Giancarlo Mauri Universit di MilanoBicocca Italy Stochastic modelling

Giancarlo Mauri Università di Milano-Bicocca (Italy) Stochastic modelling of biological systems: membrane systems in Systems Biology Pisa, 24. 10. 2007

Lab. of Bioinformatics and Natural Computing - Activities Ø Bioinformatics ü Tools for sequence analysis ü Alternative splicing prediction ü Approximation algorithms (Fingerprint Clustering with Bounded Number of Missing Values; Maximum Isomorphic Agreement Subtree; LCS) Ø DNA Computing ü Splicing systems and formal languages ü DNA word design Ø Membrane systems ü Computational power of P-systems ü Modeling of biological systems ü Stochastic software simulators (Gillespie, …) ü Brane calculi vs P-systems Pisa, 24. 10. 2007

Outline Ø Membrane systems (P-systems) ü Definition ü Computational power of P-systems Ø Modeling biological processes with Psystems Ø Stochasticity in nature ü Stochastic P-systems ü Stochastic simulation algorithms Ø Simulations with stochastic P-systems ü Ras/c. AMP/PKA pathway in yeast Pisa, 24. 10. 2007

Structure of living cells Pisa, 24. 10. 2007

Abstracting features Ø A hierarchical arrangement of organelles, with separating membranes Ø Each membrane delimits a region Ø Each region contains a multiset of elements (simple molecules, DNA sequences, other regions) Ø The chemicals/bio-elements evolve in time according to some (rewriting/combination) rules specific to each region or may be moved across the membranes Ø The rules may also dissolve/create/move regions Pisa, 24. 10. 2007
![Membrane (P-)systems [G. Paun, 1998] Ø Distributed, parallel and nondeterministic computing model Ø Basic Membrane (P-)systems [G. Paun, 1998] Ø Distributed, parallel and nondeterministic computing model Ø Basic](http://slidetodoc.com/presentation_image/f51d99337a82c163ca1f11bebb61a0f4/image-6.jpg)
Membrane (P-)systems [G. Paun, 1998] Ø Distributed, parallel and nondeterministic computing model Ø Basic ingredients: ü membrane structure (finite string of well matching parentheses) ü (multisets of) objects (symbols, strings, etc. , ) in each membrane ü evolution rules associated to each membrane: objects can evolve within membranes, move into neighboring membranes; membranes can be dissolved/divided/created/merged. ü communication of objects among membranes Pisa, 24. 10. 2007
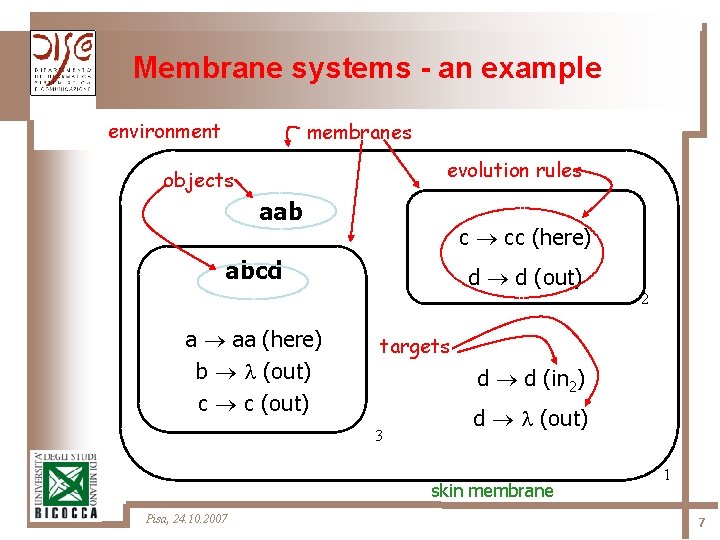
Membrane systems - an example environment membranes evolution rules objects aab c cc (here) abcd a aa (here) b (out) c c (out) d d (out) targets d d (in 2) 3 d (out) skin membrane Pisa, 24. 10. 2007 2 1 7

Membrane systems Ø The membrane structure ü separation and communication ü identification of relevant “spaces” Ø The rules ü object evolution ü object communication ü membrane evolution Ø Peculiar roles of objects ü catalysts, promoters, inhibitors Pisa, 24. 10. 2007

Computing with Membrane systems Ø Initial configuration, identified by ü membrane structure ü initial multisets of objects ü sets of evolution rules Ø Nondeterministic and maximal application of evolution rules parallel ü universal clock (synchronous) ü all membranes are simultaneously processed ü all applicable rules are (nondeterministically) applied Ø Successful computation: ü no rule can be further applied ü output (multi)sets of objects collected in a prescribed membrane or outside the system Pisa, 24. 10. 2007

Computing with membranes: the TCS perspective Ø Computability aspects ü Generated languages ü Computational power, Turing completeness, universality ü Complexity classes Ø Comparison with other models Ø Polynomial solutions to hard problems (timespace trade-off) Ø Applications: biology, bio-medicine, economics, linguistics, computer science, optimization Ø Software and simulations Pisa, 24. 10. 2007

Variants of the basic model Ø Ø Ø Ø Catalytic membrane systems Symport/antiport membrane systems Membrane systems with string objects Active membranes Polarized membranes of variable thickness Spiking neural membrane systems …. Pisa, 24. 10. 2007

Example of Symport/Antiport System (oo, out) (oo, in) 2 1 3 (o, in) oooo (oo, out; o, in) Types of rules (no transformation of objects): symport: (u, out) or (u, in) antiport: (u, out; v, in) where u, v are multisets Symbols (above example is unary) available abundantly in the environment. Pisa, 24. 10. 2007
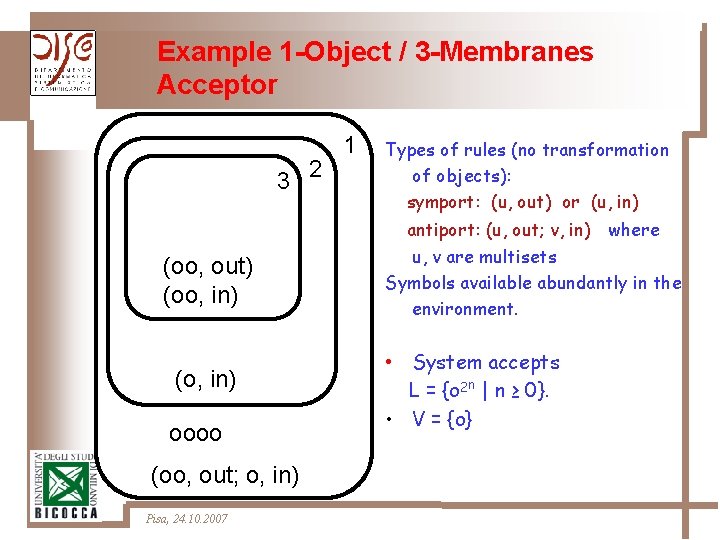
Example 1 -Object / 3 -Membranes Acceptor 3 2 (oo, out) (oo, in) (o, in) oooo (oo, out; o, in) Pisa, 24. 10. 2007 1 Types of rules (no transformation of objects): symport: (u, out) or (u, in) antiport: (u, out; v, in) where u, v are multisets Symbols available abundantly in the environment. • System accepts L = {o 2 n | n ≥ 0}. • V = {o}
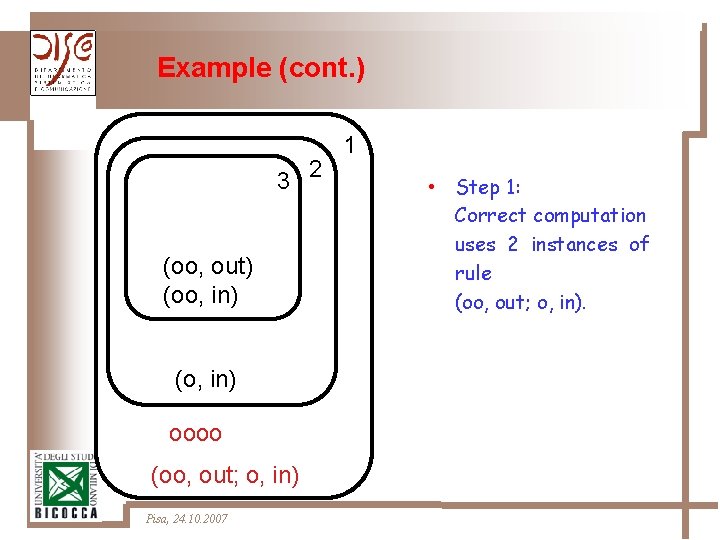
Example (cont. ) 3 2 (oo, out) (oo, in) (o, in) oooo (oo, out; o, in) Pisa, 24. 10. 2007 1 • Step 1: Correct computation uses 2 instances of rule (oo, out; o, in).
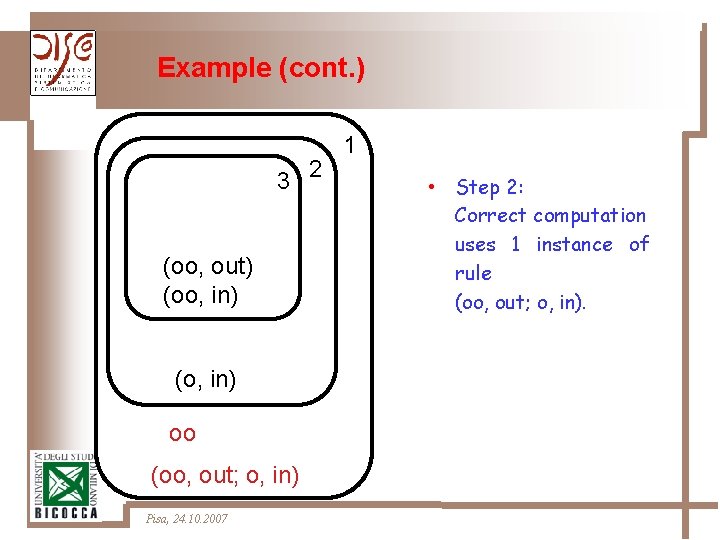
Example (cont. ) 3 2 (oo, out) (oo, in) (o, in) oo (oo, out; o, in) Pisa, 24. 10. 2007 1 • Step 2: Correct computation uses 1 instance of rule (oo, out; o, in).
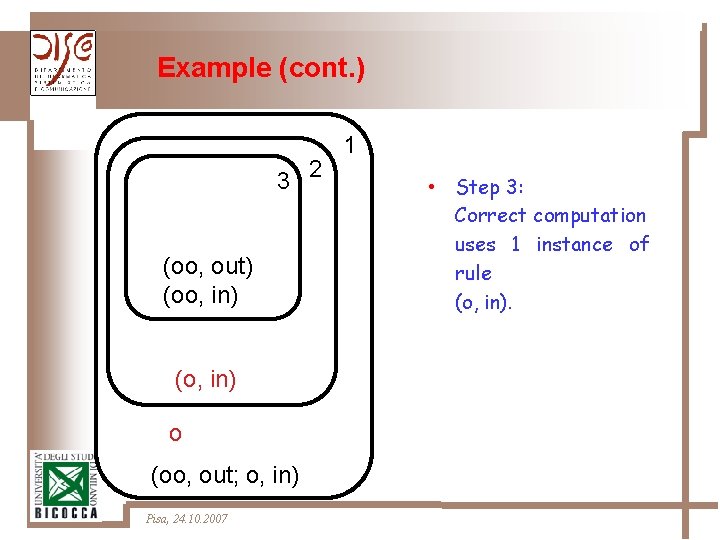
Example (cont. ) 3 2 (oo, out) (oo, in) (o, in) o (oo, out; o, in) Pisa, 24. 10. 2007 1 • Step 3: Correct computation uses 1 instance of rule (o, in).
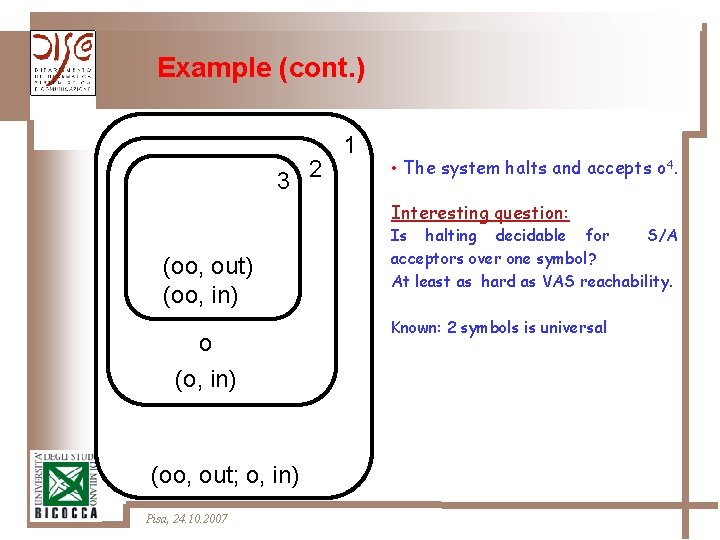
Example (cont. ) 3 2 1 • The system halts and accepts o 4. Interesting question: (oo, out) (oo, in) o (o, in) (oo, out; o, in) Pisa, 24. 10. 2007 Is halting decidable for S/A acceptors over one symbol? At least as hard as VAS reachability. Known: 2 symbols is universal

References Ø Paun, Gh. : Membrane Computing. An Introduction. Springer, Berlin (2002) Ø The P Systems Web Page: http: //psystems. disco. unimib. it/ ü bibliography ü download papers ü download simulators Pisa, 24. 10. 2007

Back to biology Ø Use of the framework of P-systems to give a formal description of specific cellular phenomena or cellular structures Ø Software tools for dynamical analysis of biological systems Pisa, 24. 10. 2007
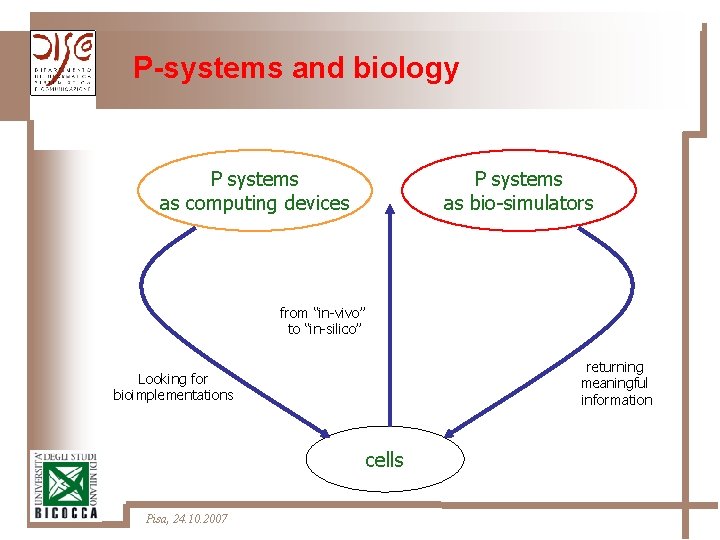
P-systems and biology P systems as computing devices P systems as bio-simulators from “in-vivo” to “in-silico” returning meaningful information Looking for bioimplementations cells Pisa, 24. 10. 2007

Systems biology Ø The “omics” era ü holistic approach to the interpretation and the analysis of biological systems (Ideker et al. , Ann. Rev. Gen. Hum. Gen. , 2001; Kitano, MIT Press, 2001): q investigation of the system at a global scale (components and interactions ! global behaviour) q integration and representation of quantitative and qualitative data q different types of perturbations Ø Modeling biological systems ü identification of structure and parameters ü analysis of system dynamics via software simulations methods for system control and design Pisa, 24. 10. 2007

From structures (syntax) to functions (semantics) in Biology Ø Bio-components as information and computational devices Ø Millions of simultaneous computational threads active (e. g. , metabolic networks, gene regulatory networks, signaling pathways) Ø Components interaction changes the future behavior Ø Interactions occur only if components are correctly located (e. g. , they are close enough or they are not divided by membranes) Pisa, 24. 10. 2007

Membrane systems and Systems Biology Ø Advantages: ü parallel and stochastic processing ü discreteness ü cellular localizations ü easily comprehensible ü scalability Pisa, 24. 10. 2007

Applications of membrane systems Ø Membrane systems as modelling tools at molecular and cellular scale: ü transport proteins Ø Na+/K+ pump, Ca 2+ channels, mechanosensitive channels ü chemical reactions Ø Belousov-Zhabotinsky, Michaelis-Menten ü cellular signaling pathways Ø EGFR, Ras/c. AMP/PKA ü bacterial colonies Ø Vibrio fischeri, Pseudomonas aeruginosa Pisa, 24. 10. 2007
Applications of membrane systems Ø Membrane systems as modelling tools at ecological scale: ü Lotka-Volterra equation Ø population dynamics ü tritrophic systems Ø plants+herbivores+carnivores ü metapopulations Ø populations living in fragmented habitats Pisa, 24. 10. 2007

Other approaches Ø ODE (global, monolithic, difficult) Ø Petri nets Ø The pi-calculus (Milner, Parrow, Walker 1992) Ø The stochastic pi-calculus (Priami 1995) Ø (Mem)Brane calculus (Cardelli 2004) Pisa, 24. 10. 2007

Stochasticity in biological systems Ø “Noise” in nature many experimental evidences of stochasticity in living systems ü observation of genes expression has shown the stochastic nature of transcription and translation [Abkowitz et al, 1996; Ozbudak et al, 2002] ü the m. RNA production is “quantal” [Hume, 2000] and occurs in random pulses [Ross et al, 1994; Walters et al, 1995] ü the protein production occurs in short bursts and at random time intervals [Yarchuk et al, 1992; Chapon, 1982] ü lysis/lysogeny switch in the -phage [Oppenheim et al, 2005] Pisa, 24. 10. 2007

Stochasticity in biological systems Ø 2 kinds of noise: ü intrinsic noise - due to the inherent nature of the biochemical interactions ü extrinsic noise - due to the external environmental conditions Ø Complex systems such as the biological ones are extremely non-linear and often exhibits many steady states, bifurcations or chaotic behavior Pisa, 24. 10. 2007

Stochasticity in biological systems Ø Stochastic models framework because: are suitable in this ü take into consideration discrete quantities of components, ü are in accordance with the stochastic processes, ü are appropriate to describe "small systems" and instability phenomena. Ø Stochastic simulation is the probe to access the different evolutions Pisa, 24. 10. 2007
![Stochastic simulation algorithm Ø Stochastic Simulation Algorithm [Gillespie, 1977] ü N chemical species Si Stochastic simulation algorithm Ø Stochastic Simulation Algorithm [Gillespie, 1977] ü N chemical species Si](http://slidetodoc.com/presentation_image/f51d99337a82c163ca1f11bebb61a0f4/image-30.jpg)
Stochastic simulation algorithm Ø Stochastic Simulation Algorithm [Gillespie, 1977] ü N chemical species Si in a single fixed volume V, with Xi current number of molecules of Si ü M chemical reactions Rμ, with reaction parameters cμ ü 2 questions: when will the next reaction occur? Which reaction will it be? ü SSA computes the probability that the next reaction in V will occur in the differential time interval (t+ , t+ +d ) and will be Rμ ü huge computational time needed: it increases with the number of reactant species (and of reaction channels) Pisa, 24. 10. 2007

Stochastic membrane systems: DPP (Pescini et al. , 2006) Ø Rules are applied according to the probability associated with them via a known function Ø At each step the probability changes dynamically looking after the systems variations Ø To have a parallel step the probability distribution is: ü fixed by the actual system state ü kept constant for the whole parallel step Pisa, 24. 10. 2007
![Stochastic simulation algorithm Ø Tau Leaping stochastic simulation method [Gillespie et al, 2006]: ü Stochastic simulation algorithm Ø Tau Leaping stochastic simulation method [Gillespie et al, 2006]: ü](http://slidetodoc.com/presentation_image/f51d99337a82c163ca1f11bebb61a0f4/image-32.jpg)
Stochastic simulation algorithm Ø Tau Leaping stochastic simulation method [Gillespie et al, 2006]: ü used to speed up SSA ü during every leap ( ) several reactions are executed ü the rules execution order, within each step, does not matter ü the complexity of the algorithm increases linearly with the number of reactant species Pisa, 24. 10. 2007

Stochastic membrane systems: DPP Ø The membrane system structure is exploited to extend the Tau Leaping procedure to multiple volumes systems ü tracing the time of each membrane as well as the time of the whole system q qualitative and quantitative evolution ü communication of objects among membranes as in standard membrane systems ü computational time increases with the number of reactant species and with the number of volumes Pisa, 24. 10. 2007

-DPP: How it works Ø There is a -leaping engine in every membrane. Ø Every membrane generates a value based on its internal state Ø The system evolves according to the smallest of the system. Ø In each membrane the probability distribution is generated according to ü ü the underlying process ü the system status. Pisa, 24. 10. 2007
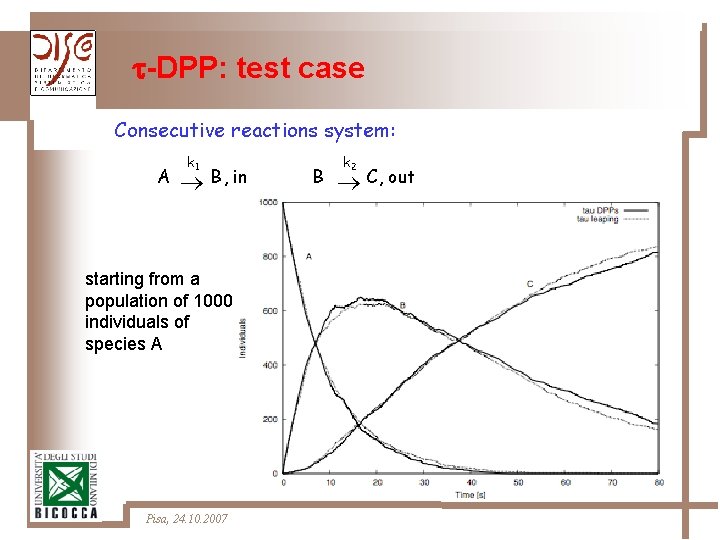
-DPP: test case Consecutive reactions system: A k 1 B, in starting from a population of 1000 individuals of species A Pisa, 24. 10. 2007 B k 2 C, out

Resources Ø Cluster with Beowulf architecture, with 30 nodes: ü AMD Athlon(TM) XP 2800+ processors ü Linux OS Ø Stochastic simulator: ü C language code ü MPI library (http: //www. mcs. anl. gov/mpi/index. html) ü mpicc compiler Pisa, 24. 10. 2007
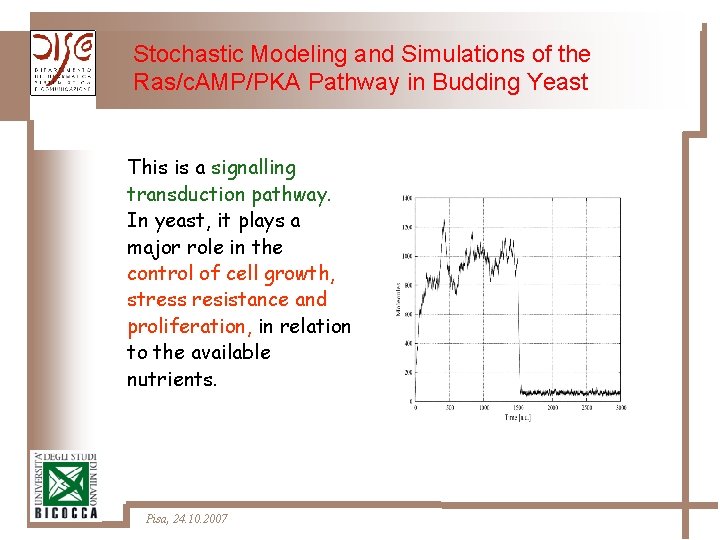
Stochastic Modeling and Simulations of the Ras/c. AMP/PKA Pathway in Budding Yeast This is a signalling transduction pathway. In yeast, it plays a major role in the control of cell growth, stress resistance and proliferation, in relation to the available nutrients. Pisa, 24. 10. 2007
Ras/c. AMP/PKA Pathway Cyclic AMP plays a key regulatory role in yeast growth: High c. AMP Required for growth and cell cycle progression. Low stress resistance. Low c. AMP Arrest of growth and cell cycle in G 1/G 0, accumulation of trehalose and glicogen. Stress resistance A moderate PKA activity is required for growth. PKA activates protein synthesis, ribosomes and r. RNA synthesis and synthesis of Cln 3. Pisa, 24. 10. 2007

Ras/c. AMP/PKA Pathway The Ras/c. AMP/PKA pathway in yeast trasduces two different signals: 1) Allows the G 1/S transition at START through a “nutrient sensing” mechanism ( still not defined. . ), likely through a modulation of the activity of Cdc 25 or Ira proteins, and also regulates Ps (cell size required for entry in S phase) 2) Generates a peak of PKA activity (mediated by a fast c. AMP increase) in response to addition of fermentable sugars (glucose and fructose). This signalling regulates the transition between respiratory and fermentative metabolism. Pisa, 24. 10. 2007 39
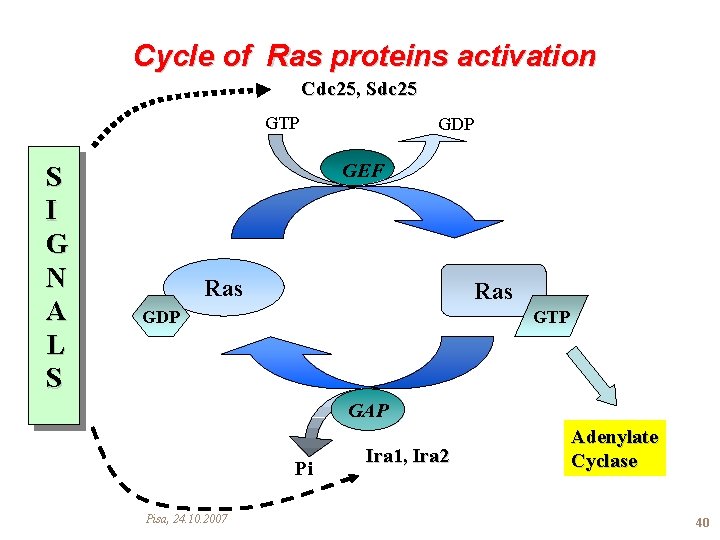
Cycle of Ras proteins activation Cdc 25, Sdc 25 GTP S I G N A L S GDP GEF Ras GDP GTP GAP Pi Pisa, 24. 10. 2007 Ira 1, Ira 2 Adenylate Cyclase 40
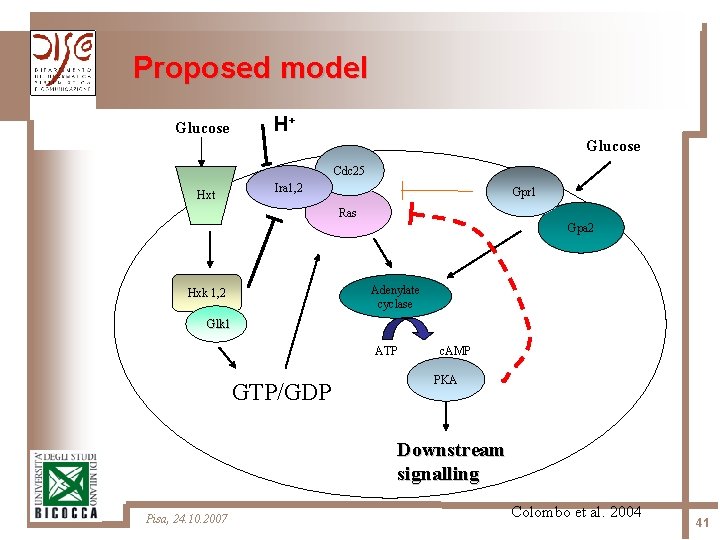
Proposed model H+ Glucose Cdc 25 Ira 1, 2 Hxt Gpr 1 Ras Gpa 2 Adenylate cyclase Hxk 1, 2 Glk 1 ATP GTP/GDP c. AMP PKA Downstream signalling Pisa, 24. 10. 2007 Colombo et al. 2004 41
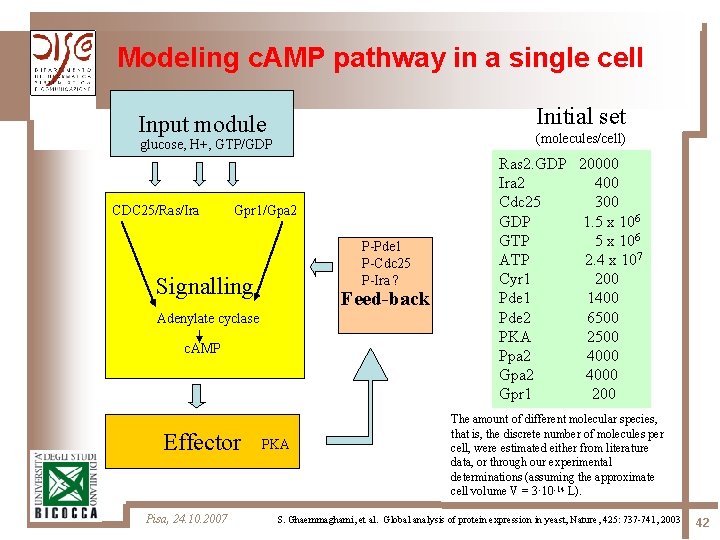
Modeling c. AMP pathway in a single cell Initial set Input module (molecules/cell) glucose, H+, GTP/GDP CDC 25/Ras/Ira Gpr 1/Gpa 2 P-Pde 1 P-Cdc 25 P-Ira ? Signalling Feed-back Adenylate cyclase c. AMP Effector Pisa, 24. 10. 2007 PKA Ras 2. GDP 20000 Ira 2 400 Cdc 25 300 GDP 1. 5 x 106 GTP 5 x 106 ATP 2. 4 x 107 Cyr 1 200 Pde 1 1400 Pde 2 6500 PKA 2500 Ppa 2 4000 Gpr 1 200 The amount of different molecular species, that is, the discrete number of molecules per cell, were estimated either from literature data, or through our experimental determinations (assuming the approximate cell volume V = 3· 10 -14 L). S. Ghaemmaghami, et al. Global analysis of protein expression in yeast, Nature, 425: 737 -741, 2003 42

Methodology 4 main logical modules have been identified in the model: Ø the switch cycle of Ras 2 protein, involving the GEF Cdc 25 and the GAP Ira 2 Ø the synthesis of c. AMP, via activation of adenylate cyclase Cyr 1 Ø the activation of PKA, via the reversible binding between its regulatory subunits R and c. AMP, and subsequent dissociation of the tetramer PKA Ø the activity of phosphodiesterases Pde 1 and Pde 2, which determines the feedback mechanisms for c. AMP degradation Pisa, 24. 10. 2007

Ras/c. AMP/PKA signaling pathway Ø Using tau-DPPs, we can simulate systems structured by several volumes, tracing the simulated time of the compartments as well as time line of the whole system. This gives us the possibility to quantitatively and qualitatively describe biological systems. Ø Our model was able to simulate properly the Ras protein cycle, the activation of adenylate cyclase, the production of cyclic AMP and the activation of c. AMP-dependent protein kinase in a single yeast cell. The results are compared with the experimental data and give information on the key regulatory elements of this signalling network. Pisa, 24. 10. 2007

Ras/c. AMP/PKA signaling pathway Ø The model involves: ü 34 rules ü 30 molecular species ü 1 major feedback ü many Michaelis Menten schemes Pisa, 24. 10. 2007

Ras/c. AMP/PKA signaling pathway Reaction Reagents Products Constant r 1 Ras 2·GDP+Cdc 25 Ras 2·GDP·Cdc 25 1 r 2 Ras 2·GDP·Cdc 25 Ras 2·GDP+Cdc 25 1 r 3 Ras 2·GDP·Cdc 25 Ras 2·Cdc 25+GDP 1. 5 r 4 Ras 2·Cdc 25+GDP Ras 2·GDP·Cdc 25 1 r 5 Ras 2·Cdc 25+GTP Ras 2·GTP·Cdc 25 1 r 6 Ras 2·GTP·Cdc 25 Ras 2·Cdc 25+GTP 1 r 7 Ras 2·GTP·Cdc 25 Ras 2·GTP+Cdc 25 1 r 8 Ras 2·GTP+Cdc 25 Ras 2·GTP·Cdc 25 1 r 9 Ras 2·GTP+ ra 2 Ras 2·GTP·Ira 2 3· 10− 2 r 10 Ras 2·GTP·Ira 2 Ras 2·GDP+Ira 2 0. 7 r 11 Ras 2·GTP+CYR Ras 2·GTP·CYR 1 10− 3 r 12 Ras 2·GTP·CYR 1+ATP Ras 2·GTP·CYR 1+c. AMP 0. 21· 10− 5 r 13 Ras 2·GTP·CYR Ira 2 Ras 2·GDP+CYR 1+Ira 2 10− 3 Pisa, 24. 10. 2007
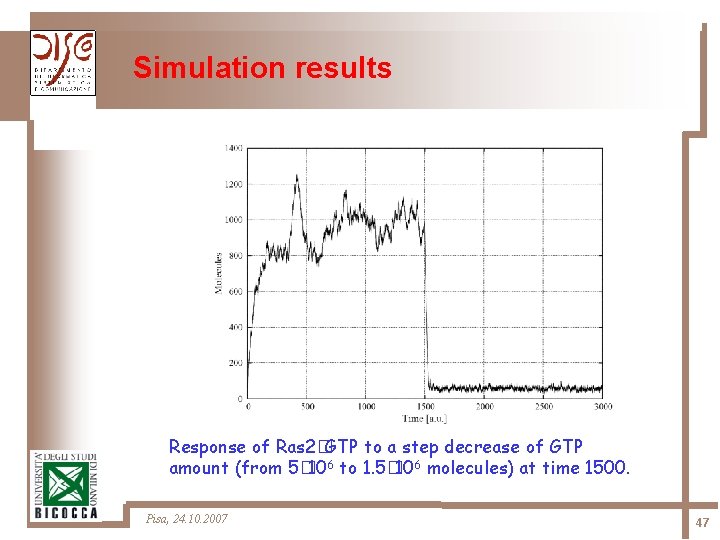
Simulation results Response of Ras 2�GTP to a step decrease of GTP amount (from 5� 106 to 1. 5� 106 molecules) at time 1500. Pisa, 24. 10. 2007 47

Simulation results Sensitivity of Ras 2�GTP. Left : dependence on the rate of dissociation of Cdc 25, (a) k 7 = 0. 2; (b) k 7 = 0. 5; (c) k 7 = 1. 0; (d) k 7 = 1. 5; (e) k 7 = 2. 0. Right: dependence on the activity of Ira 2, (a) k 10 = 0. 25; (b) k 10 = 0. 5; (c) k 10 = 0. 7; (d) k 10 = 0. 9; (e) k 10 = 1. 1. Pisa, 24. 10. 2007 48

Simulation results Sensitivity of Ras 2�GTP module. Pisa, 24. 10. 2007 49
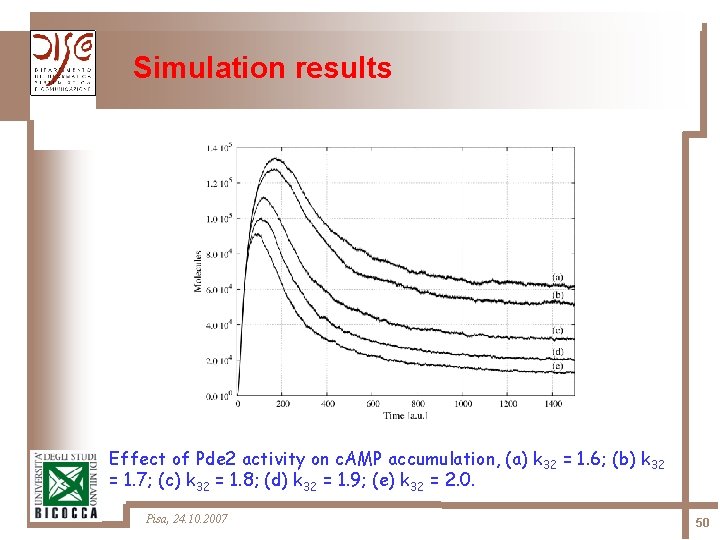
Simulation results Effect of Pde 2 activity on c. AMP accumulation, (a) k 32 = 1. 6; (b) k 32 = 1. 7; (c) k 32 = 1. 8; (d) k 32 = 1. 9; (e) k 32 = 2. 0. Pisa, 24. 10. 2007 50

Simulation results Effect of Pde 1 activity on c. AMP accumulation, (a) k 28 = 1. 7; (b) k 28 = 2. 0; (c) k 28 = 3. 0; (d) k 28 = 4. 0; (e) k 28 = 5. 0. Pisa, 24. 10. 2007 51
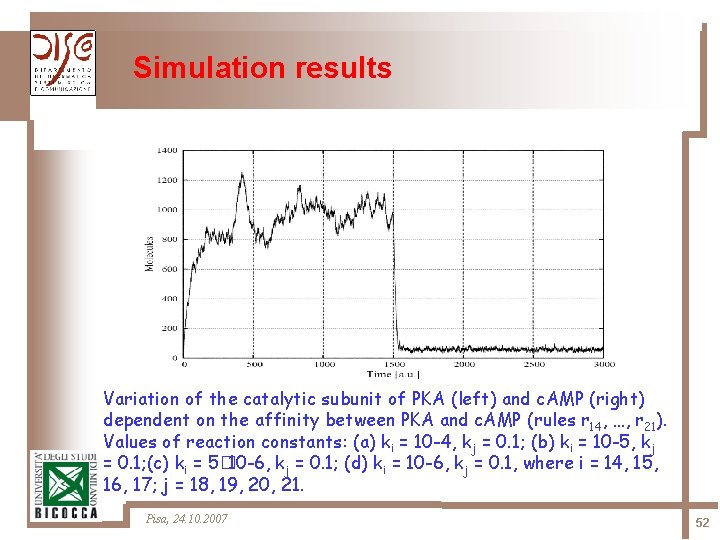
Simulation results Variation of the catalytic subunit of PKA (left) and c. AMP (right) dependent on the affinity between PKA and c. AMP (rules r 14, …, r 21). Values of reaction constants: (a) ki = 10 -4, kj = 0. 1; (b) ki = 10 -5, kj = 0. 1; (c) ki = 5� 10 -6, kj = 0. 1; (d) ki = 10 -6, kj = 0. 1, where i = 14, 15, 16, 17; j = 18, 19, 20, 21. Pisa, 24. 10. 2007 52
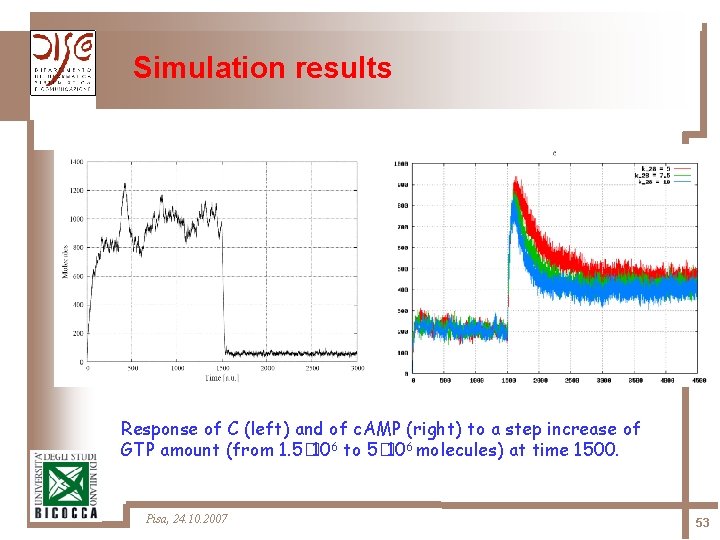
Simulation results Response of C (left) and of c. AMP (right) to a step increase of GTP amount (from 1. 5� 106 to 5� 106 molecules) at time 1500. Pisa, 24. 10. 2007 53

Simulation results Effect of different GTP input values on c. AMP accumulation (left) and on Ras 2�GTP and PKA activity (right). Pisa, 24. 10. 2007 54
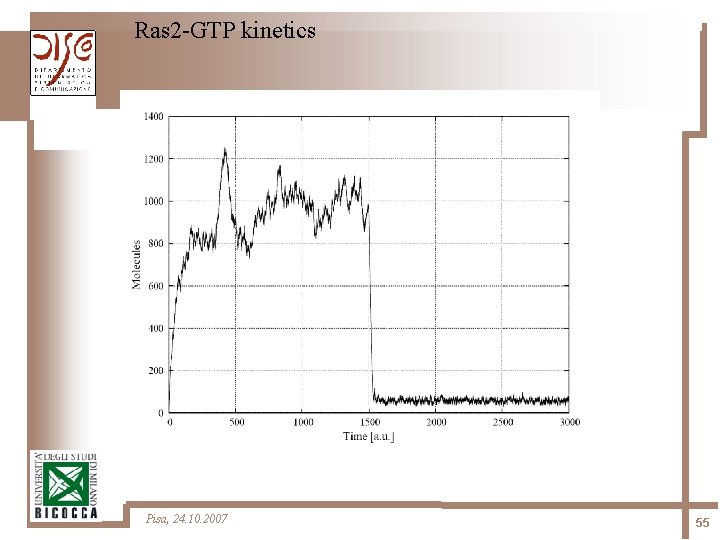
Ras 2 -GTP kinetics Pisa, 24. 10. 2007 55

Work in progress Ø Simulation tools ü DPP optimization ü integration of Genetic Algorithms ü topological distribution of molecular species in distinct cellular regions and/or presence of large signalling complexes localized in internal membranes Ø Analysis ü communicating classes and beyond ü role of “noise” in Molecular Dynamics-Computing Pisa, 24. 10. 2007

Work in progress Ø Biological Systems Modelling: ü Implementation of Gpr 1/Gpa 2 glucose sensing system (that requires t-DPP) ü Introduction of multiple feed-back levels (Ira, Cdc 25, Ras) ü Extensive sensitivity analysis, stability /instability behaviour etc. ü Biofilms formation in Pseudomonas aeruginosa and Escherichia coli colonies Pisa, 24. 10. 2007

References I. I. Ardelean, D. Besozzi, M. H. Garzon, G. Mauri, S. Roy P System Models for Mechanosensitive Channels In "Applications of Membrane Computing”, Springer, 2005 D. Pescini, D. Besozzi, C. Zandron, G. Mauri Analysis and Simulation of Dynamics in Probabilistic P Systems Proc. DNA 11, LNCS 3892, Springer, 2006 D. Besozzi, P. Cazzaniga, D. Pescini, G. Mauri Seasonal variance in P system models for metapopulations First Int. Conf. on Bio-Inspired Computing: Theory and Applications Pisa, 24. 10. 2007

References P. Cazzaniga, D. Pescini, D. Besozzi, G. Mauri Tau leaping stochastic simulation method in P Systems Proc. WMC 7, LNCS 4361, Springer, 2006 E. Martegani et al. Identification of an intracellular signalling complex for Ras/CAMP pathway in yeast: experimental evidences and modelling 25 th International Specialised Symposium on Yeasts, 2006 P. Cazzaniga, D. Besozzi, E. Martegani, S. Colombo, G. Mauri Stochastic modelling of the Ras/c. AMP signal transduction pathway in yeast Journal of Biotechnology, to appear Pisa, 24. 10. 2007

The research team Daniela Besozzi Paolo Cazzaniga Dario Pescini Giancarlo Mauri Università degli Studi di Milano Bicocca Dipartimento di Informatica, Sistemistica e Comunicazione Università degli Studi di Milano Dipartimento di Informatica e Comunicazione Sonia Colombo Enzo Martegani Università degli Studi di Milano. Bicocca Dipartimento di Biotecnologie e Bioscienze Pisa, 24. 10. 2007

Thanks! Pisa, 24. 10. 2007 61
- Slides: 61