Genomics of Ferns and Lycophytes Chapter 6 Structure



















- Slides: 34

Genomics of Ferns and Lycophytes

Chapter 6: Structure and evolution of fern plastid genomes Paul G. Wolf and Jessie M. Roper

Question: What is the inheritance of the chloroplast genome in ferns? Marchantia cp genome • ca. 150 kb, circular molecule • large and small single copy regions separated by inverted repeat • gene number and order +/- conserved across land plants

Generally in land plants: maternal (via the egg, excluded via sperm) • maternal with some biparental in Angiosperms • paternal in Gymnosperms • ferns? Phyllitis (Aspleniaceae) – biparental Osmunda (Osmundaceae) – maternal Polystichum (Drypoteridaceae) – maternal Pteridium Dennstaedtiaceae) – maternal Pellaea (Pteridaceae) – maternal “During insemination in Ceratopteris richardii [Pteridaceae], the sperm cytoskeleton and flagella rearrange, and the coils of the cell extend while entering the neck canal. . All cellular components, except plastids, enter the egg cytoplasm” Lopez-Smith and Renzagalia, 2008 (Sexual Plant Reproduction)

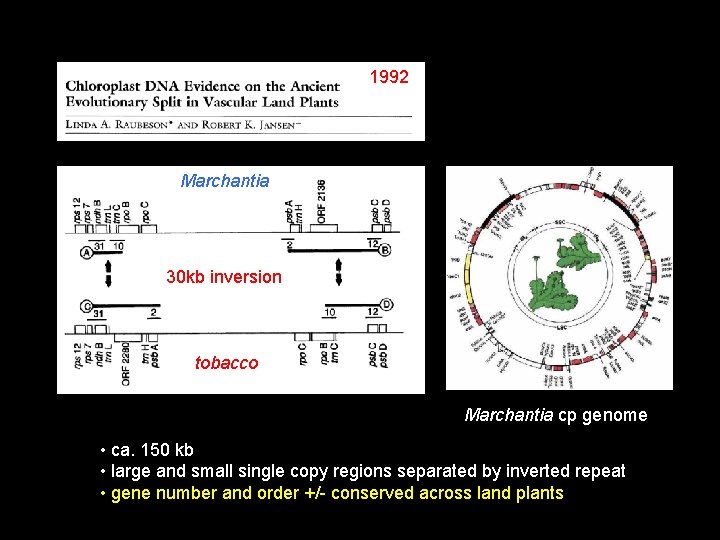
1992 Marchantia 30 kb inversion tobacco Marchantia cp genome • ca. 150 kb • large and small single copy regions separated by inverted repeat • gene number and order +/- conserved across land plants

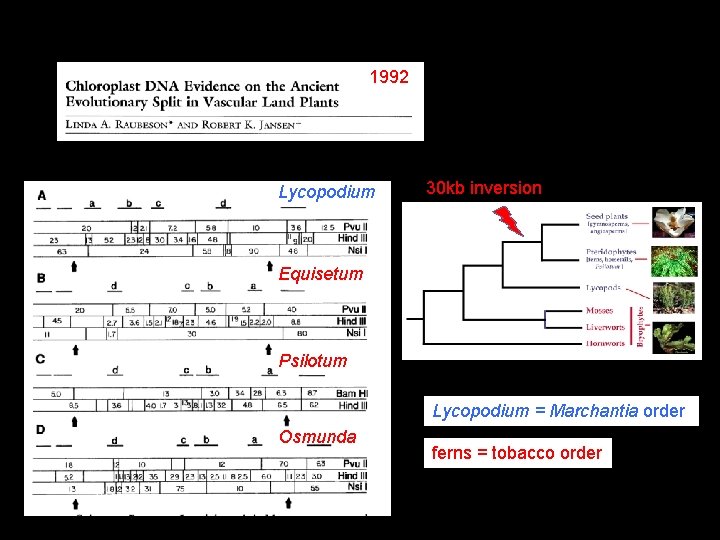
1992 Lycopodium 30 kb inversion Equisetum Psilotum Lycopodium = Marchantia order Osmunda ferns = tobacco order

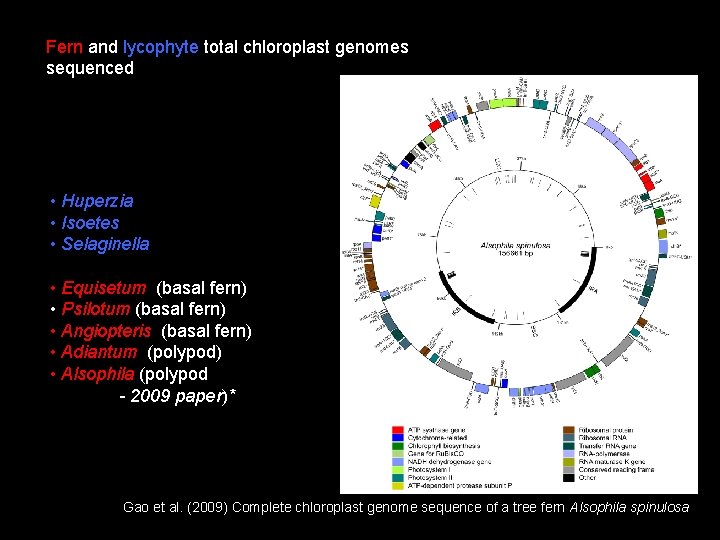
Fern and lycophyte total chloroplast genomes sequenced • Huperzia • Isoetes • Selaginella • Equisetum (basal fern) • Psilotum (basal fern) • Angiopteris (basal fern) • Adiantum (polypod) • Alsophila (polypod - 2009 paper)* Gao et al. (2009) Complete chloroplast genome sequence of a tree fern Alsophila spinulosa

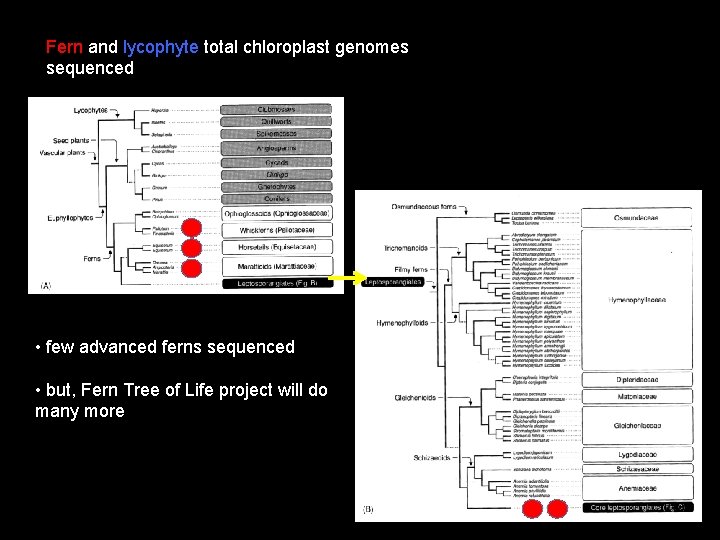
Fern and lycophyte total chloroplast genomes sequenced • few advanced ferns sequenced • but, Fern Tree of Life project will do many more

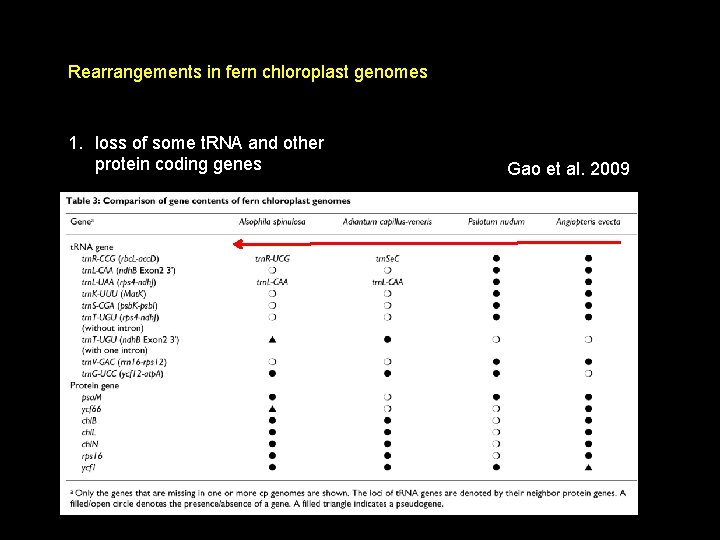
Rearrangements in fern chloroplast genomes 1. loss of some t. RNA and other protein coding genes Gao et al. 2009

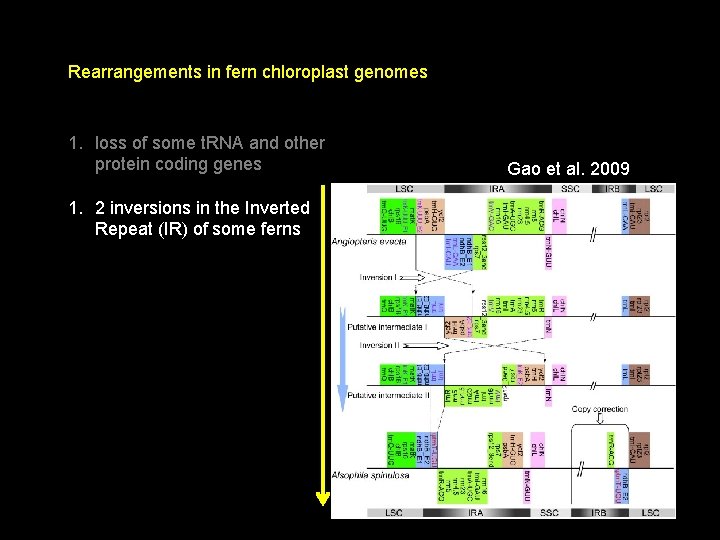
Rearrangements in fern chloroplast genomes 1. loss of some t. RNA and other protein coding genes 1. 2 inversions in the Inverted Repeat (IR) of some ferns Gao et al. 2009

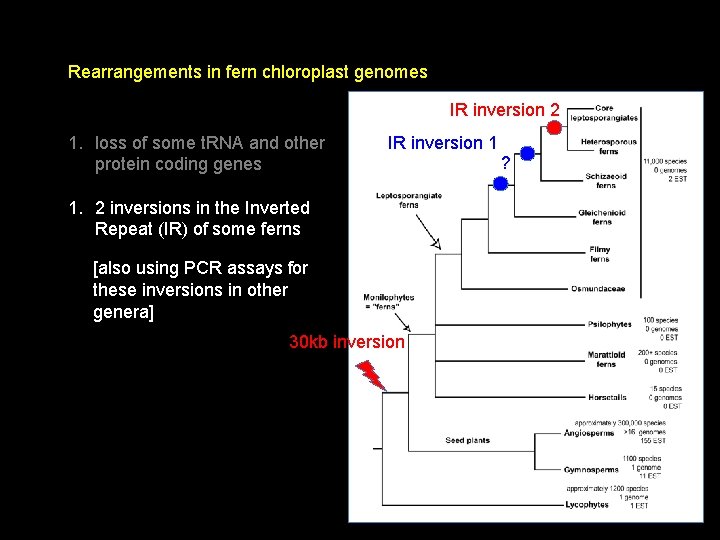
Rearrangements in fern chloroplast genomes IR inversion 2 1. loss of some t. RNA and other protein coding genes IR inversion 1 1. 2 inversions in the Inverted Repeat (IR) of some ferns [also using PCR assays for these inversions in other genera] 30 kb inversion ?

Chapter 7: Evolution of the nuclear genome of ferns and lycophytes Takuya Nakazato, Michael S. Barker, Loren H. Rieseberg, and Gerald J. Gastony Unfurling fern biology in the genomics age (Bio. Science, 2010) Michael S. Barker and Paul G. Wolf

Academic family tree of Gerald J. Gastony

Rolla and Alice Tryon 1950 s and 1990 s Is there an “Alice Tryon Women in Science” bequest for Botany Department?

Academic family tree of Gerald J. Gastony Rieseberg Nakazato Barker

The neglected fern and lycophyte nuclear genomes - or the “crying ferns” 1. 1 genetic linkage map - Ceratopteris 1. 4 EST libraries – Selaginella (2), Ceratopteris, Adiantum 2. 3 BAC libraries - Selaginella (2), Ceratopteris 3. 1 nuclear genome sequencing project in the works - Selaginella

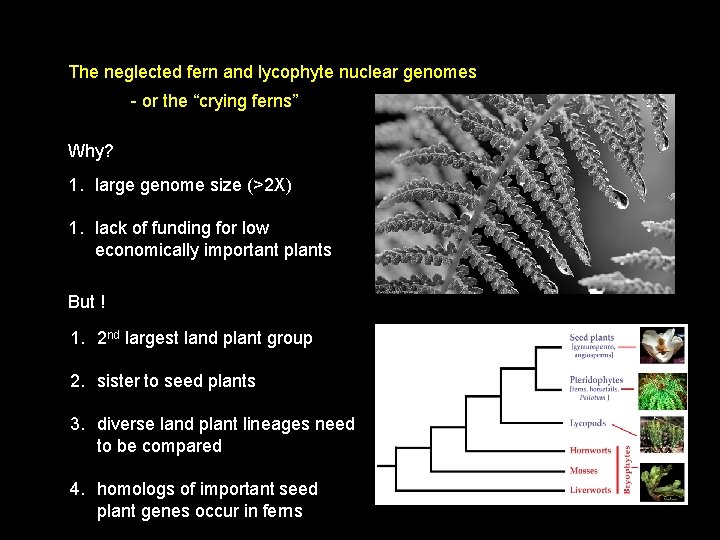
The neglected fern and lycophyte nuclear genomes - or the “crying ferns” Why? 1. large genome size (>2 X) 1. lack of funding for low economically important plants

The neglected fern and lycophyte nuclear genomes - or the “crying ferns” Why? 1. large genome size (>2 X) 1. lack of funding for low economically important plants But ! 1. 2 nd largest land plant group 2. sister to seed plants 3. diverse land plant lineages need to be compared 4. homologs of important seed plant genes occur in ferns

A short history of the study of the fern genome Haploid chromosome number • 57 in ferns vs. 16 in angiosperms [ > 14 = polyploid (Grant, 1981) ] Ophioglossum (adder’s-tongue fern) - 2 n = 1440 (96 ploid) in O. reticulatum

A short history of the study of the fern genome Haploid chromosome number • 57 in ferns vs. 16 in angiosperms [ > 14 = polyploid (Grant, 1981) ] Questions: How does this fern choreograph meiosis with an n > 600? Has it ever been observed? Do large n's lead to more aborted or nonviable spores?

A short history of the study of the fern genome Haploid chromosome number • 57 in ferns vs. 16 in angiosperms [ > 14 = polyploid (Grant, 1981) ] • 13. 6 in heterosporous ferns is exception • heterosporous lycophytes << homosporous lycophytes • heterosporous seed plants << homosporous ferns & allies Therefore, homosporous ferns acquire high chromosome number to select for increased heterozygosity via polyploidy Hypothesis of Klekowski & Baker (1966)

A short history of the study of the fern genome Two lines of evidence did not support this hypothesis 1. Isozyme analysis indicated widespread silencings of genes – diploid numbers of copies 1. nn 2. Most homosporous ferns are outcrossing Therefore, homosporous ferns acquire high chromosome number to select for increased heterozygosity via polyploidy Hypothesis of Klekowski & Baker (1966)

A short history of the study of the fern genome Homosporous ferns acquired high chromosome numbers with diploid gene expression via repeated cycles of polyploidization and subsequent gene silencing without chromosome loss Hypothesis of Chris Haufler (1987)

A short history of the study of the fern genome Many lines of evidence support this as the working hypothesis in ferns 1. Pseudogenes in nuclear genes in Polystichum 1. FISH detection of multiple dispersed chromosomal locations of r. DNA in Ceratopteris 1. +/- Genetic linkage map analysis in Ceratopteris Homosporous ferns acquired high chromosome numbers with diploid gene expression via repeated cycles of polyploidization and subsequent gene silencing without chromosome loss Hypothesis of Chris Haufler (1987)

The future of fern genomics? Ceratopteris has emerged as the “model” organism for fern genomics Study of the origin of polyploidy (neo- and paleo-) Correlating genomic changes to speciation and development Two examples using Ceratopteris 1. Nakazato et al. (2006) genetic linkage analysis 1. Barker (2010) EST analysis

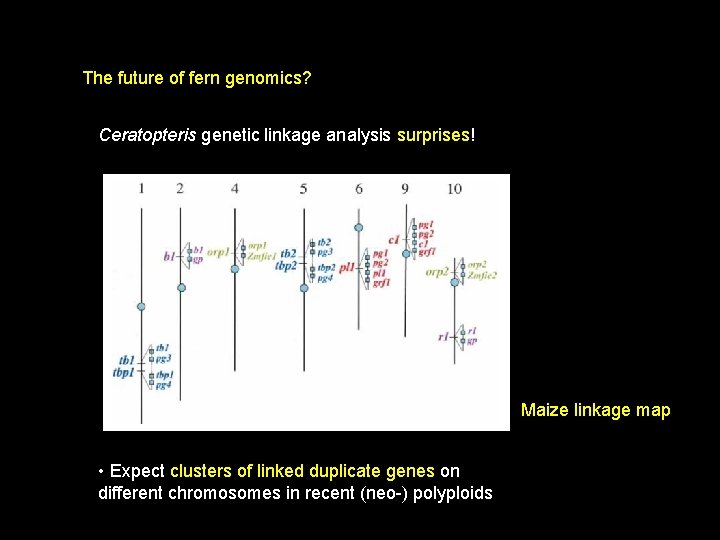
The future of fern genomics? Ceratopteris genetic linkage analysis • 700 genetic markers • 85% multiple copies • 24% single copy – low! • large numbers of duplicate genes on different chromosomes

The future of fern genomics? Ceratopteris genetic linkage analysis surprises! Maize linkage map • Expect clusters of linked duplicate genes on different chromosomes in recent (neo-) polyploids

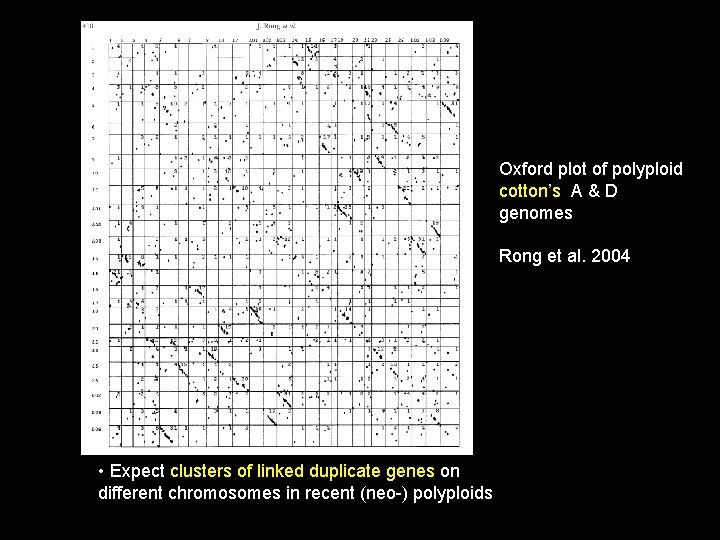
Oxford plot of polyploid cotton’s A & D genomes Rong et al. 2004 • Expect clusters of linked duplicate genes on different chromosomes in recent (neo-) polyploids

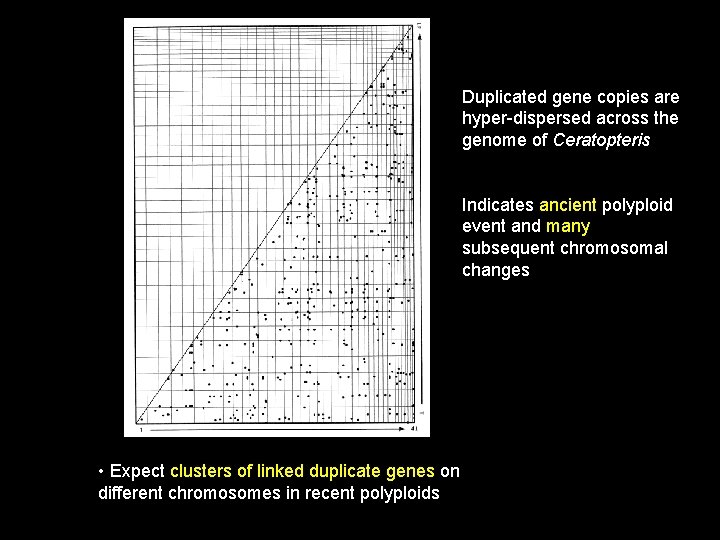
Duplicated gene copies are hyper-dispersed across the genome of Ceratopteris Indicates ancient polyploid event and many subsequent chromosomal changes • Expect clusters of linked duplicate genes on different chromosomes in recent polyploids

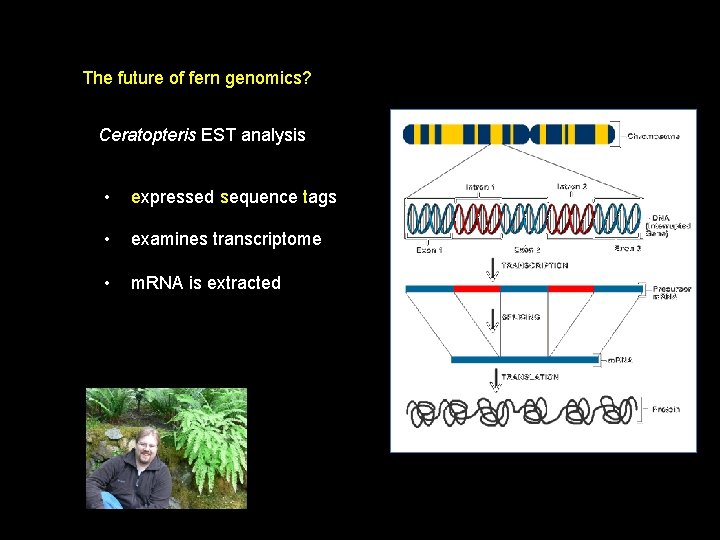
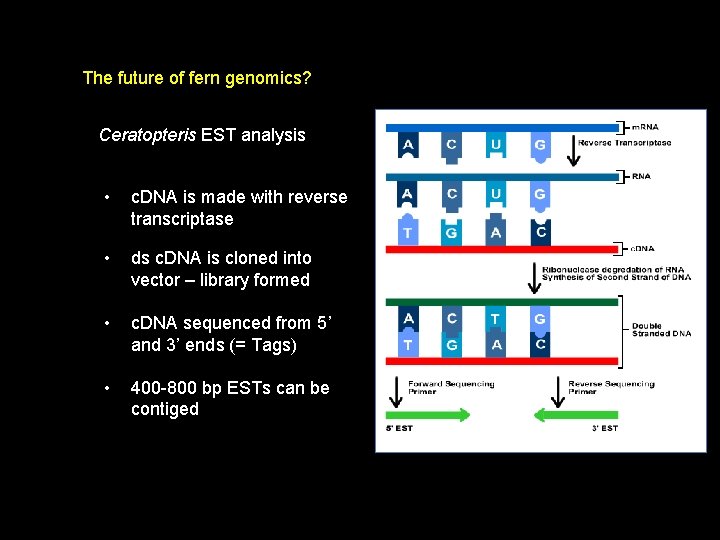
The future of fern genomics? Ceratopteris EST analysis • expressed sequence tags • examines transcriptome • m. RNA is extracted

The future of fern genomics? Ceratopteris EST analysis • c. DNA is made with reverse transcriptase • ds c. DNA is cloned into vector – library formed • c. DNA sequenced from 5’ and 3’ ends (= Tags) • 400 -800 bp ESTs can be contiged

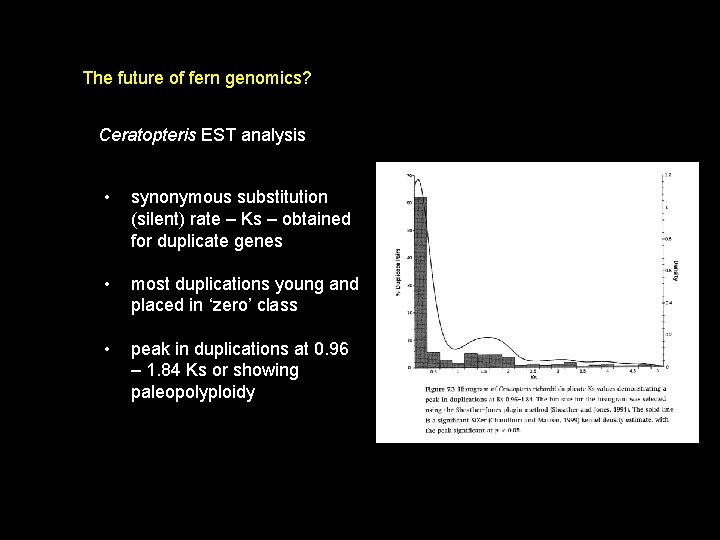
The future of fern genomics? Ceratopteris EST analysis • synonymous substitution (silent) rate – Ks – obtained for duplicate genes • most duplications young and placed in ‘zero’ class • peak in duplications at 0. 96 – 1. 84 Ks or showing paleopolyploidy

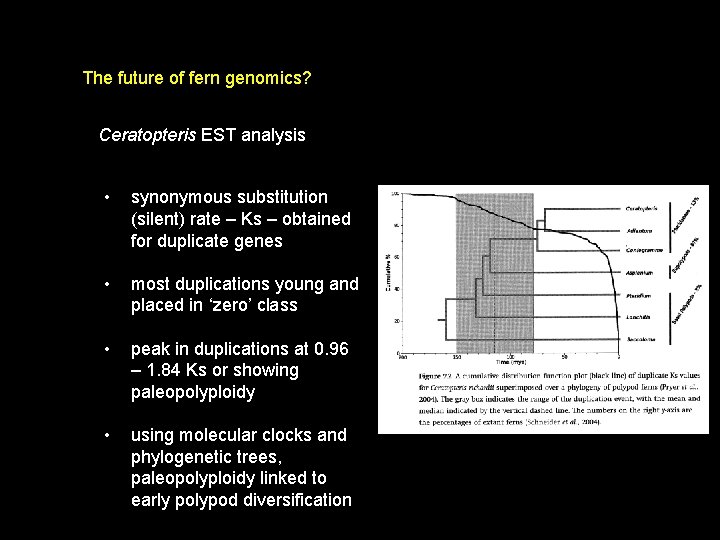
The future of fern genomics? Ceratopteris EST analysis • synonymous substitution (silent) rate – Ks – obtained for duplicate genes • most duplications young and placed in ‘zero’ class • peak in duplications at 0. 96 – 1. 84 Ks or showing paleopolyploidy • using molecular clocks and phylogenetic trees, paleopolyploidy linked to early polypod diversification

Question Set 1 Question Set 2 1. Ferns and fern allies are diverse and old; is 1. What are the justifications for selecting it really appropriate to expect that all have Ceratopteris richardii as a model their nuclear genomes evolving by same organism for ferns? Do the “idiosyncratic” “rules”? features of its genome affect generalization to ferns? 1. You have been given a blank check to sequence the fern genome of your 2. Could maintaining large amounts of choice. Which would you choose and why? physical genetic material be What methods would you use? disadvantageous for fern evolution? Could it be related to slow speciation 2. Why is the fate of most duplicate genes to rates, compared to angiosperms? Or, on eventually become silenced? Could the other hand, could the silenced genes mutations accumulate in both copies at the hold the key to the long history of fern same rate causing subfunctionalization, evolution? where mutations cause the two copies to functionally be diminished to one over 1. Can high chromosome numbers in ferns time? and lycophytes simply be an outcome of the ‘stringent bivalent pairing’ that is 3. If you are really interested in known in the group? How might that idea understanding the process of speciation, be further examined or tested? would ferns be the better choice relative to angiosperms?