Grid Logical Grid Middleware Node Geographically (e. g. UKGrid) Physical Gigabit IP Network Node A Grid vs The Grid A Grid of resources, not just compute resources but databases, digital libraries, instruments, workflows, documents … These configurations are dynamic Resources discovered, combined, used and disbanded as and when needed or available.
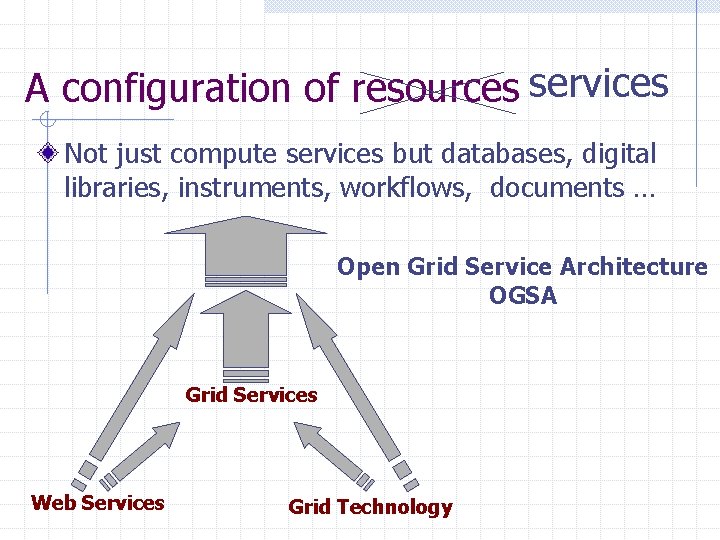
A configuration of resources services Not just compute services but databases, digital libraries, instruments, workflows, documents … Open Grid Service Architecture OGSA Grid Services Web Services Grid Technology

Bio Services Domain Oriented Services Basic Bio. Grid Services Grid Resource Services Common Services Base Services Fabric Services • • Drug Discovery Microbial Engineering Molecular Ecology Oncology Research • • Integrated Databases Sequence Analysis Protein Interactions Cell Simulation • • • Compute Services Pipeline Services Data Archive Service Database Hosting Workflow Enactment Event notification

What We Need to Create Grid Bio applications enablement software layer n n Provide application’s access to Grid services Provides OS independent services Grid enabled version of bioinformatics data management tools (e. g. DL, SRS, etc. ) n n Need to support virtual databases via Grid services Grid support for commercial databases Bioinformatics applications “plug-in” modules n n End user tools for a variety of domains Support major existing Bio IT platforms

Requirements for the Bio. Grid Open and extendable architecture n n Enable tie in to service stack at appropriate points Not just access via Portals Leverage scripting tools in wide use for Bioinformatics n Create Bio. Grid services bindings for PERL and Python Address data federation and integration n Leverage work of IBM, Lion Bio. Sciences, DAS, Bio. MOBY, etc. Match the biology workflow and tool chain n n Create high-level Bio. Grid services to address critical stages in existing workflow Support composibility of new Bio. Grid tools with existing tool chain elements

Some Bio. Grid Challenges Scalable human bioinformatics expertise n n Best people working on the important problems Exploit collaboration technology to create world class teams Robust local bioinformatics computing environment n n Best systems administrators and high-end technologies Embed local resources into the Grid via portal technologies Access to leading edge bioinformatics software and databases customized to user needs n n Core content from top scientists and developers Integrated access to biological databases Worldwide access to robust computing and database infrastructure n n Leverage Grid technology to provide worldwide access Integrate purpose built systems and service providers

Reality Checks!! The Technology is Ready n Not true — its emerging w Building middleware, Advancing Standards, Developing, Dependability w Building demonstrators. w The computational grid is in advance of the data intensive middleware w Integration and curation are probably the obstacles w But!! It doesn’t have to be all there to be useful. We know how we will use grid services n No — Disruptive technology w Lower the barriers of entry.

Reality Checks!! It’s the only game n Not true — I 3 C, Bio. MOBY, bio. DAS, OMG LSR w Grid and Web service merge makes integration likely. One Size Fits All n Not true w Addressed by a minimum set of composable virtual services, But starting with Globus It’s only for “big” science n No — “small” science collaborates too! Biology is not unique! n Astro. Grid

Not a silver bullet! Its just middleware not magic Data quality Content management of databases (controlled vocabularies) Provenance and versioning policies Appropriate use of tools Computational inaccessibility of free text annotation Database accessibility through means other than point and click web interfaces. Independent of the Grid!
Life Sciences Grid (LSG) http: //people. cs. uchicago. edu/~dangulo/LSG/
- Slides: 47