Graphs for workflow Genome Workflow Compile time IncludeExclude

- Slides: 8

Graphs for workflow

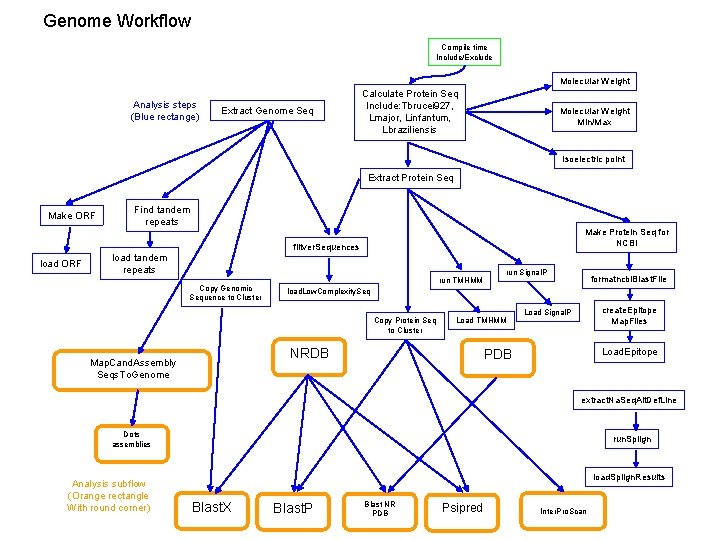
Genome Workflow Compile time Include/Exclude Molecular Weight Analysis steps (Blue rectange) Extract Genome Seq Calculate Protein Seq Include: Tbrucei 927, Lmajor, Linfantum, Lbraziliensis Molecular Weight Min/Max Isoelectric point Extract Protein Seq Make ORF Find tandem repeats Make Protein Seq for NCBI filtver. Sequences load ORF load tandem repeats Copy Genomic Sequence to Cluster run TMHMM formatncbi. Blast. File load. Low. Complexity. Seq Copy Protein Seq to Cluster Load TMHMM NRDB Map. Cand. Assembly Seqs. To. Genome run Signal. P create. Epitope Map. Files Load Signal. P Load. Epitope PDB extract. Na. Seq. Alt. Def. Line Dots assemblies Analysis subflow (Orange rectangle With round corner) run. Splign load. Splign. Results Blast. X Blast. P Blast NR PDB Psipred Inter. Pro. Scan

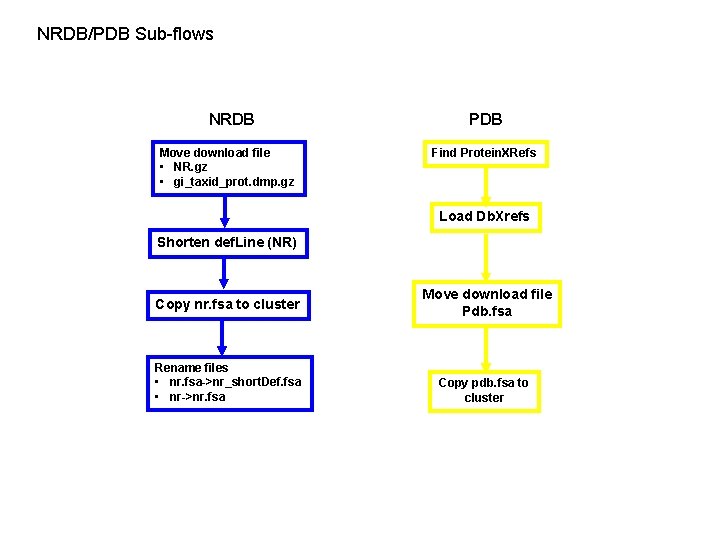
NRDB/PDB Sub-flows NRDB Move download file • NR. gz • gi_taxid_prot. dmp. gz PDB Find Protein. XRefs Load Db. Xrefs Shorten def. Line (NR) Copy nr. fsa to cluster Rename files • nr. fsa->nr_short. Def. fsa • nr->nr. fsa Move download file Pdb. fsa Copy pdb. fsa to cluster

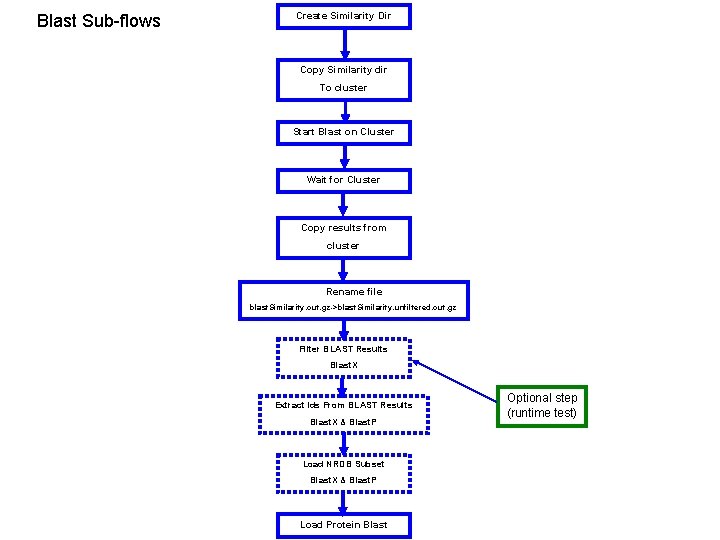
Blast Sub-flows Create Similarity Dir Copy Similarity dir To cluster Start Blast on Cluster Wait for Cluster Copy results from cluster Rename file blast. Similarity. out. gz->blast. Similarity. unfiltered. out. gz Filter BLAST Results Blast. X Extract Ids From BLAST Results Blast. X & Blast. P Load NRDB Subset Blast. X & Blast. P Load Protein Blast Optional step (runtime test)

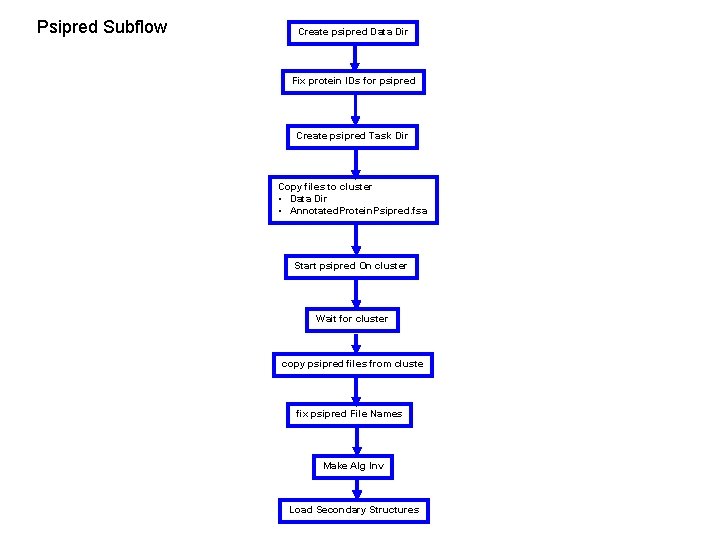
Psipred Subflow Create psipred Data Dir Fix protein IDs for psipred Create psipred Task Dir Copy files to cluster • Data Dir • Annotated. Protein. Psipred. fsa Start psipred On cluster Wait for cluster copy psipred files from cluste fix psipred File Names Make Alg Inv Load Secondary Structures

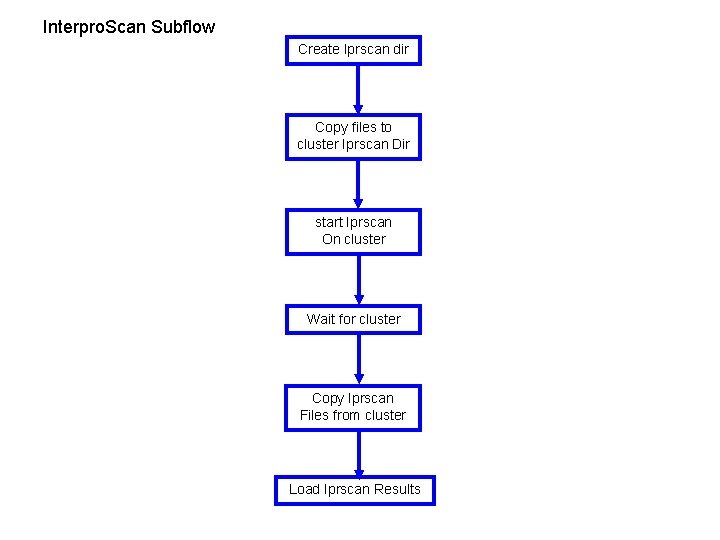
Interpro. Scan Subflow Create Iprscan dir Copy files to cluster Iprscan Dir start Iprscan On cluster Wait for cluster Copy Iprscan Files from cluster Load Iprscan Results

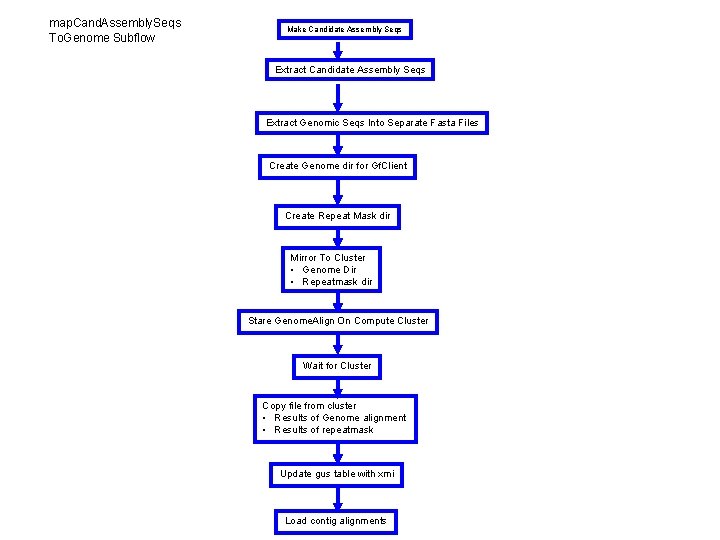
map. Cand. Assembly. Seqs To. Genome Subflow Make Candidate Assembly Seqs Extract Genomic Seqs Into Separate Fasta Files Create Genome dir for Gf. Client Create Repeat Mask dir Mirror To Cluster • Genome Dir • Repeatmask dir Stare Genome. Align On Compute Cluster Wait for Cluster Copy file from cluster • Results of Genome alignment • Results of repeatmask Update gus table with xmi Load contig alignments

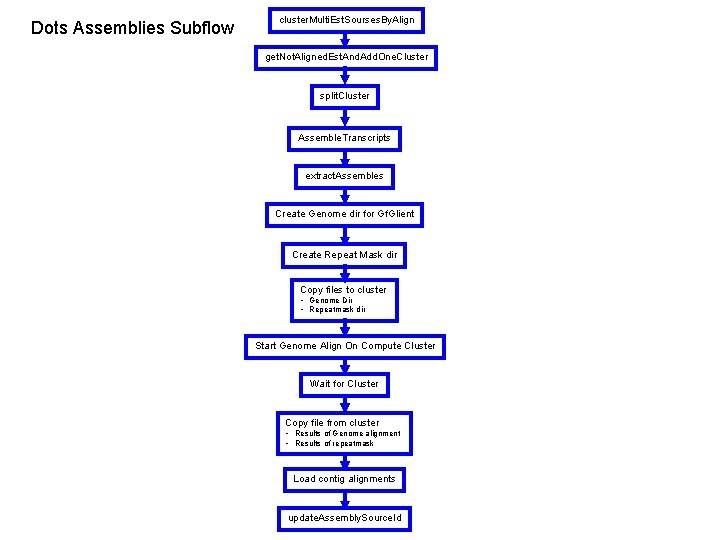
Dots Assemblies Subflow cluster. Multi. Est. Sourses. By. Align get. Not. Aligned. Est. And. Add. One. Cluster split. Cluster Assemble. Transcripts extract. Assembles Create Genome dir for Gf. Glient Create Repeat Mask dir Copy files to cluster • Genome Dir • Repeatmask dir Start Genome Align On Compute Cluster Wait for Cluster Copy file from cluster • Results of Genome alignment • Results of repeatmask Load contig alignments update. Assembly. Source. Id