Genotyping and Genetic Maps Bas Heijmans Leiden University

Genotyping and Genetic Maps Bas Heijmans Leiden University Medical Centre The Netherlands

Pedigree file in linkage format 1 1 2 2 2 1 2 3 4 5 0 0 1 1 1 0 0 2 2 2 1 1 2 1 1 1 3 2 2 0 0 5 6 5 2 4 3 4 0 0 8 7 7

Pedigree file in linkage format ) er rk 1 s d m fa 1 1 2 2 2 i ily pe n rso 1 2 3 4 5 id t fa 0 0 1 1 1 r he t x o se m 0 0 2 2 2 1 1 2 1 e s ea dis 1 1 1 1 1 tu a t s 1 3 2 2 0 0 5 6 5 ma rd a( t a e rk 2 4 3 4 0 0 8 7 7 ma

Marker choice for genome-wide linkage scans Short tandem repeats (STR, a. k. a. microsatellites) because: • High heterozygosity (1 STR ~ 5 SNPs) • There are more than enough (1/30 kb thus >>1/c. M) • Reliable genetic maps (Marshfield, Decode) • Optimized marker sets, spacing down to 5 c. M (Marshfield/Applied Biosystems) • Reasonably automated measurement (2 persons 40, 000 checked genotypes in database per week) • Low cost per genotype (<$0. 15 for consumables) • Reasonable success and error rates (>92% and <0. 8%)
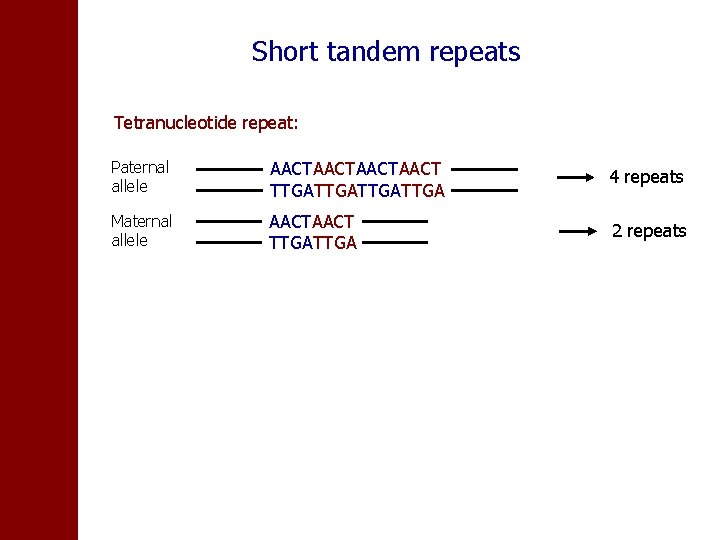
Short tandem repeats Tetranucleotide repeat: Paternal allele AACTAACT TTGATTGA 4 repeats Maternal allele AACT TTGA 2 repeats
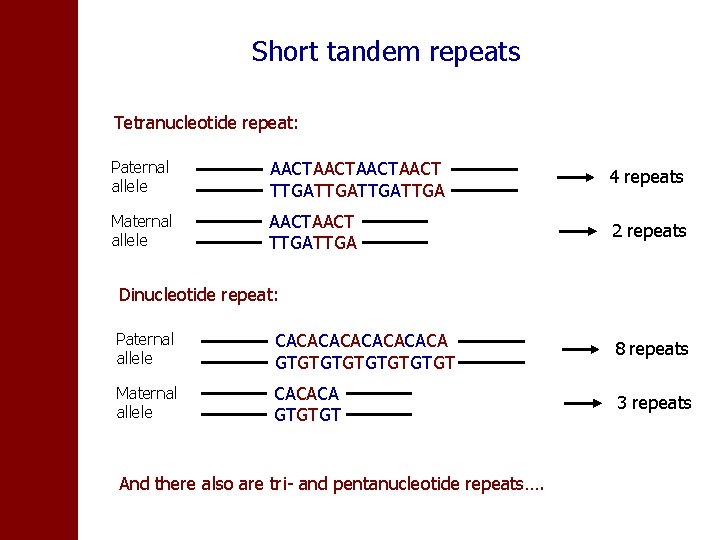
Short tandem repeats Tetranucleotide repeat: Paternal allele AACTAACT TTGATTGA 4 repeats Maternal allele AACT TTGA 2 repeats Dinucleotide repeat: Paternal allele CACACACA GTGTGTGT 8 repeats Maternal allele CACACA GTGTGT 3 repeats And there also are tri- and pentanucleotide repeats….

Principle of genotyping methods • Short tandem repeats length differences CACACACA GTGTGTGT CACACA GTGTGT • SNPs only sequence difference G C A T • Destruction restriction site (RFLP) • Hybridization differences (Taq. Man) • One base-pair sequencing reactionprimer extension (Sequenom, Orchid) • Ligation assay (Illumina) • VNTR, insertion/deletion polymorphisms (1 bp to ~300 bp for Alu repeat)
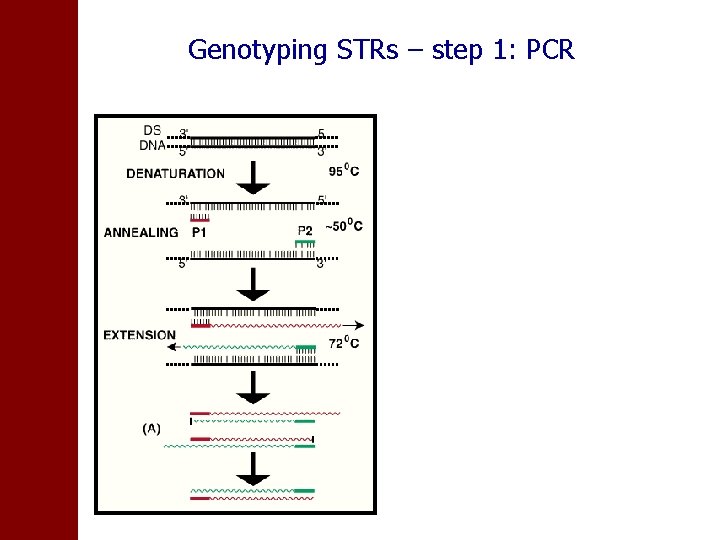
Genotyping STRs – step 1: PCR
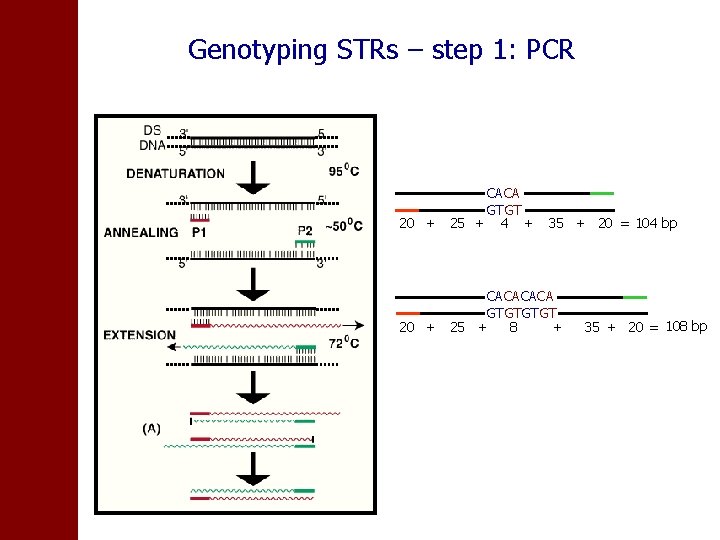
Genotyping STRs – step 1: PCR CACA GTGT 20 + 25 + 4 + 35 + 20 = 104 bp CACA GTGT 20 + 25 + 8 + 35 + 20 = 108 bp
Genotyping STRs – step 1: PCR in practice genomic DNA + primers + Taq DNA polymerase + d. NTPs (ACGT) + buffer

Genotyping STRs – step 2: electophoresis Detect length differences Agarose or polyacrylamide slab gel • DNA is negatively charged • Longer fragments migrate slower than shorter ones through polymer network. — electrode + electrode

To scan the whole human genome… • 1 short tandem repeat every 10 c. M • makes 400 markers per individual • Assuming 1000 individuals (preferably 1000 s) • One whole genome scan = 400, 000 genotypings

Not like this…….

Not like this……. but like this 96 -well plates 384 -well plates

Not like this…….

Not like this……. but like this

Not like this…….

Not like this……. but like this

Electrophoresis using automated sequencer • 96 capillaries (no lanes) (ABI 3700) • Put in machine and all goes automatically • Primers are labelled with fluorescent dye • Machine detects PCR products through a laser start Typically 15 markers in one capillary: CACA GTGT CACA TCTC AGAG Detector Laser 2. 5 h TGTGTG ACACAC + A bit later

Through-put A 384 -well plate taking about one night • 384 samples minus 16 controls = 368 • 15 markers per sample • makes 5520 genotypes (if succes rate 100%)
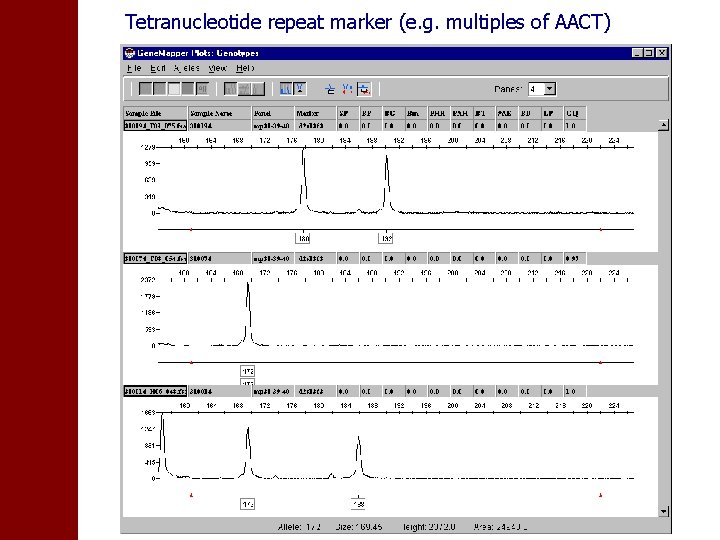
Tetranucleotide repeat marker (e. g. multiples of AACT)

• Detected length of PCR product depends on machine • Standards are used to correct this (CEPH DNA samples) • Take this into account when analysing data from different machines/labs
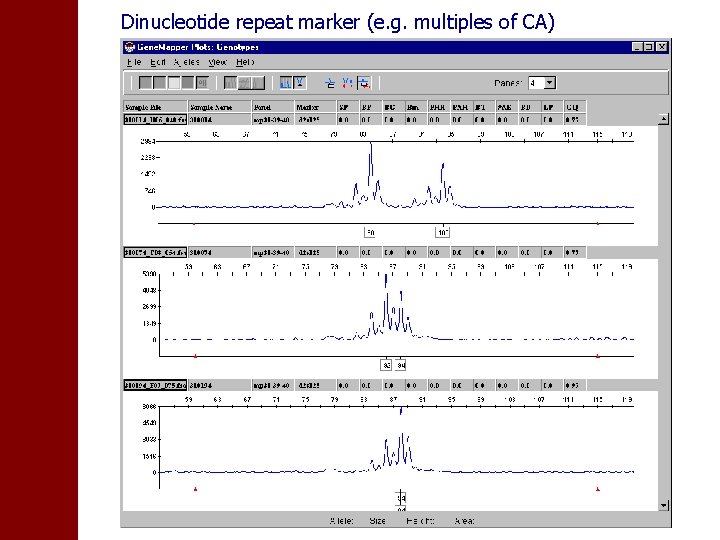
Dinucleotide repeat marker (e. g. multiples of CA)

• Dinucleotide repeats give less clean pictures but in practice this is no problem as long as pattern is always the same • However, markers not in standard 10 c. M screening sets often are more problematic (different stutter patterns for different samples, non-constant ratio ‘real peak’/plus-A peak) increased error rates?
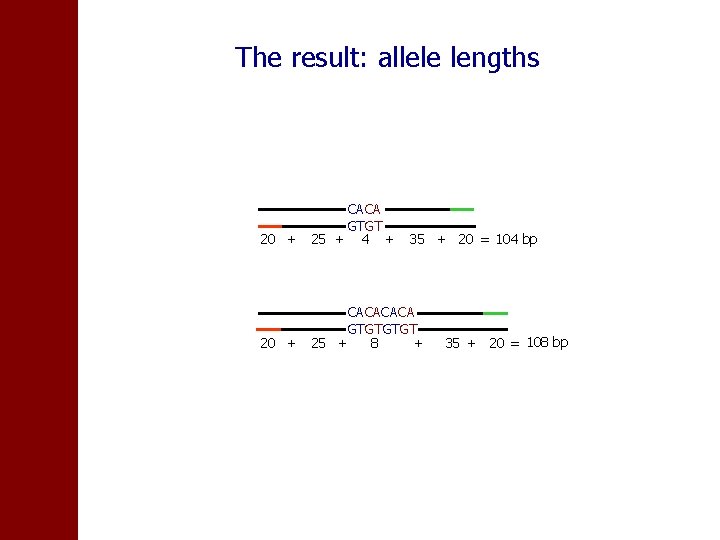
The result: allele lengths CACA GTGT 20 + 25 + 4 + 35 + 20 = 104 bp CACA GTGT 20 + 25 + 8 + 35 + 20 = 108 bp

Pedigree file in linkage format a er R 1 1 2 2 2 1 2 3 4 5 0 0 1 1 1 0 0 2 2 2 1 1 2 1 1 102 106 104 0 0 111 112 111 aw k ar t da m 104 110 106 110 0 0 118 114 ed er b um n Re 1 3 2 2 0 0 5 6 5 2 4 3 4 0 0 8 7 7 ta a d

Genetic map of measured markers For IBD estimation using Merlin or other software • Pedigree file • Genetic map

Markers measured on chromosome 19 16 markers d 19 s 247 d 19 s 1034 d 19 s 391 d 19 s 865 d 19 s 394 d 19 s 588 d 19 s 49 d 19 s 433 d 19 s 47 d 19 s 420 d 19 s 178 apoc 2 d 19 s 246 d 19 s 180 d 19 s 210 d 19 s 254

Genetic maps Available from • Marshfield Center for Medical Genetics http: //research. marshfieldclinic. org/genetics/ • Decode Genetics (most accurate) Supplemental data to Kong et al. Nat Genet 2002; 31: 241 -7. see F: BasGenotyping&MapsDecode. Map. xls

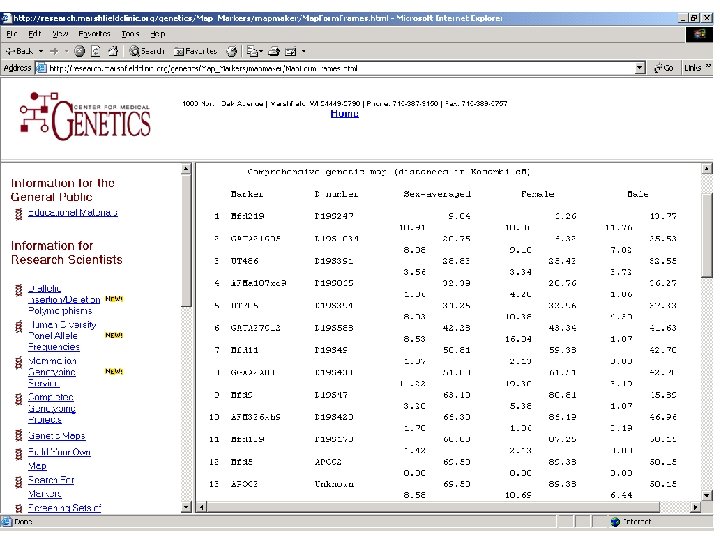
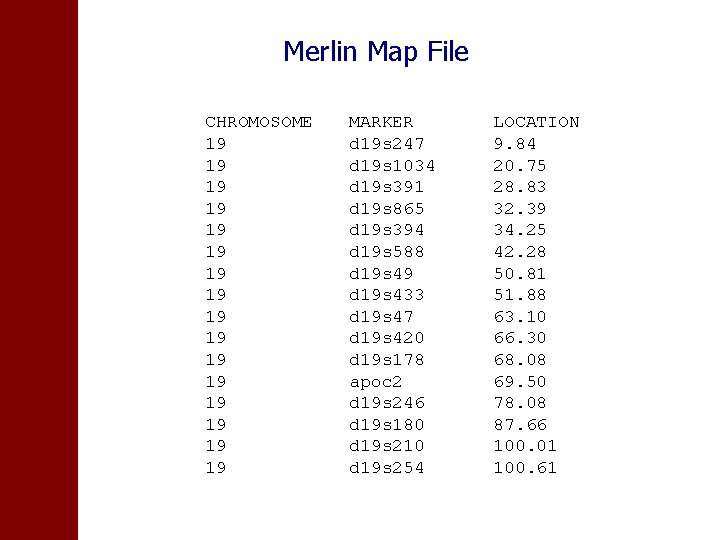
Merlin Map File CHROMOSOME 19 19 19 19 MARKER d 19 s 247 d 19 s 1034 d 19 s 391 d 19 s 865 d 19 s 394 d 19 s 588 d 19 s 49 d 19 s 433 d 19 s 47 d 19 s 420 d 19 s 178 apoc 2 d 19 s 246 d 19 s 180 d 19 s 210 d 19 s 254 LOCATION 9. 84 20. 75 28. 83 32. 39 34. 25 42. 28 50. 81 51. 88 63. 10 66. 30 68. 08 69. 50 78. 08 87. 66 100. 01 100. 61
- Slides: 38