Genomics of Gene Regulation ANSC 497 B Ross

Genomics of Gene Regulation ANSC 497 B Ross Hardison Nov. 10, 2009
DNA sequences involved in regulation of gene transcription Protein-DNA interactions Chromatin effects
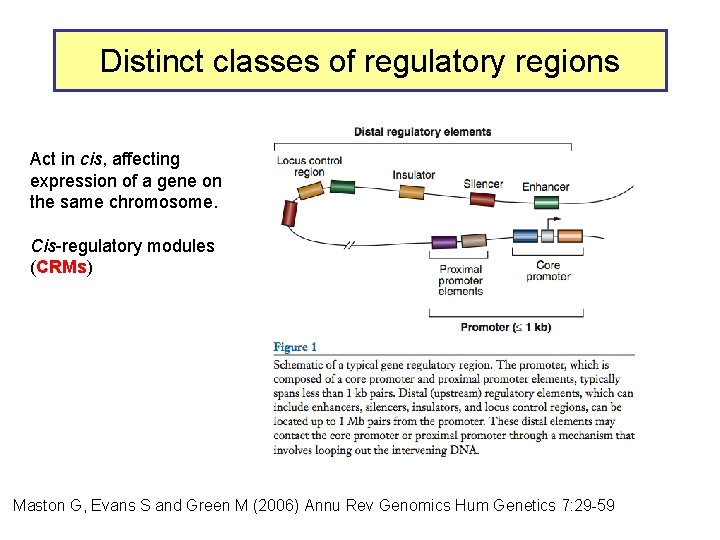
Distinct classes of regulatory regions Act in cis, affecting expression of a gene on the same chromosome. Cis-regulatory modules (CRMs) Maston G, Evans S and Green M (2006) Annu Rev Genomics Hum Genetics 7: 29 -59
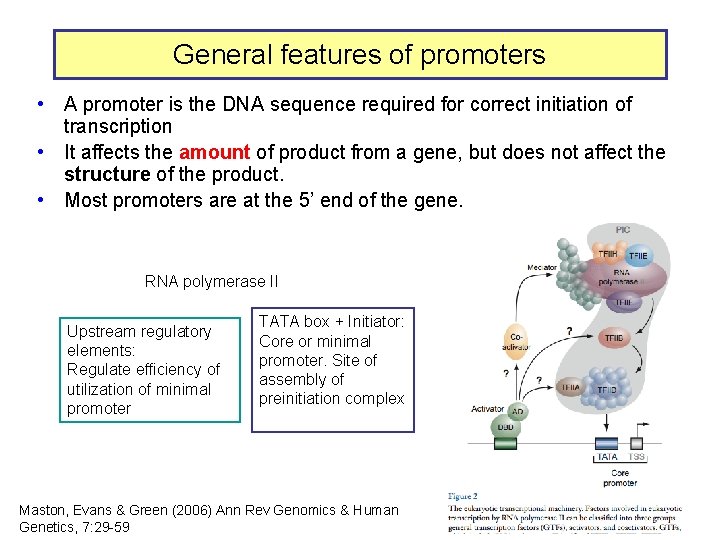
General features of promoters • A promoter is the DNA sequence required for correct initiation of transcription • It affects the amount of product from a gene, but does not affect the structure of the product. • Most promoters are at the 5’ end of the gene. RNA polymerase II Upstream regulatory elements: Regulate efficiency of utilization of minimal promoter TATA box + Initiator: Core or minimal promoter. Site of assembly of preinitiation complex Maston, Evans & Green (2006) Ann Rev Genomics & Human Genetics, 7: 29 -59

Conventional view of eukaryotic gene promoters Maston, Evans & Green (2006) Ann Rev Genomics & Human Genetics, 7: 29 -59
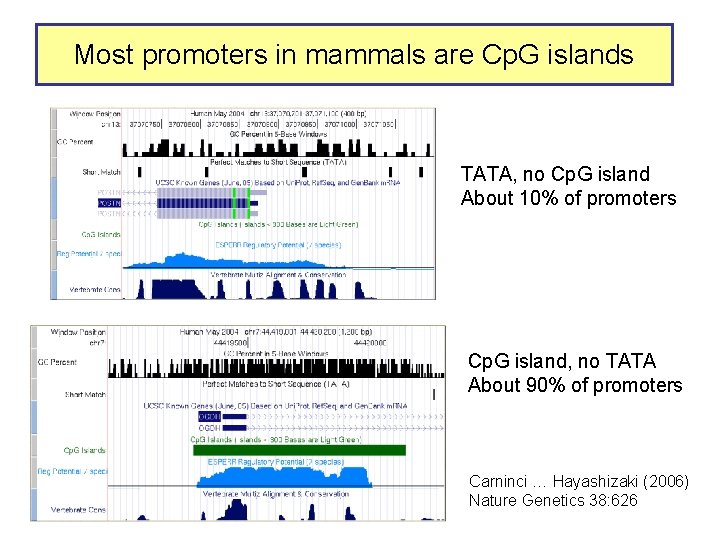
Most promoters in mammals are Cp. G islands TATA, no Cp. G island About 10% of promoters Cp. G island, no TATA About 90% of promoters Carninci … Hayashizaki (2006) Nature Genetics 38: 626
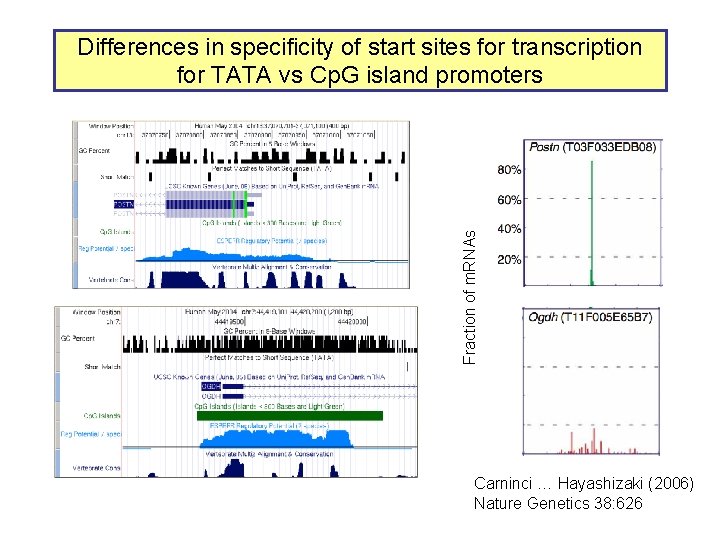
Fraction of m. RNAs Differences in specificity of start sites for transcription for TATA vs Cp. G island promoters Carninci … Hayashizaki (2006) Nature Genetics 38: 626
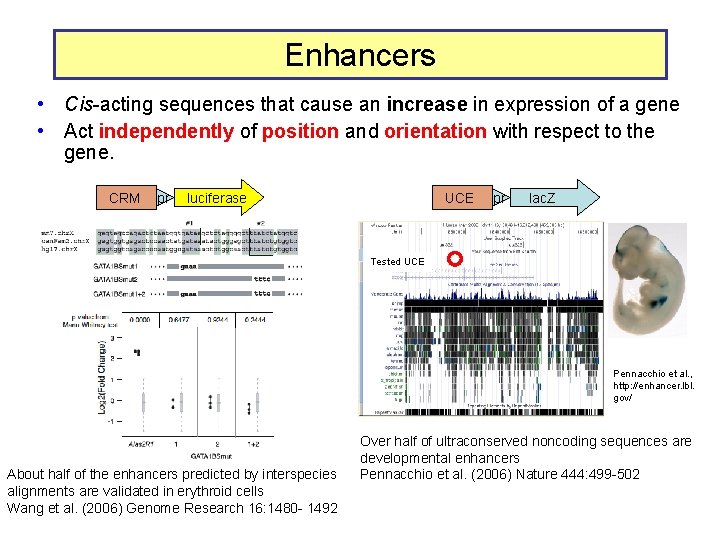
Enhancers • Cis-acting sequences that cause an increase in expression of a gene • Act independently of position and orientation with respect to the gene. CRM pr luciferase UCE pr lac. Z Tested UCE Pennacchio et al. , http: //enhancer. lbl. gov/ About half of the enhancers predicted by interspecies alignments are validated in erythroid cells Wang et al. (2006) Genome Research 16: 1480 - 1492 Over half of ultraconserved noncoding sequences are developmental enhancers Pennacchio et al. (2006) Nature 444: 499 -502
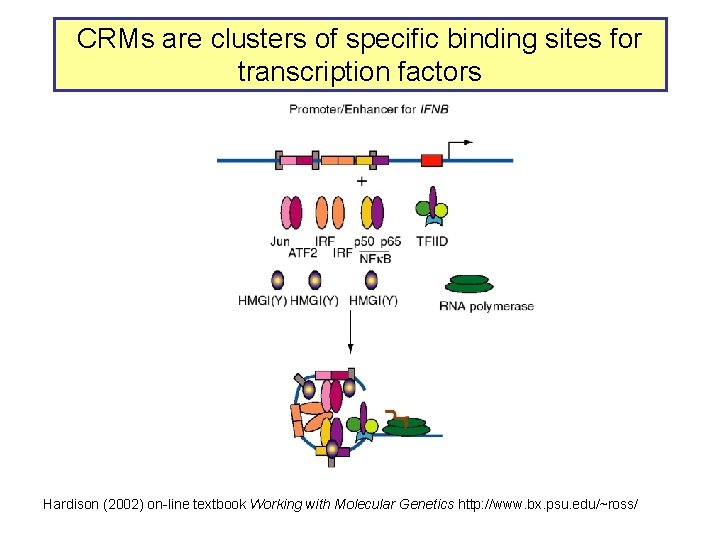
CRMs are clusters of specific binding sites for transcription factors Hardison (2002) on-line textbook Working with Molecular Genetics http: //www. bx. psu. edu/~ross/
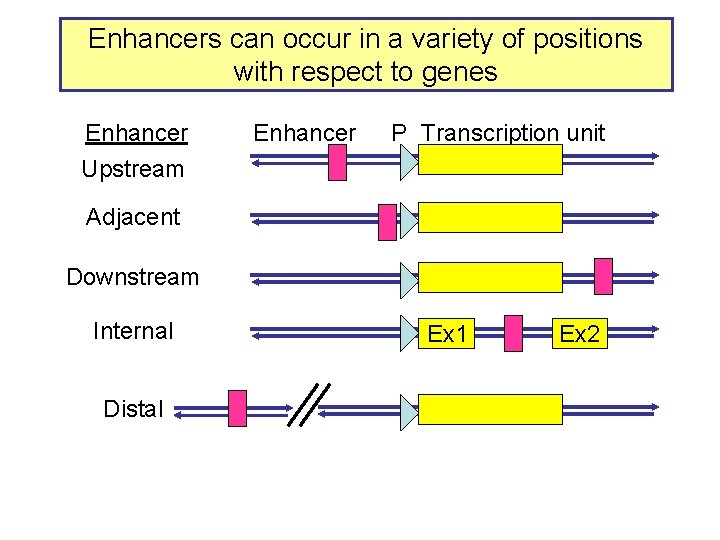
Enhancers can occur in a variety of positions with respect to genes Enhancer P Transcription unit Upstream Adjacent Downstream Internal Distal Ex 1 Ex 2
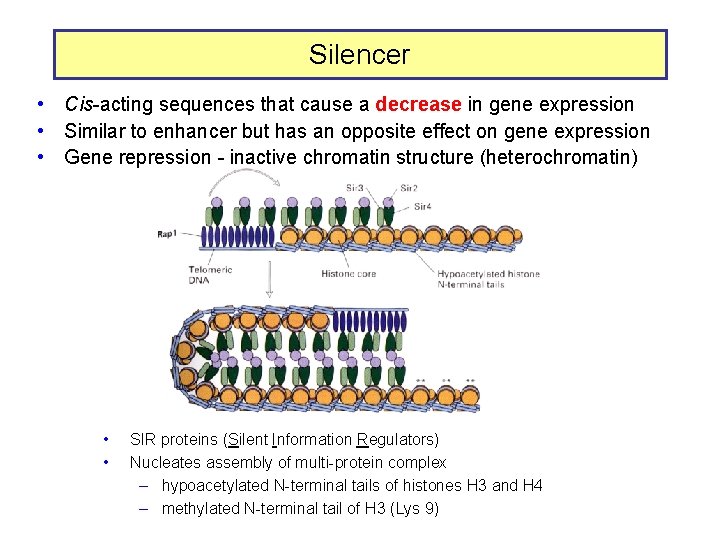
Silencer • Cis-acting sequences that cause a decrease in gene expression • Similar to enhancer but has an opposite effect on gene expression • Gene repression - inactive chromatin structure (heterochromatin) • • SIR proteins (Silent Information Regulators) Nucleates assembly of multi-protein complex – hypoacetylated N-terminal tails of histones H 3 and H 4 – methylated N-terminal tail of H 3 (Lys 9)
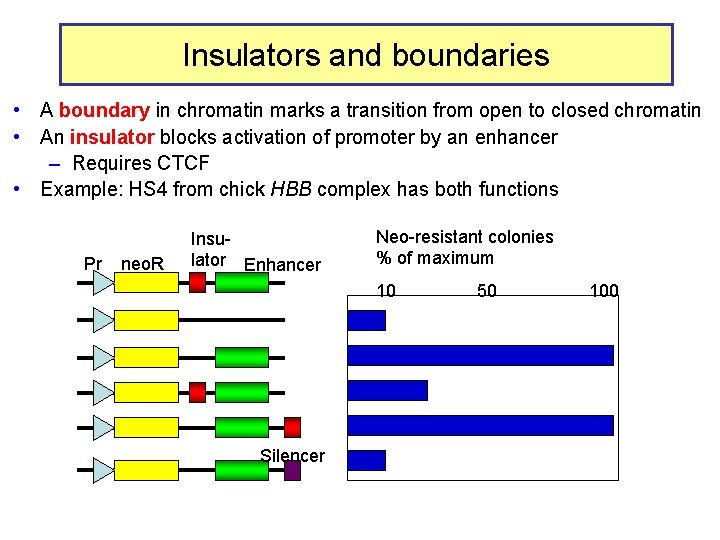
Insulators and boundaries • A boundary in chromatin marks a transition from open to closed chromatin • An insulator blocks activation of promoter by an enhancer – Requires CTCF • Example: HS 4 from chick HBB complex has both functions Pr neo. R Insulator Enhancer Neo-resistant colonies % of maximum 10 Silencer 50 100
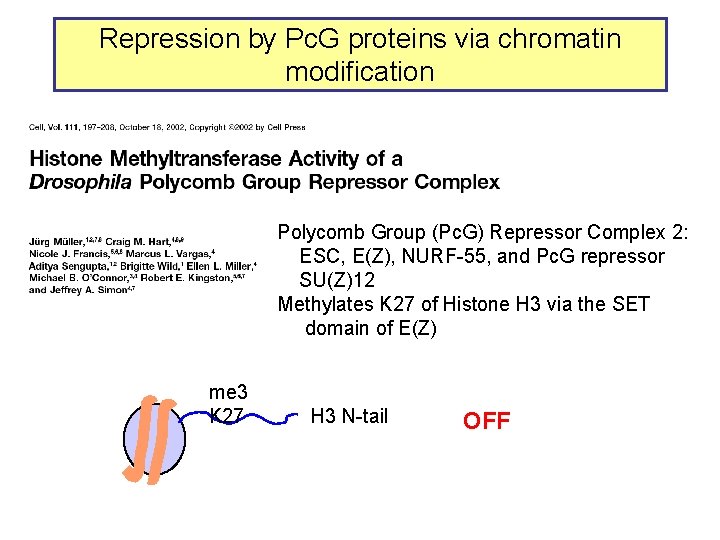
Repression by Pc. G proteins via chromatin modification Polycomb Group (Pc. G) Repressor Complex 2: ESC, E(Z), NURF-55, and Pc. G repressor SU(Z)12 Methylates K 27 of Histone H 3 via the SET domain of E(Z) me 3 K 27 H 3 N-tail OFF
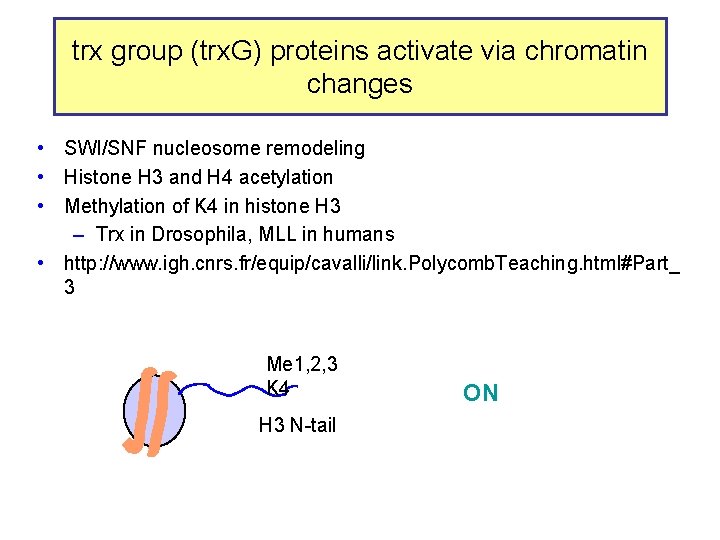
trx group (trx. G) proteins activate via chromatin changes • SWI/SNF nucleosome remodeling • Histone H 3 and H 4 acetylation • Methylation of K 4 in histone H 3 – Trx in Drosophila, MLL in humans • http: //www. igh. cnrs. fr/equip/cavalli/link. Polycomb. Teaching. html#Part_ 3 Me 1, 2, 3 K 4 H 3 N-tail ON
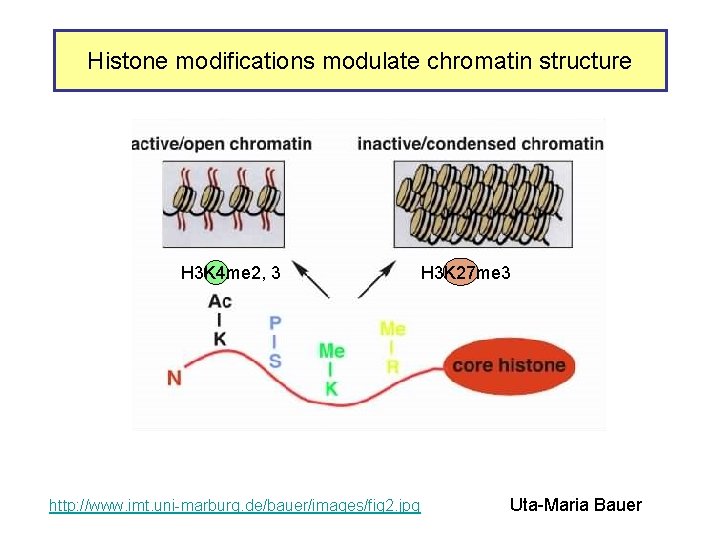
Histone modifications modulate chromatin structure H 3 K 4 me 2, 3 http: //www. imt. uni-marburg. de/bauer/images/fig 2. jpg H 3 K 27 me 3 Uta-Maria Bauer
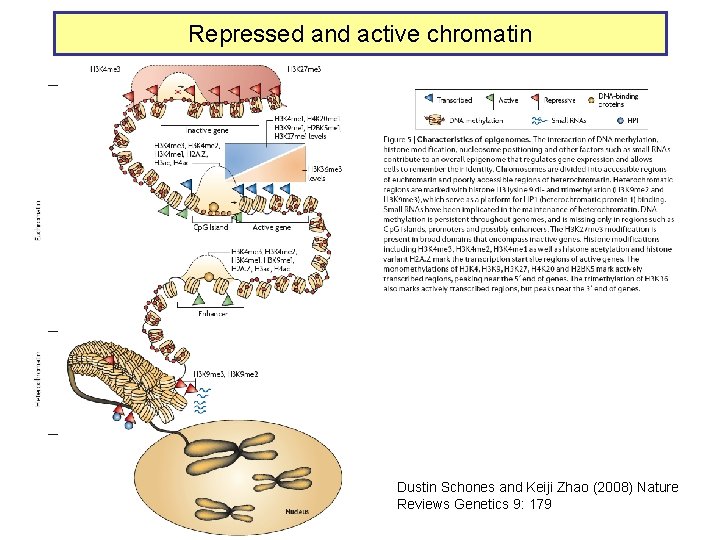
Repressed and active chromatin Dustin Schones and Keiji Zhao (2008) Nature Reviews Genetics 9: 179
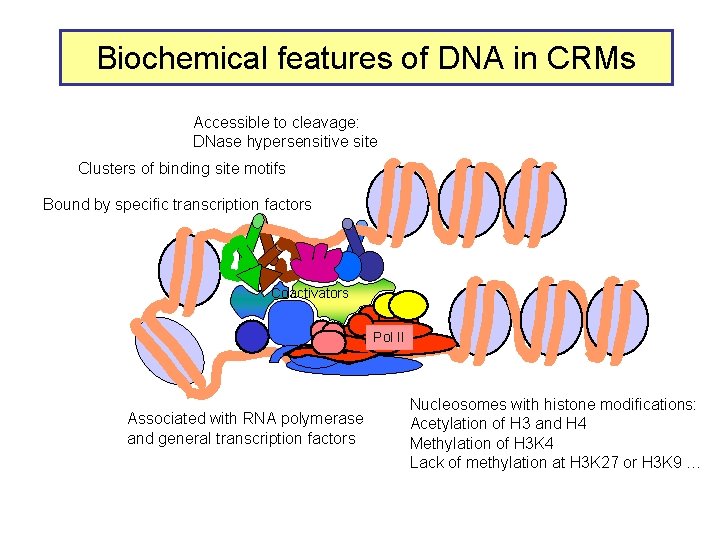
Biochemical features of DNA in CRMs Accessible to cleavage: DNase hypersensitive site Clusters of binding site motifs Bound by specific transcription factors Coactivators Pol IIa II Associated with RNA polymerase and general transcription factors Nucleosomes with histone modifications: Acetylation of H 3 and H 4 Methylation of H 3 K 4 Lack of methylation at H 3 K 27 or H 3 K 9 …

Methods in Genomics of Gene Regulation
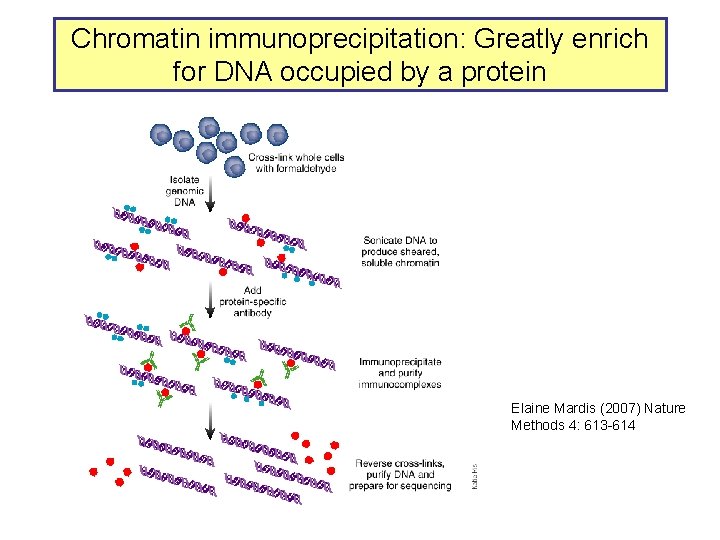
Chromatin immunoprecipitation: Greatly enrich for DNA occupied by a protein Elaine Mardis (2007) Nature Methods 4: 613 -614
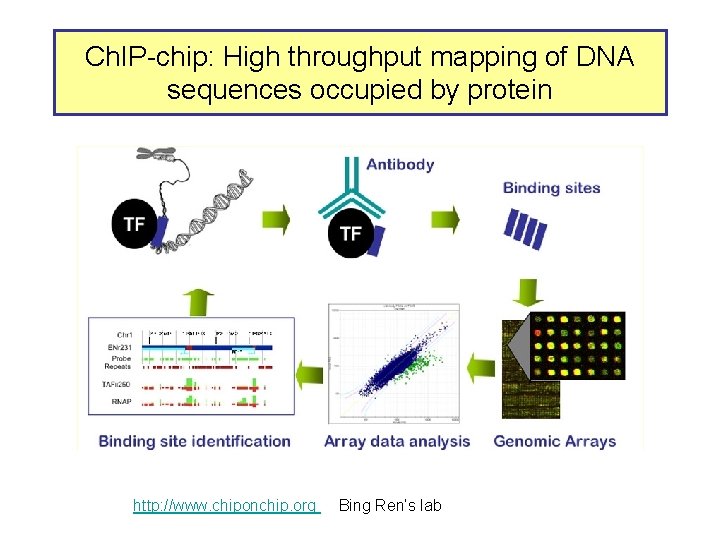
Ch. IP-chip: High throughput mapping of DNA sequences occupied by protein http: //www. chiponchip. org Bing Ren’s lab
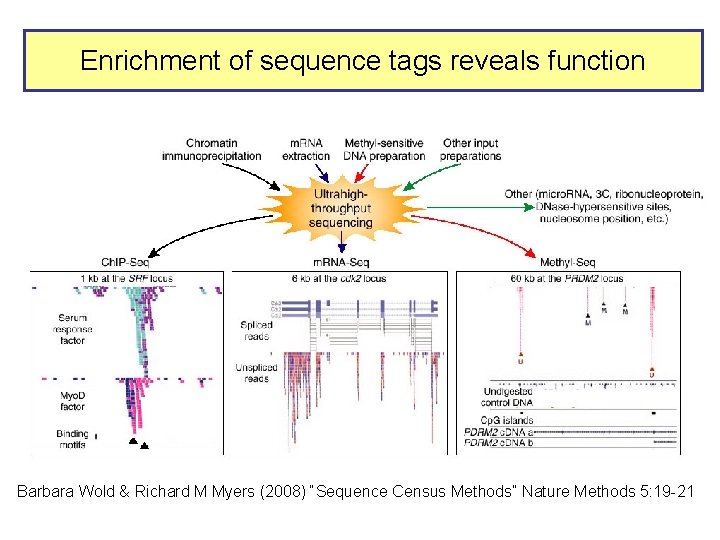
Enrichment of sequence tags reveals function Barbara Wold & Richard M Myers (2008) “Sequence Census Methods” Nature Methods 5: 19 -21
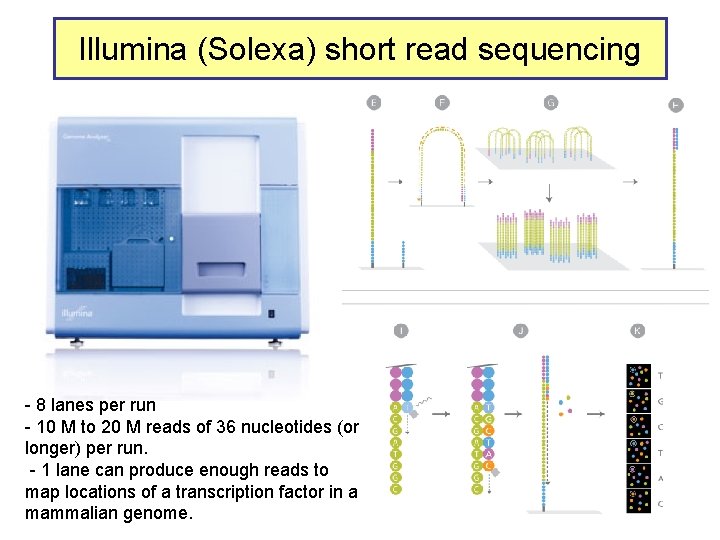
Illumina (Solexa) short read sequencing - 8 lanes per run - 10 M to 20 M reads of 36 nucleotides (or longer) per run. - 1 lane can produce enough reads to map locations of a transcription factor in a mammalian genome.
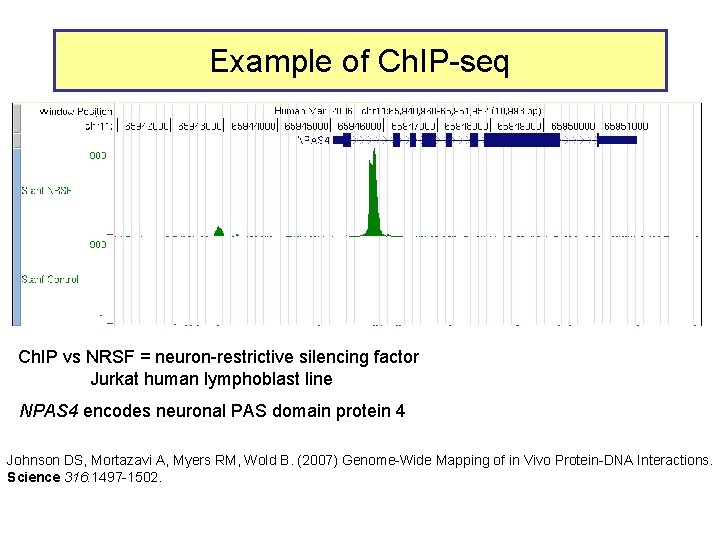
Example of Ch. IP-seq Ch. IP vs NRSF = neuron-restrictive silencing factor Jurkat human lymphoblast line NPAS 4 encodes neuronal PAS domain protein 4 Johnson DS, Mortazavi A, Myers RM, Wold B. (2007) Genome-Wide Mapping of in Vivo Protein-DNA Interactions. Science 316: 1497 -1502.
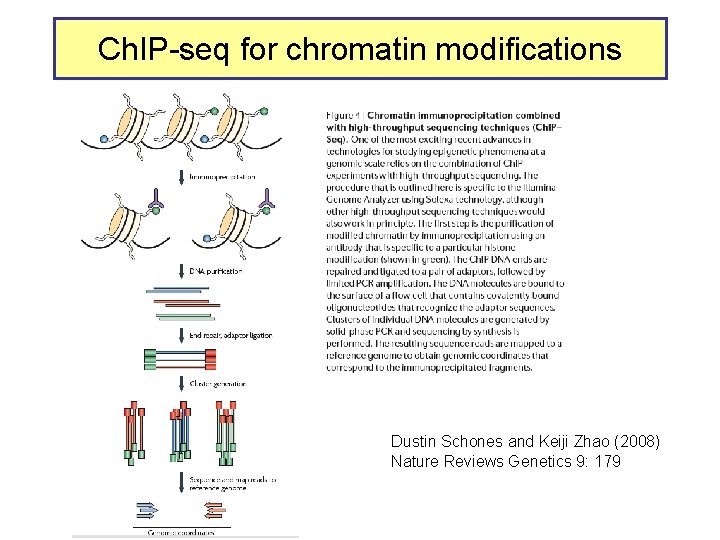
Ch. IP-seq for chromatin modifications Dustin Schones and Keiji Zhao (2008) Nature Reviews Genetics 9: 179
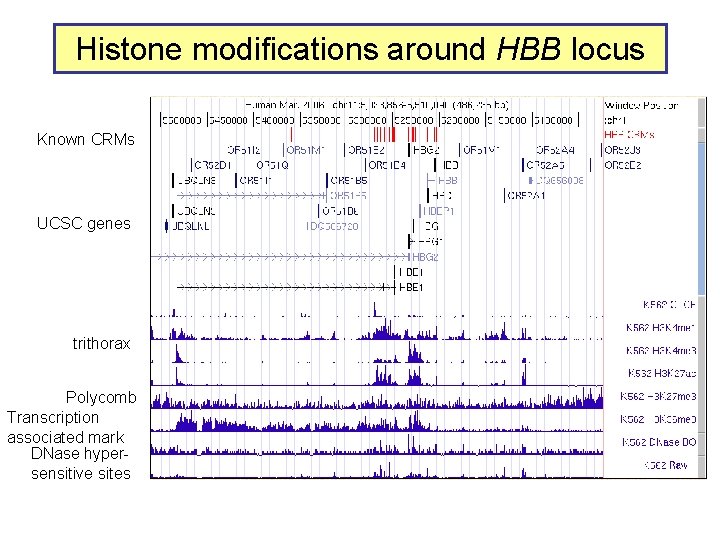
Histone modifications around HBB locus Known CRMs UCSC genes trithorax Polycomb Transcription associated mark DNase hypersensitive sites
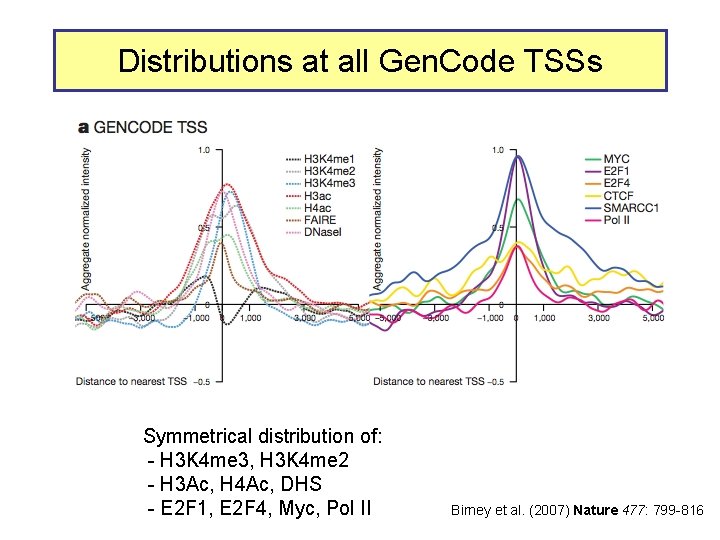
Distributions at all Gen. Code TSSs Symmetrical distribution of: - H 3 K 4 me 3, H 3 K 4 me 2 - H 3 Ac, H 4 Ac, DHS - E 2 F 1, E 2 F 4, Myc, Pol II Birney et al. (2007) Nature 477: 799 -816
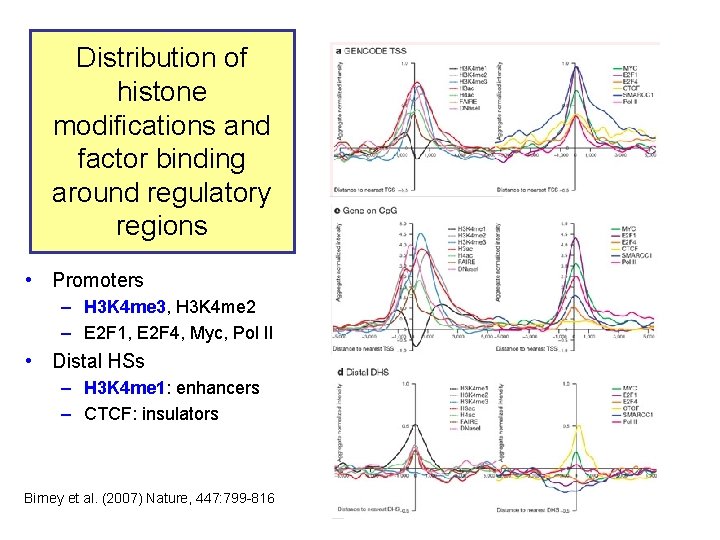
Distribution of histone modifications and factor binding around regulatory regions • Promoters – H 3 K 4 me 3, H 3 K 4 me 2 – E 2 F 1, E 2 F 4, Myc, Pol II • Distal HSs – H 3 K 4 me 1: enhancers – CTCF: insulators Birney et al. (2007) Nature, 447: 799 -816

Enhancers predicted from chromatin signatures (2009) Nature 459: 108 -112
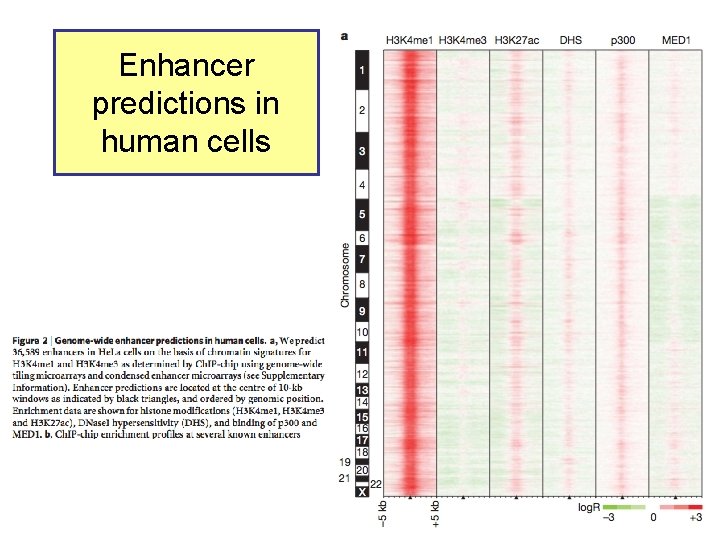
Enhancer predictions in human cells
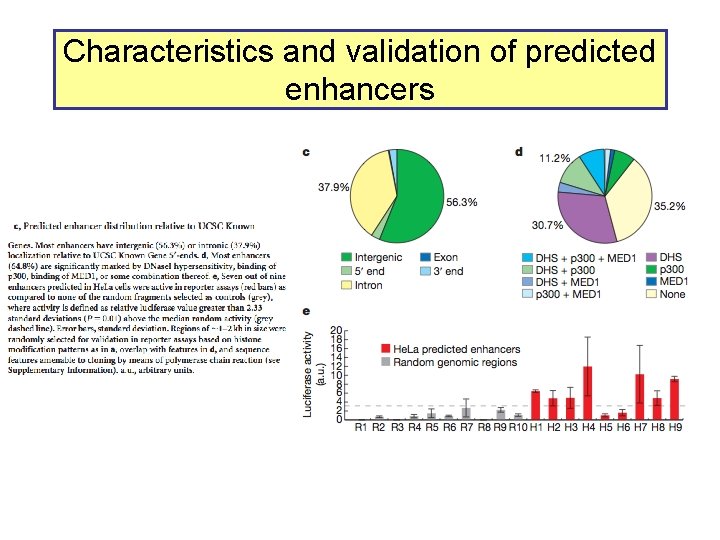
Characteristics and validation of predicted enhancers

Data Resources for Genomics of Gene Regulation

UCSC Genome Browser • Visualize data described in publications, e. g. – Expression data • Affymetrix gene arrays, GNF, Su et al. 2004 – Regulation • • • Kim et al. 2005, PICs (TAF 1) Kim et al. , 2008, CTCF Boyle et al. , 2008, DNase hypersensitive sites Heintzman et al. , 2009, Enhancers predicted by H 3 K 4 me 1 Mikkelsen et al. , 2007, Chromatin modifications in pluripotent and lineage-committed cells ENCODE project, Production phase – Expression • • Affy high density tiling arrays RNA-seq from several sources (CSHL, Helicos) – Regulation • • Broad histone modifications HAIB DNA methylation Open Chromatin UW DNase HS HAIB TFBS Yale TFBS SUNY RBP
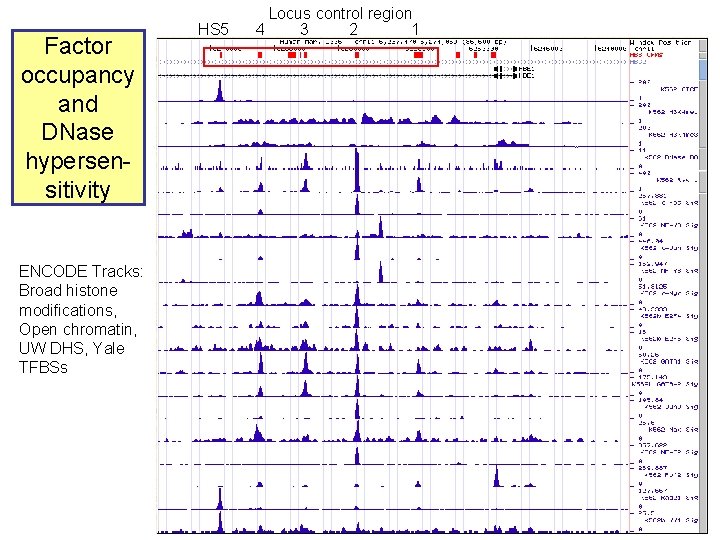
Factor occupancy and DNase hypersensitivity ENCODE Tracks: Broad histone modifications, Open chromatin, UW DHS, Yale TFBSs HS 5 Locus control region 4 3 2 1

Collated sets of published regulatory regions • http: //www. bx. psu. edu/~ross/dataset/Reguldata. html • Noncoding DNA segments with high regulatory potential • PRPs: Intersection of the High RP segments and the PRe. Mods (clusters of conserved transcription factor binding site motifs) • Most constrained DNA segments, phast. Cons • DNase hypersensitive sites in CD 4+ T cells • DNA segments occupied by CTCF in primary fibroblasts • Preinitiation complexes (TAF 1) in IMR 90 cells • Predicted erythroid cis-regulatory modules

Gene. Track • Genomic data analysis and integration – Istvan Albert, Frank Pugh, et al. , PSU – http: //genetrack. bx. psu. edu/ • Install on your system • Gallery of data for visualization – Yeast H 2 AZ nucleosome predictions, 454 sequencing – Drosophila H 2 AZ nucleosome predictions, 454 sequencing
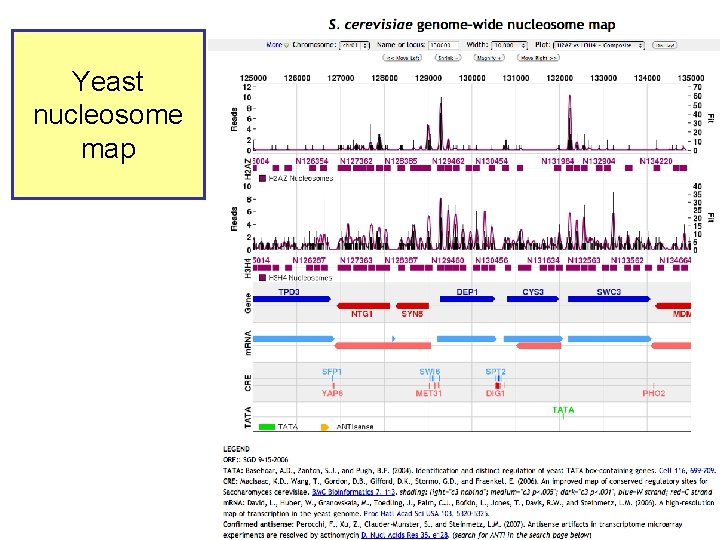
Yeast nucleosome map
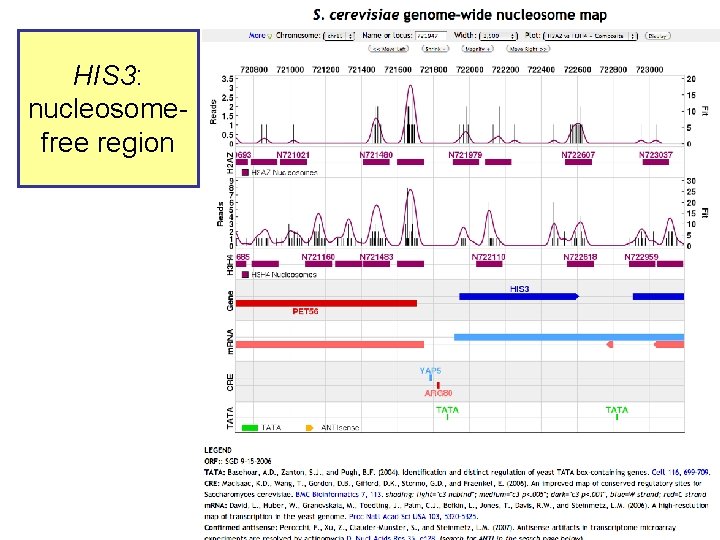
HIS 3: nucleosomefree region

mod. ENCODE http: //www. modencode. org/ Worm and Fly Gene annotations Expression Chromatin modifications TFBs in vivo, etc.

Experimental Tests in the Genomics of Gene Regulation
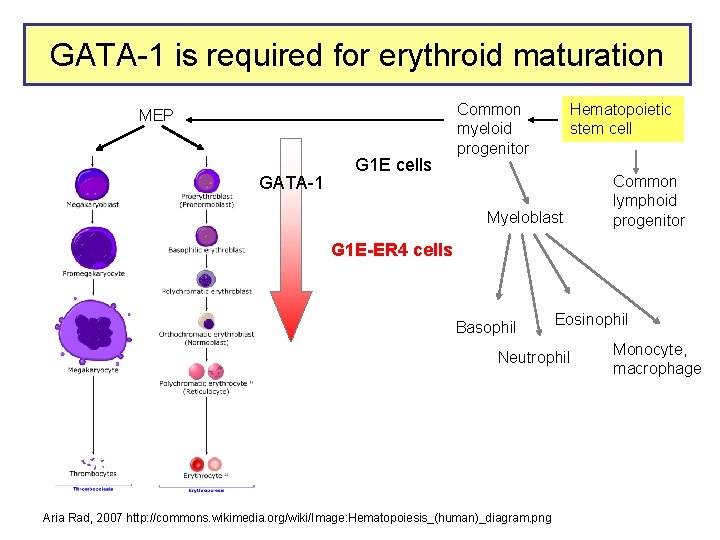
GATA-1 is required for erythroid maturation MEP GATA-1 G 1 E cells Common myeloid progenitor Hematopoietic stem cell Myeloblast Common lymphoid progenitor G 1 E-ER 4 cells Basophil Eosinophil Neutrophil Aria Rad, 2007 http: //commons. wikimedia. org/wiki/Image: Hematopoiesis_(human)_diagram. png Monocyte, macrophage
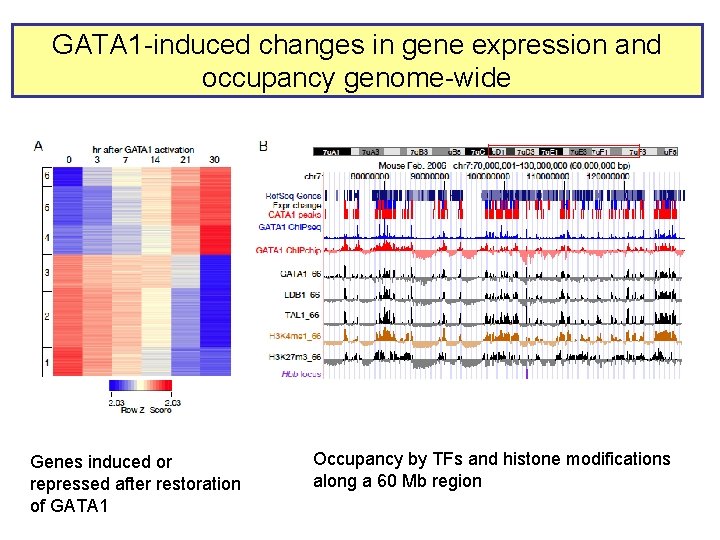
GATA 1 -induced changes in gene expression and occupancy genome-wide Genes induced or repressed after restoration of GATA 1 Occupancy by TFs and histone modifications along a 60 Mb region
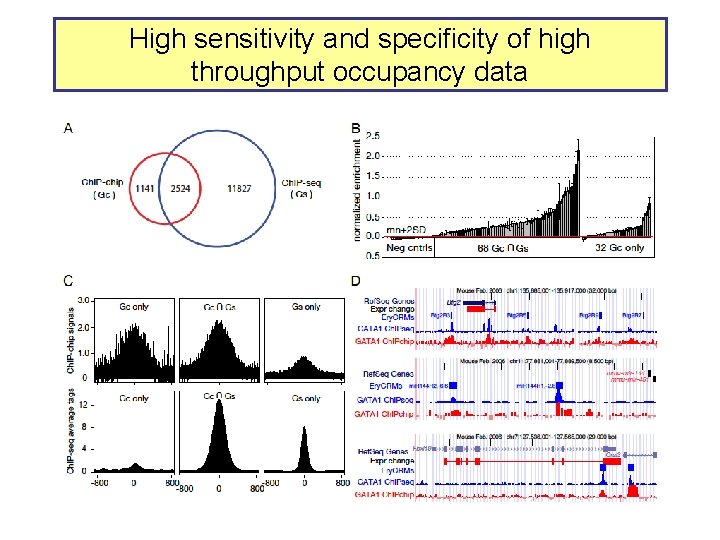
High sensitivity and specificity of high throughput occupancy data
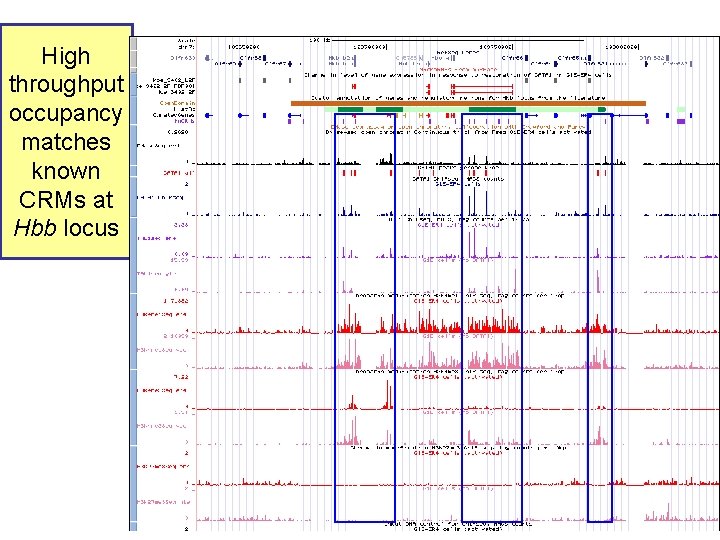
High throughput occupancy matches known CRMs at Hbb locus
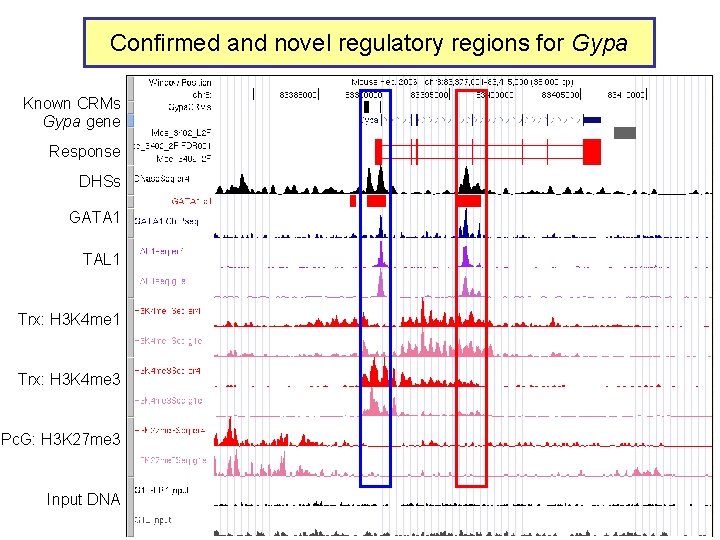
Confirmed and novel regulatory regions for Gypa Known CRMs Gypa gene Response DHSs GATA 1 TAL 1 Trx: H 3 K 4 me 3 Pc. G: H 3 K 27 me 3 Input DNA
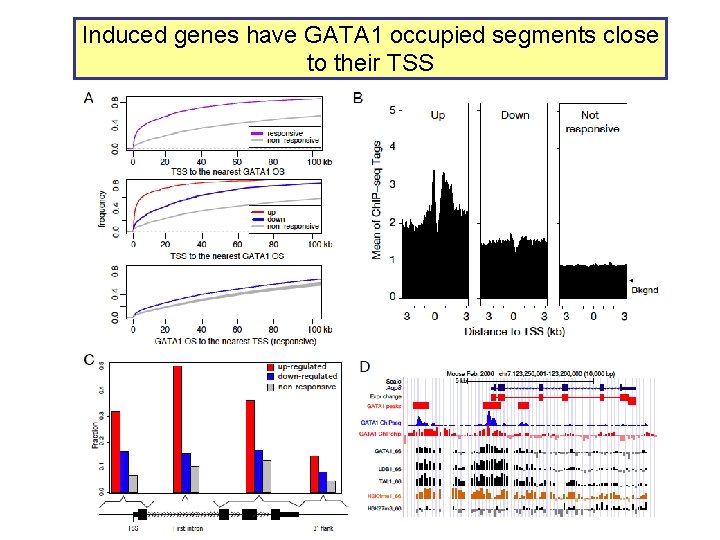
Induced genes have GATA 1 occupied segments close to their TSS
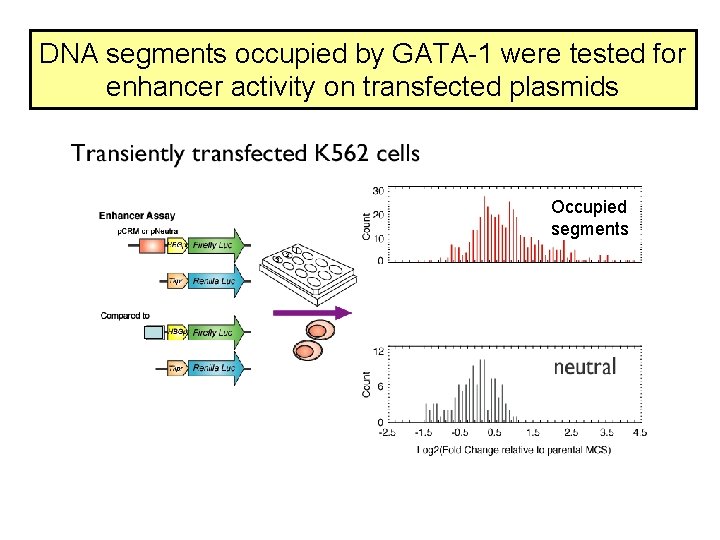
DNA segments occupied by GATA-1 were tested for enhancer activity on transfected plasmids Occupied segments
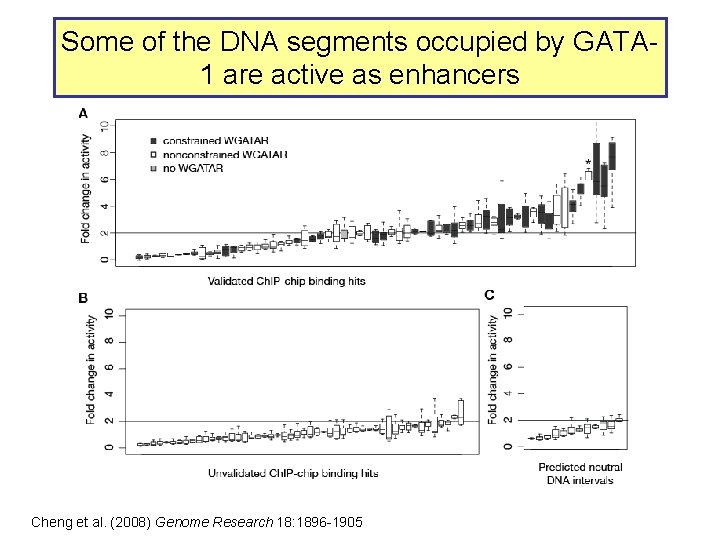
Some of the DNA segments occupied by GATA 1 are active as enhancers Cheng et al. (2008) Genome Research 18: 1896 -1905
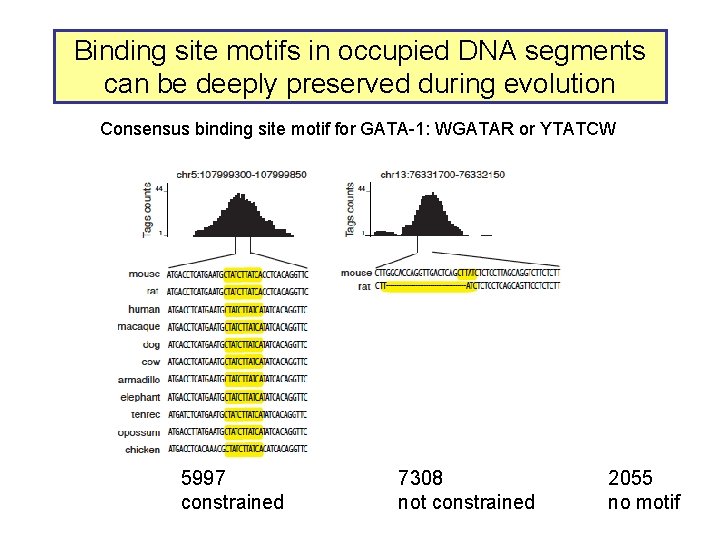
Binding site motifs in occupied DNA segments can be deeply preserved during evolution Consensus binding site motif for GATA-1: WGATAR or YTATCW 5997 constrained 7308 not constrained 2055 no motif
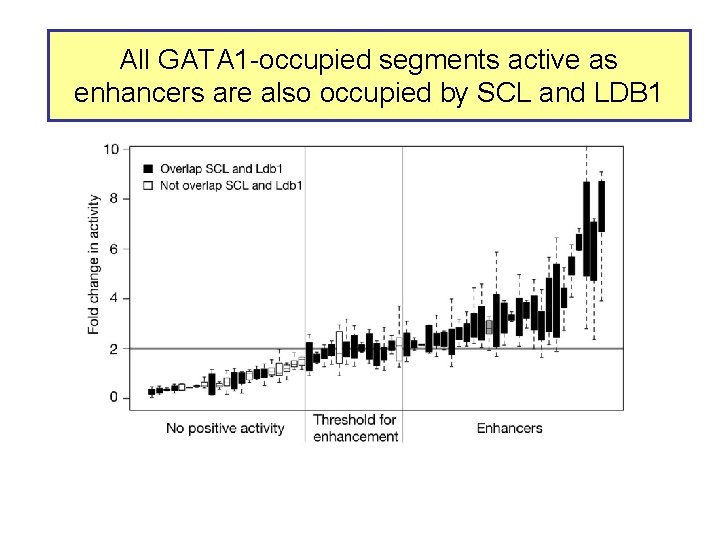
All GATA 1 -occupied segments active as enhancers are also occupied by SCL and LDB 1

Genetic Determinants of Variation in Gene Expression

Variation of gene expression among individuals • Levels of expression of many genes vary in humans (and other species) • Variation in expression is heritable • Determinants of variability map to discrete genomic intervals • Often multiple determinants • This variation indicates an abundance of cis-regulatory variation in the human genome • For example: – Microarray expression analyses of 3554 genes in 14 families • Morley M … Cheung VG (2004) Nature 430: 743 -747 - Expression analysis of about 16 Hap. Map individuals • Storey et al. (2007) AJHG 80: 502 -509 – Expression analysis of all 270 individuals genotypes in Hap. Map • Stranger BE … Dermitzakis E (2007) Nature Genetics 39: 1217 -1224
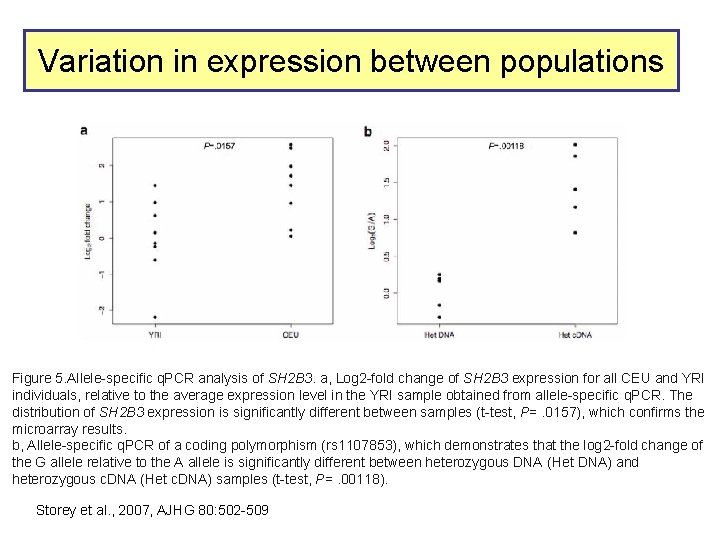
Variation in expression between populations Figure 5. Allele-specific q. PCR analysis of SH 2 B 3. a, Log 2 -fold change of SH 2 B 3 expression for all CEU and YRI individuals, relative to the average expression level in the YRI sample obtained from allele-specific q. PCR. The distribution of SH 2 B 3 expression is significantly different between samples (t-test, P=. 0157), which confirms the microarray results. b, Allele-specific q. PCR of a coding polymorphism (rs 1107853), which demonstrates that the log 2 -fold change of the G allele relative to the A allele is significantly different between heterozygous DNA (Het DNA) and heterozygous c. DNA (Het c. DNA) samples (t-test, P=. 00118). Storey et al. , 2007, AJHG 80: 502 -509

Mapping determinants of expression variation • • Stranger et al. , 2007, Nature Genetics 39: 1217 -1224 Expression analysis of EBV-transformed lymphoblastoid cells from all 270 individuals genotypes in Hap. Map – – • 30 Caucasian trios (90) of European descent in Utah (CEU) 30 Yoruba trios (90) from Ibadan, Nigeria (YRI) 45 unrelated Chinese individuals from Beijing Univ (CHB) 45 unrelated Japanese individuals from Tokyo (JPT) Measure levels of expression of 47, 294 probes (about 24, 000 genes) in each individual – Focus on 13, 643 genes “selected on criteria of variance and population differentiation” • • Already know genotypes at about 2. 2 million SNPs for each individual (Hap. Map) Test for significant association of variation at each SNP with variation in expression of each gene – Linear regression model – Spearman rank correlation test • Evaluate significance of regression P values by 10, 000 permutations of the data, focus on those associations above the 0. 001 permutation threshold
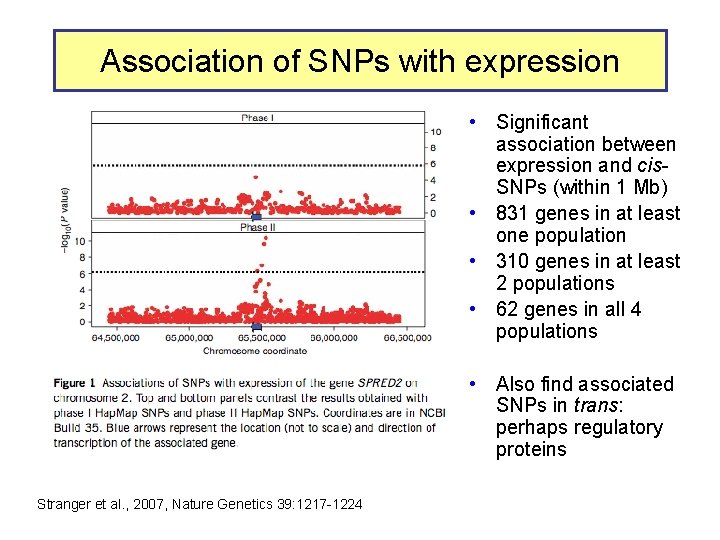
Association of SNPs with expression • Significant association between expression and cis. SNPs (within 1 Mb) • 831 genes in at least one population • 310 genes in at least 2 populations • 62 genes in all 4 populations • Also find associated SNPs in trans: perhaps regulatory proteins Stranger et al. , 2007, Nature Genetics 39: 1217 -1224
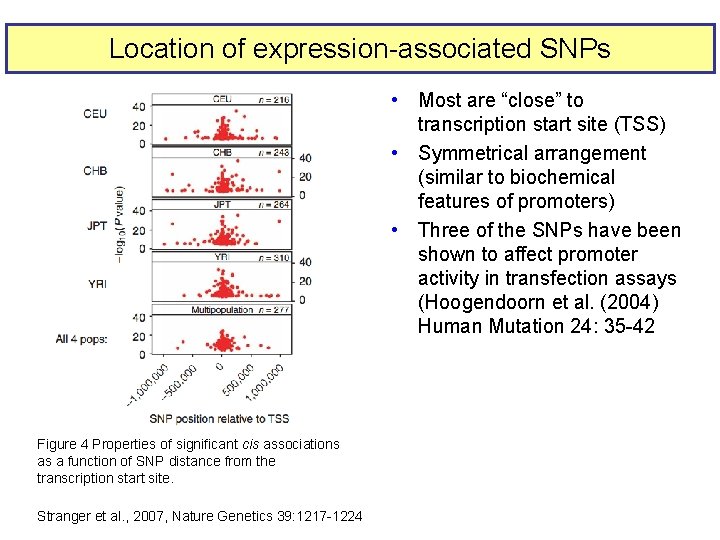
Location of expression-associated SNPs • Most are “close” to transcription start site (TSS) • Symmetrical arrangement (similar to biochemical features of promoters) • Three of the SNPs have been shown to affect promoter activity in transfection assays (Hoogendoorn et al. (2004) Human Mutation 24: 35 -42 Figure 4 Properties of significant cis associations as a function of SNP distance from the transcription start site. Stranger et al. , 2007, Nature Genetics 39: 1217 -1224

Relevance to human health • "We predict that variants in regulatory regions make a greater contribution to complex disease than do variants that affect protein sequence” – Manolis Dermitzakis, Science. Daily
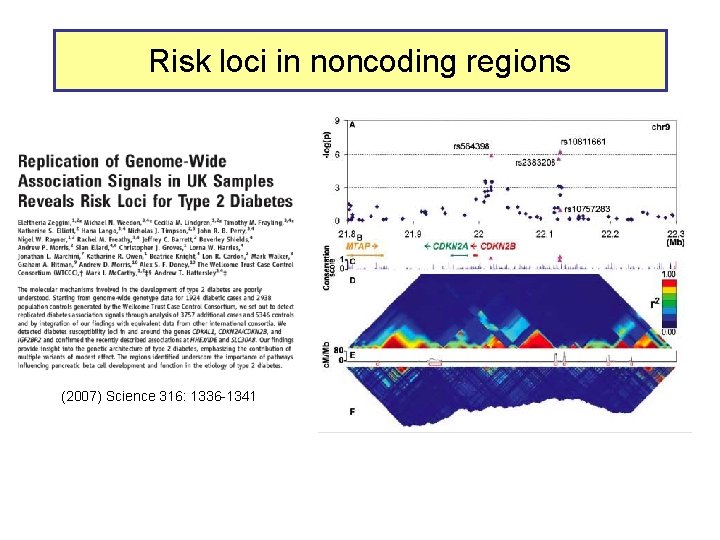
Risk loci in noncoding regions (2007) Science 316: 1336 -1341
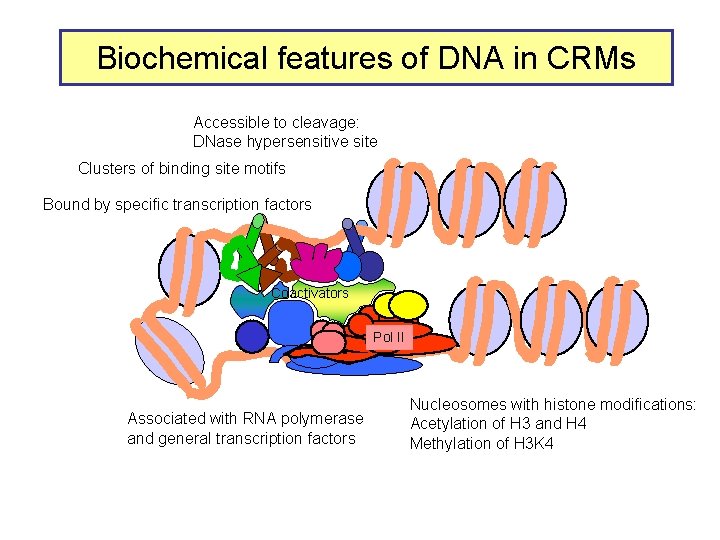
Biochemical features of DNA in CRMs Accessible to cleavage: DNase hypersensitive site Clusters of binding site motifs Bound by specific transcription factors Coactivators Pol IIa II Associated with RNA polymerase and general transcription factors Nucleosomes with histone modifications: Acetylation of H 3 and H 4 Methylation of H 3 K 4
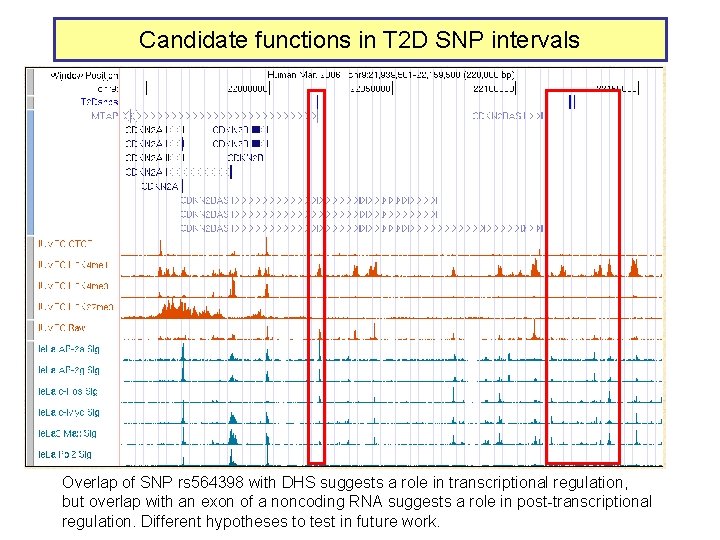
Candidate functions in T 2 D SNP intervals Overlap of SNP rs 564398 with DHS suggests a role in transcriptional regulation, but overlap with an exon of a noncoding RNA suggests a role in post-transcriptional regulation. Different hypotheses to test in future work.
- Slides: 59