Glucocorticoid receptor crosstalk with NFk B in airway

Glucocorticoid receptor crosstalk with NF-k. B in airway cells – analyzing the cistromes BIOS 6660 Genomic Data Analysis with R and Bioconductor Anthony Gerber MD, Ph. D. October 20, 1015

The Nuclear Receptor family • Transcription factors • Many are ligand activated • Only class of transcription factors that can be targeted by small molecules in the clinic • Clinical targets include estrogen, androgen, mineralocorticoid, Vitamins D, glucocorticoid and thyroid receptors, RXR, PPAR • Major interest in developing selective ligands/modulators to enable improved therapeutic windows
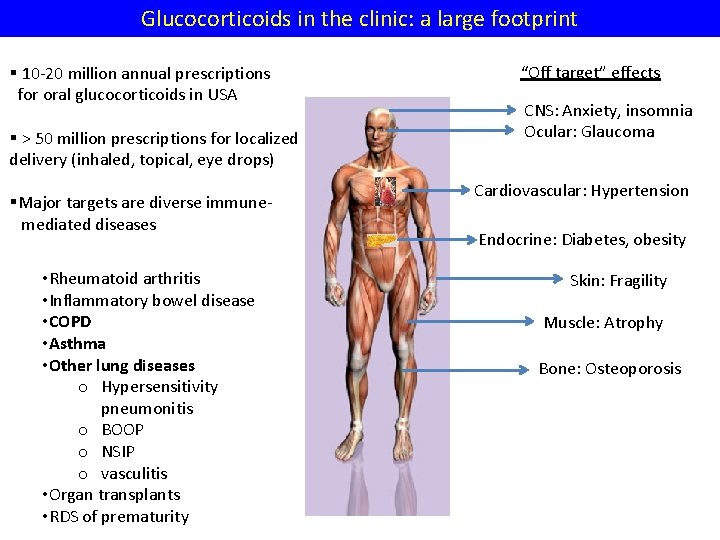
Glucocorticoids in the clinic: a large footprint § 10 -20 million annual prescriptions for oral glucocorticoids in USA § > 50 million prescriptions for localized delivery (inhaled, topical, eye drops) §Major targets are diverse immunemediated diseases • Rheumatoid arthritis • Inflammatory bowel disease • COPD • Asthma • Other lung diseases o Hypersensitivity pneumonitis o BOOP o NSIP o vasculitis • Organ transplants • RDS of prematurity “Off target” effects CNS: Anxiety, insomnia Ocular: Glaucoma Cardiovascular: Hypertension Endocrine: Diabetes, obesity Skin: Fragility Muscle: Atrophy Bone: Osteoporosis

Balancing disease symptoms with glucocorticoid side effects in the clinic: A 50 year old female with severe, persistent asthma No oral glucocorticoid use • lung function < 50% of normal • short of breath after walking 2 blocks • unable to go up a flight of stairs • frequent coughing episodes • 1 -2 ER visits per quarter • 2 -3 hospitalizations per year Taking oral glucocorticoids • lung function ~80% of normal • no shortness of breath after 10 blocks • able to go up 2 flights of stairs • no hospitalizations or ED visits • 20 pound weight gain • lower extremity edema • irritability • high blood sugars • increased risk of osteoporosis There is a major unmet need for improved glucocorticoid-based therapies
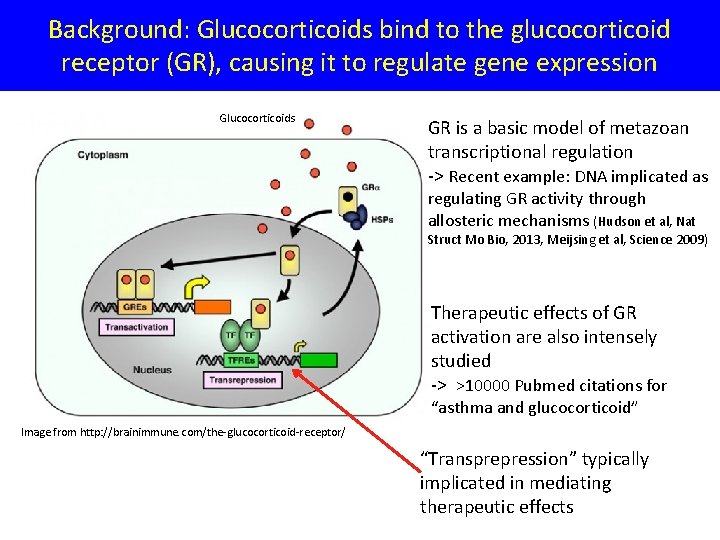
Background: Glucocorticoids bind to the glucocorticoid receptor (GR), causing it to regulate gene expression Glucocorticoids GR is a basic model of metazoan transcriptional regulation -> Recent example: DNA implicated as regulating GR activity through allosteric mechanisms (Hudson et al, Nat Struct Mo Bio, 2013, Meijsing et al, Science 2009) Therapeutic effects of GR activation are also intensely studied -> >10000 Pubmed citations for “asthma and glucocorticoid” Image from http: //brainimmune. com/the-glucocorticoid-receptor/ “Transprepression” typically implicated in mediating therapeutic effects
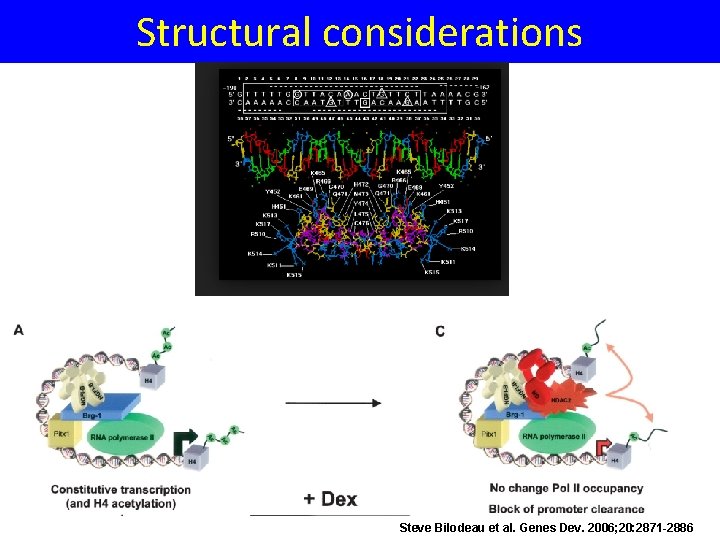
Structural considerations Steve Bilodeau et al. Genes Dev. 2006; 20: 2871 -2886
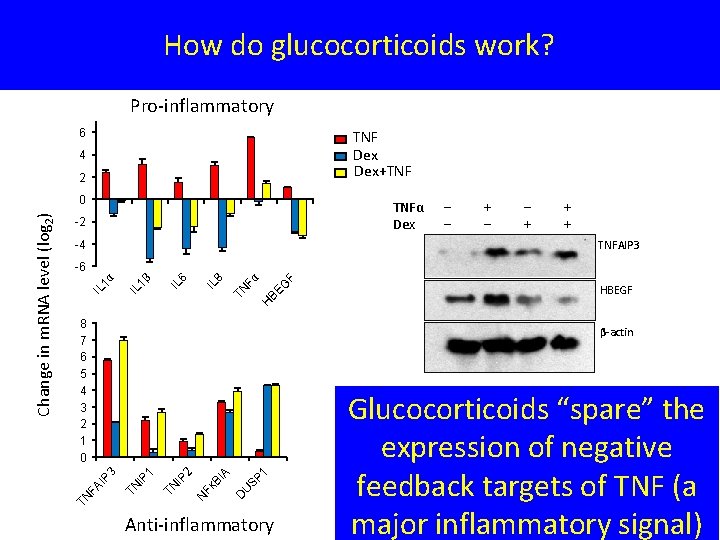
How do glucocorticoids work? Pro-inflammatory 6 TNF Dex+TNF 4 2 TNFα Dex -2 -4 ‒ ‒ + + + TNFAIP 3 HBEGF H BE G F Fα TN IL 8 IL 6 1β IL IL 1α -6 8 7 6 5 4 3 2 1 0 SP 1 U BI Fᴋ N D A 2 IP TN 1 IP TN IP 3 β-actin FA TN Change in m. RNA level (log 2) 0 Anti-inflammatory Glucocorticoids “spare” the expression of negative feedback targets of TNF (a major inflammatory signal)
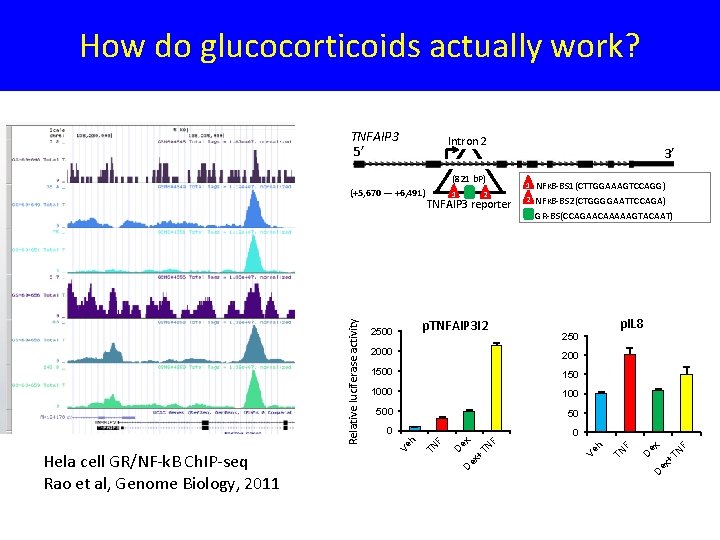
How do glucocorticoids actually work? Intron 2 NFᴋB-BS 1(CTTGGAAAGTCCAGG) 2 NFᴋB-BS 2(CTGGGGAATTCCAGA) GR-BS(CCAGAACAAAAAGTACAAT) 0 0 TN ex + D D F 50 ex 500 ex + 100 D 1000 D 150 F 1500 ex 200 F 2000 TN p. IL 8 250 F TNFAIP 3 reporter 1 TN 2 p. TNFAIP 3 I 2 2500 Ve h Relative luciferase activity Hela cell GR/NF-k. B Ch. IP-seq Rao et al, Genome Biology, 2011 1 Ve h (821 b. P) (+5, 670 — +6, 491) 3’ TN TNFAIP 3 5’
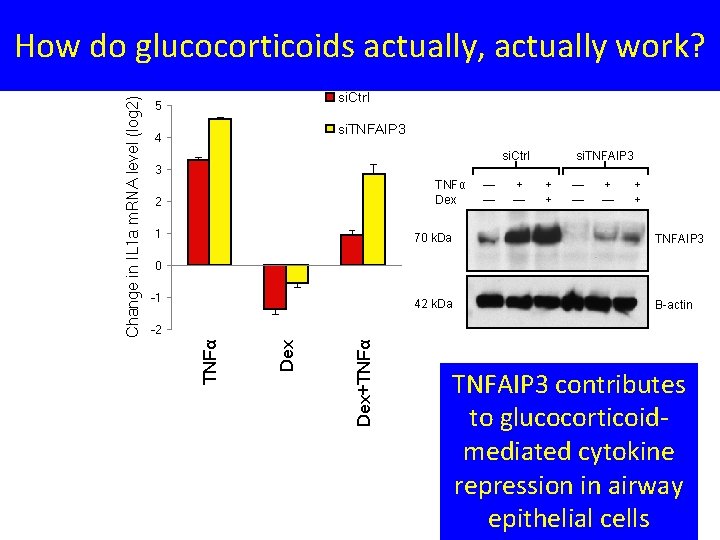
si. Ctrl 5 B si. TNFAIP 3 4 si. Ctrl si. TNFAIP 3 3 TNFα Dex 2 1 — — + — + + 70 k. Da TNFAIP 3 42 k. Da Β-actin 0 -1 Dex+TNFα Dex -2 TNFα Change in IL 1 a m. RNA level (log 2) How do glucocorticoids actually, actually work? TNFAIP 3 contributes to glucocorticoidmediated cytokine repression in airway epithelial cells

How do glucocorticoids work and what prevents them from working in asthma? Since GR interactions with DNA define GR activity study GR interactions with DNA in airway cells Since GR interactions with inflammatory factors are important for GC efficacy Study DNA-based interactions between GR and NF-k. B No current data on GR cistrome in airway cells…
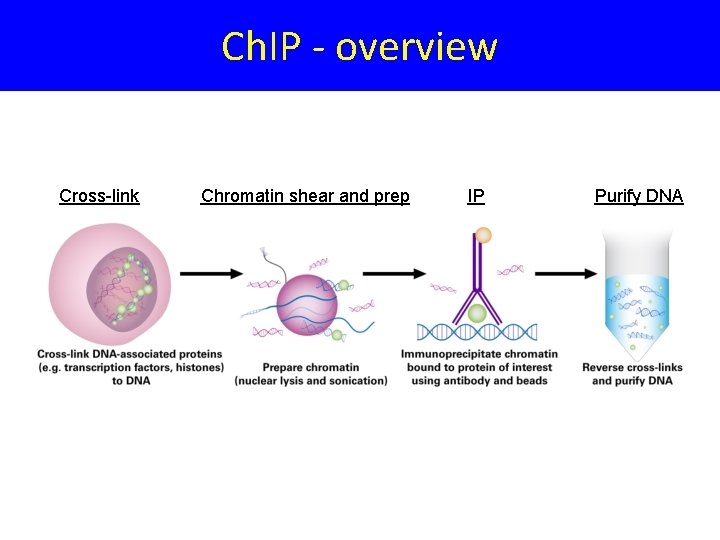
Ch. IP - overview Cross-link Chromatin shear and prep IP Purify DNA
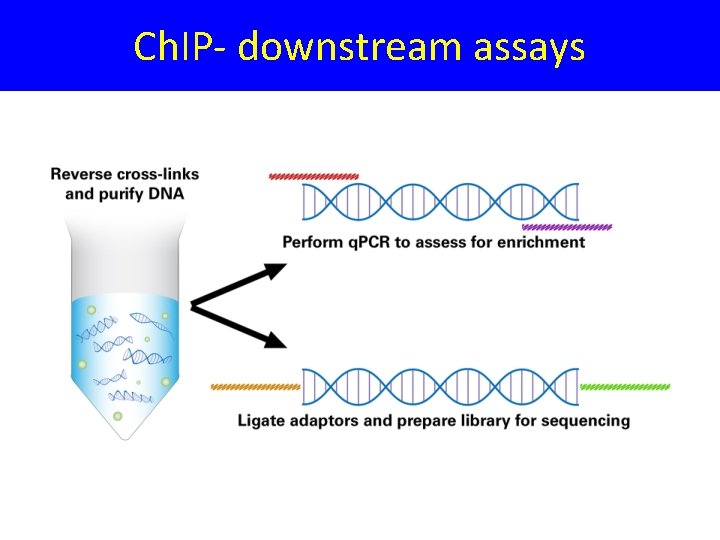
Ch. IP- downstream assays
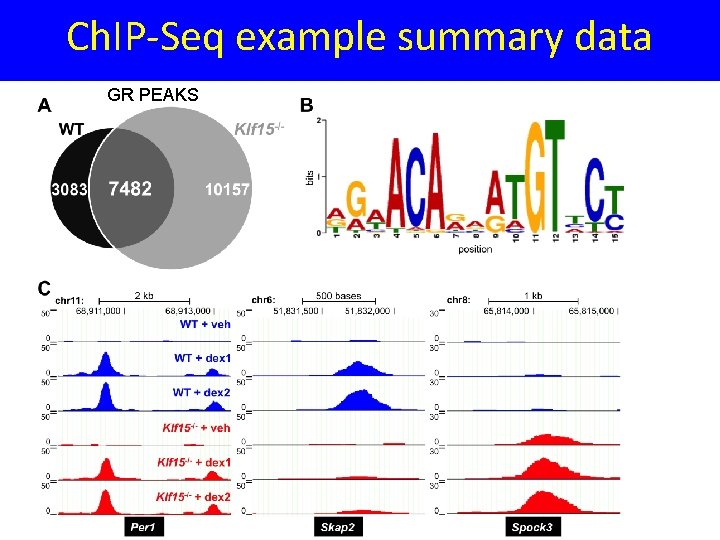
Ch. IP-Seq example summary data GR PEAKS
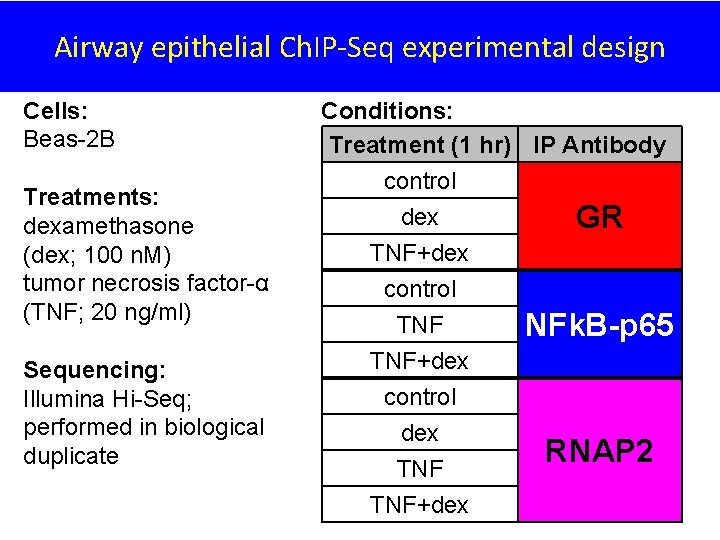
Airway epithelial Ch. IP-Seq experimental design Cells: Beas-2 B Treatments: dexamethasone (dex; 100 n. M) tumor necrosis factor-α (TNF; 20 ng/ml) Sequencing: Illumina Hi-Seq; performed in biological duplicate Conditions: Treatment (1 hr) IP Antibody control dex GR TNF+dex control dex TNF+dex NFk. B-p 65 RNAP 2
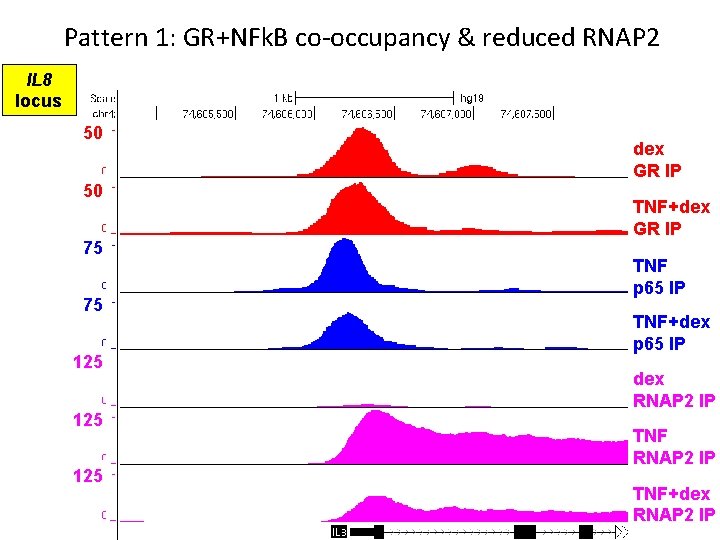
Pattern 1: GR+NFk. B co-occupancy & reduced RNAP 2 IL 8 locus 50 50 75 75 125 125 dex GR IP TNF+dex GR IP TNF p 65 IP TNF+dex p 65 IP dex RNAP 2 IP TNF+dex RNAP 2 IP
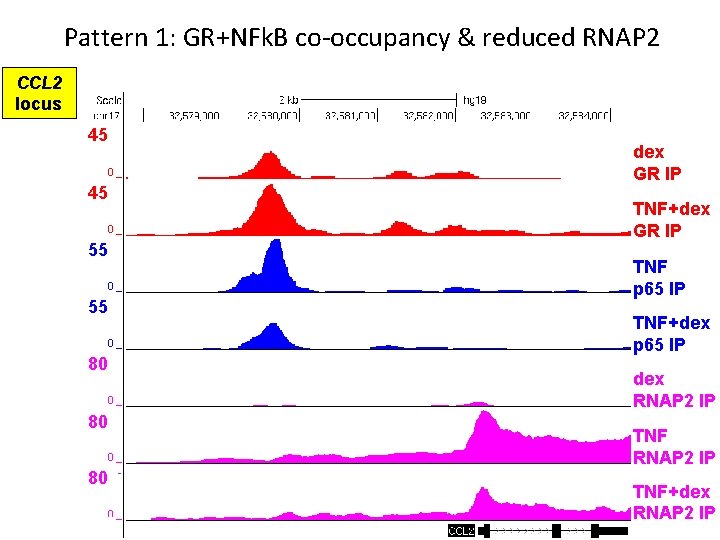
Pattern 1: GR+NFk. B co-occupancy & reduced RNAP 2 CCL 2 locus 45 45 55 55 80 80 80 dex GR IP TNF+dex GR IP TNF p 65 IP TNF+dex p 65 IP dex RNAP 2 IP TNF+dex RNAP 2 IP

Pattern 1 summary • GR binds pro-inflammatory gene in absence of TNF • GR occupancy is maintained/enhanced in presence of NFk. B • GR+NFk. B co-occupancy reduces RNAP 2 recruitment NET EFFECT = repression of pro-inflammatory transcription
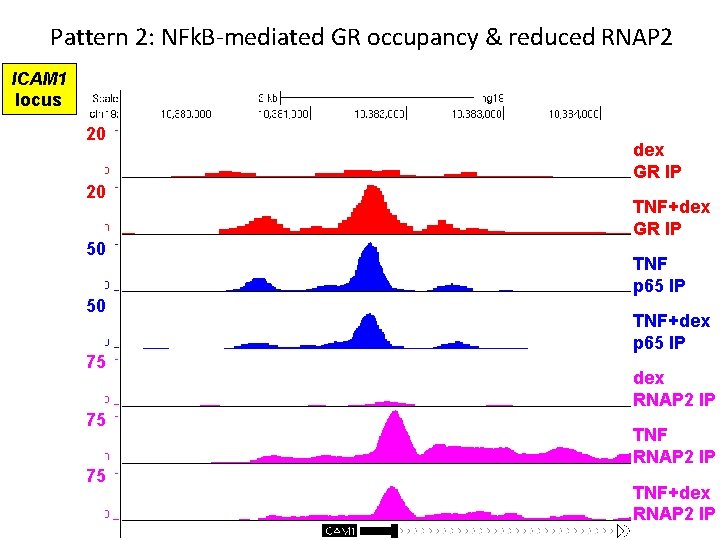
Pattern 2: NFk. B-mediated GR occupancy & reduced RNAP 2 ICAM 1 locus 20 20 50 50 75 75 75 dex GR IP TNF+dex GR IP TNF p 65 IP TNF+dex p 65 IP dex RNAP 2 IP TNF+dex RNAP 2 IP

Pattern 2 summary • GR binds pro-inflammatory gene only in presence of TNF • Role of NFk. B in GR recruitment unclear, possibly indirect • GR+NFk. B co-occupancy reduces RNAP 2 recruitment NET EFFECT = repression of pro-inflammatory transcription
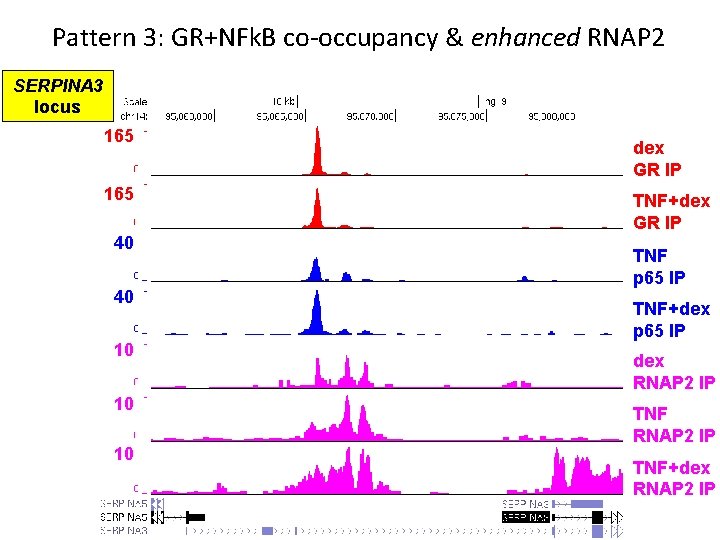
Pattern 3: GR+NFk. B co-occupancy & enhanced RNAP 2 SERPINA 3 locus 165 40 40 10 10 10 dex GR IP TNF+dex GR IP TNF p 65 IP TNF+dex p 65 IP dex RNAP 2 IP TNF+dex RNAP 2 IP
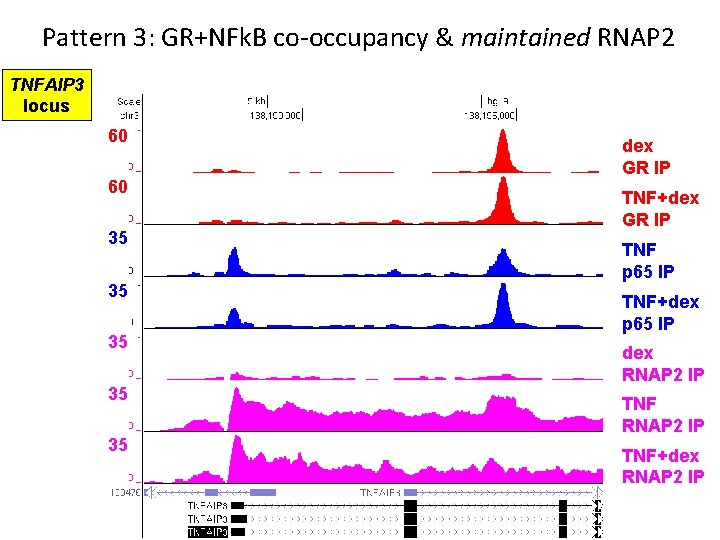
Pattern 3: GR+NFk. B co-occupancy & maintained RNAP 2 TNFAIP 3 locus 60 60 35 35 35 dex GR IP TNF+dex GR IP TNF p 65 IP TNF+dex p 65 IP dex RNAP 2 IP TNF+dex RNAP 2 IP

Pattern 3 Summary • GR binds anti-inflammatory genes with dex+TNF • GR+NFk. B co-occupancy does not appear antagonistic • GR+NFk. B co-occupancy enhances or maintains RNAP 2 recruitment NET EFFECT = activation (or sparing) of antiinflammatory transcription

What do we want from our Ch. IP data? BASIC 1. Identification of peaks for GR and p 65 under each condition and r values 2. Identification of differential GR binding vehicle vs. dex 3. Identification of differential binding p 65 binding vehicle vs. TNF treatment

What do we want from our Ch. IP data? BASIC 1. Identification of peaks for GR and p 65 under each condition and r values 2. Identification of differential GR binding vehicle vs. dex 3. Identification of differential binding p 65 binding vehicle vs. TNF treatment

What do we want from our Ch. IP data? ADVANCED 1. Differential GR binding (dex vs dex + TNF) 2. Differential p 65 binding (TNF vs TNF + dex) 3. RNAP 2 patterns 1. 2. 3. 4. 5. 6. Increased at TSS with dex vs vehicle Closest GR peak Increased at TSS with TNF and TNF + dex > TNF Increased at TSS with TNF and TNF + dex < TNF Closest p 65 peak to TSS for patterns 3 and 4 Closest GR peak to TSS for patterns 3 and 4

What do we want from our Ch. IP data? Collaboration Level! 1. Compare GR binding between Gerber lab data set and paper from Rao et al (Genome Biology, 2011) 2. Compare p 65 binding between Gerber lab data set and paper from Rao et al (Identification of differential GR binding vehicle vs. dex 3. Compare regulatory outcomes – i. e. correlate with RNAP 2 occupancy

QUESTIONS?
- Slides: 27