Genomics Bioinformatics Medicine http biochem 158 stanford edu

Genomics, Bioinformatics & Medicine http: //biochem 158. stanford. edu/ Gene Expression http: //biochem 158. stanford. edu/Gene%20 Expression. html Doug Brutlag Professor Emeritus of Biochemistry & Medicine Stanford University School of Medicine Doug Brutlag 2011

Leveraging Genomic Information Novel Diagnostics DNA Microchips & Microarrays Gene Expression - RNA Proteomics - Protein Understanding Metabolism Cell Signaling Differentiation Disease Doug Brutlag 2011

Gene Regulatory Mechanisms • Transcriptional Mechanisms – Type of promoters & RNA polymerase – Control of Transcription – Transcription Factors and TFBS • RNA processing – – – Capping Splicing and Alternative Splicing Poly-Adenylation RNA export to cytoplasm RNA degradation rates • Micro RNAs (mi. RNAs) inhibit translation and degrade m. RNA • Silencer RNAs (si. RNAs or RNAi) degrading m. RNA • Epigenetic Mechanisms – DNA methylation – Histone modifications • Acetylation • Methylation • Phosphorylation, etc. – Chromatin remodeling Doug Brutlag 2011
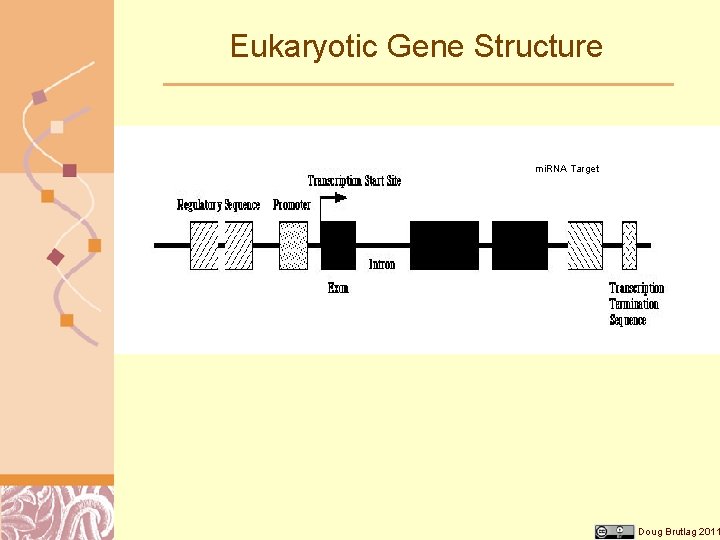
Eukaryotic Gene Structure mi. RNA Target Doug Brutlag 2011
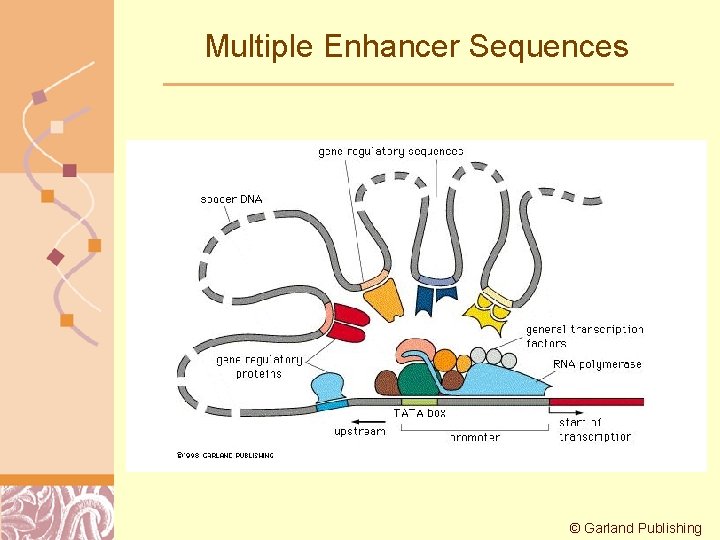
Multiple Enhancer Sequences © Garland Publishing Doug Brutlag 2011
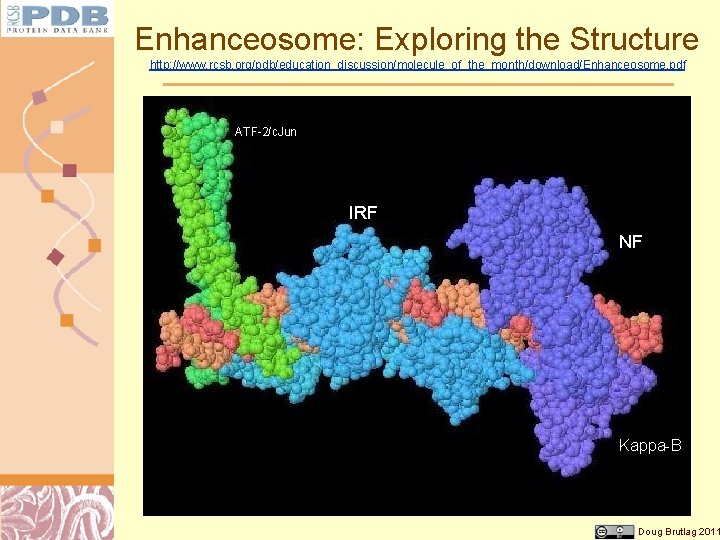
Enhanceosome: Exploring the Structure http: //www. rcsb. org/pdb/education_discussion/molecule_of_the_month/download/Enhanceosome. pdf ATF-2/c. Jun IRF NF Kappa-B Doug Brutlag 2011
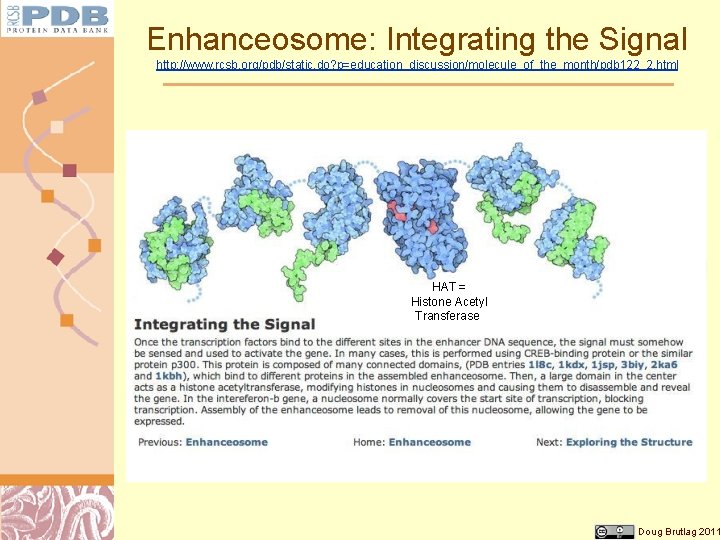
Enhanceosome: Integrating the Signal http: //www. rcsb. org/pdb/static. do? p=education_discussion/molecule_of_the_month/pdb 122_2. html HAT = Histone Acetyl Transferase Doug Brutlag 2011
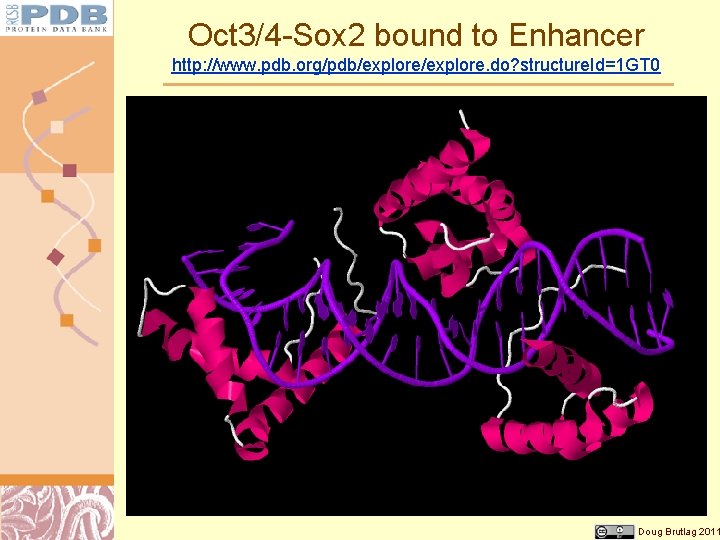
Oct 3/4 -Sox 2 bound to Enhancer http: //www. pdb. org/pdb/explore. do? structure. Id=1 GT 0 Doug Brutlag 2011
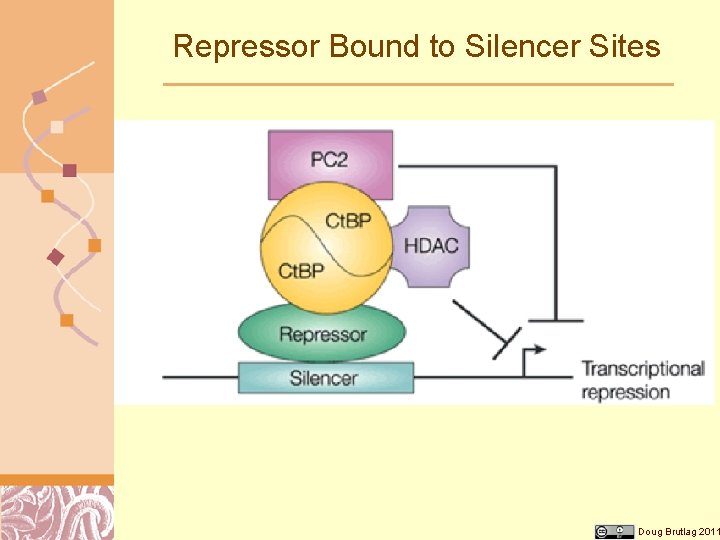
Repressor Bound to Silencer Sites Doug Brutlag 2011
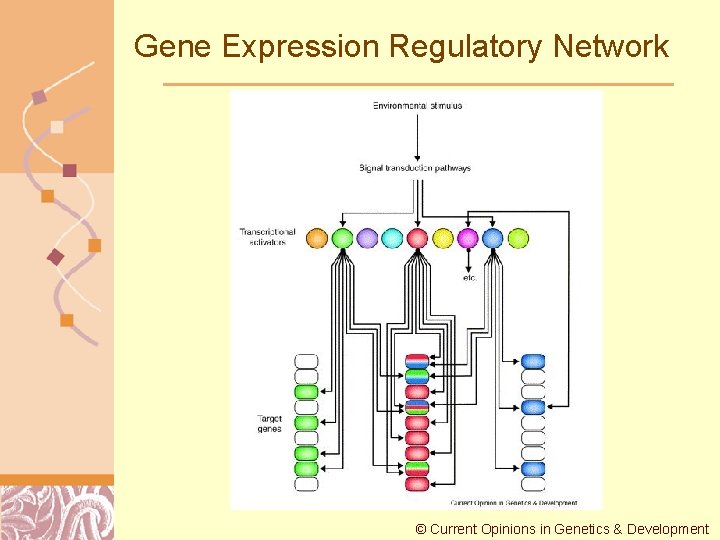
Gene Expression Regulatory Network © Current Opinions in Genetics & Development Doug Brutlag 2011
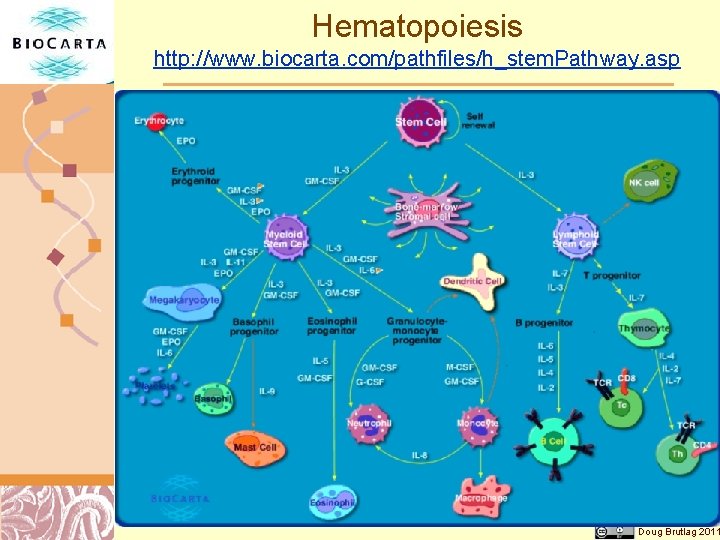
Hematopoiesis http: //www. biocarta. com/pathfiles/h_stem. Pathway. asp Doug Brutlag 2011
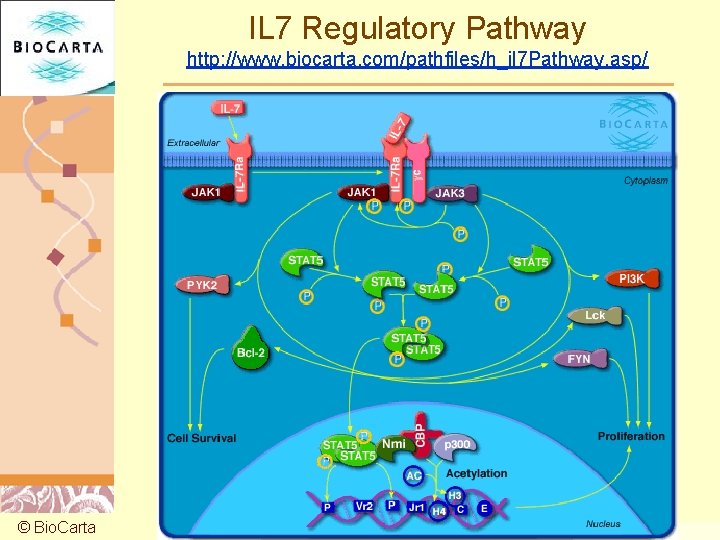
IL 7 Regulatory Pathway http: //www. biocarta. com/pathfiles/h_il 7 Pathway. asp/ © Bio. Carta Doug Brutlag 2011
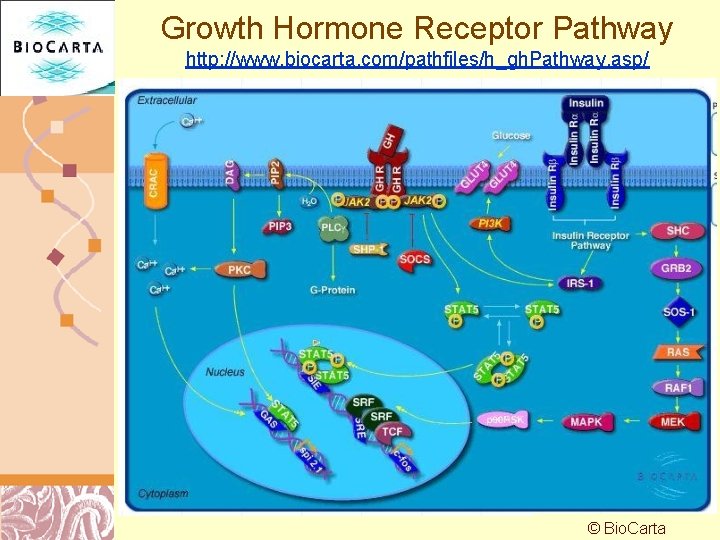
Growth Hormone Receptor Pathway http: //www. biocarta. com/pathfiles/h_gh. Pathway. asp/ © Bio. Carta Doug Brutlag 2011

DNA Microarrays & DNA Chips Accelerate Gene Expression. Analysis • Parallel Analyses – Analyze entire genomes instead of single genes – Analyze expression of entire genome – Analyze genetic polymorphisms (SNPs) • Miniaturization • Automation Doug Brutlag 2011
Diagnosis Using DNA Arrays http: //www. affymetrix. com/ © Affymetrix Doug Inc. Brutlag 2011

Affymetrix Arrays http: //www. affymetrix. com/ Any collection of 25 mers (1, 200, 000) can be synthesized in 100 steps © Affymetrix Doug Inc. Brutlag 2011
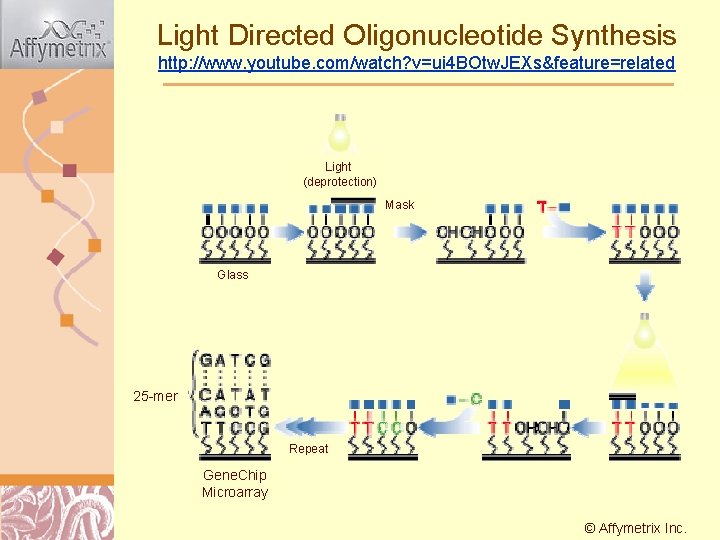
Light Directed Oligonucleotide Synthesis http: //www. youtube. com/watch? v=ui 4 BOtw. JEXs&feature=related Light (deprotection) Mask Glass 25 -mer Repeat Gene. Chip Microarray © Affymetrix Doug Inc. Brutlag 2011
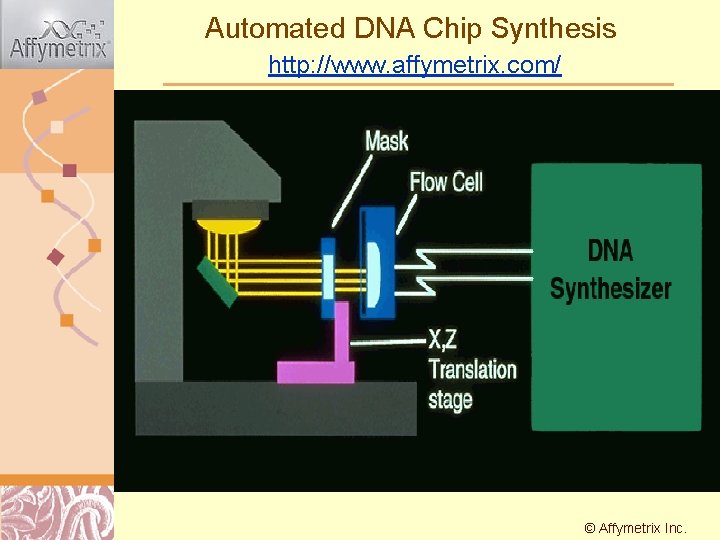
Automated DNA Chip Synthesis http: //www. affymetrix. com/ © Affymetrix Doug Inc. Brutlag 2011
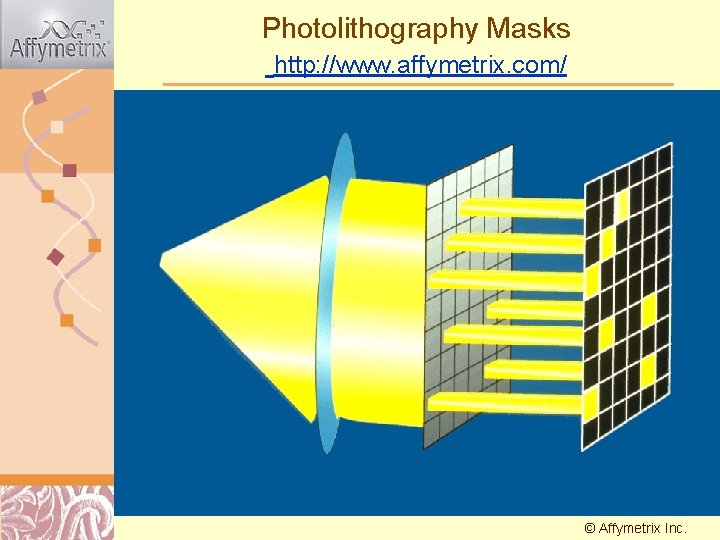
Photolithography Masks http: //www. affymetrix. com/ © Affymetrix Doug Inc. Brutlag 2011
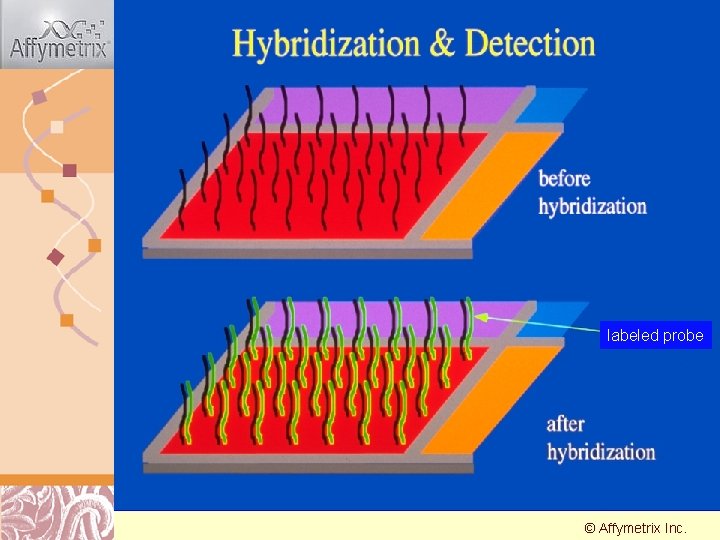
Hybridization & Detection labeled probe © Affymetrix Doug Inc. Brutlag 2011
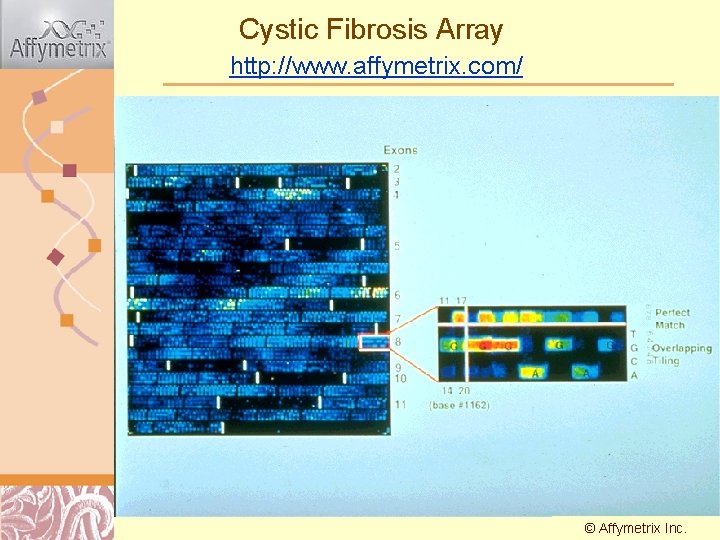
Cystic Fibrosis Array http: //www. affymetrix. com/ © Affymetrix Doug Inc. Brutlag 2011

Microarrayer in Pat Brown’s Lab http: //cmgm. stanford. edu/pbrown/ Doug Brutlag 2011

High Precision DNA Printing Arrayit Corporation Doug Brutlag 2011

Mechanical Spotting Microarrays http: //www. arrayit. com/ Arrayit Doug Corporation Brutlag 2011

DNA Chips are used to Measure Gene Expression Micro. Array analysis of whole genome gene expression Clustering of genes based on their expression pattern Searching for conserved sequence motifs regulating the expression Doug Brutlag 2011
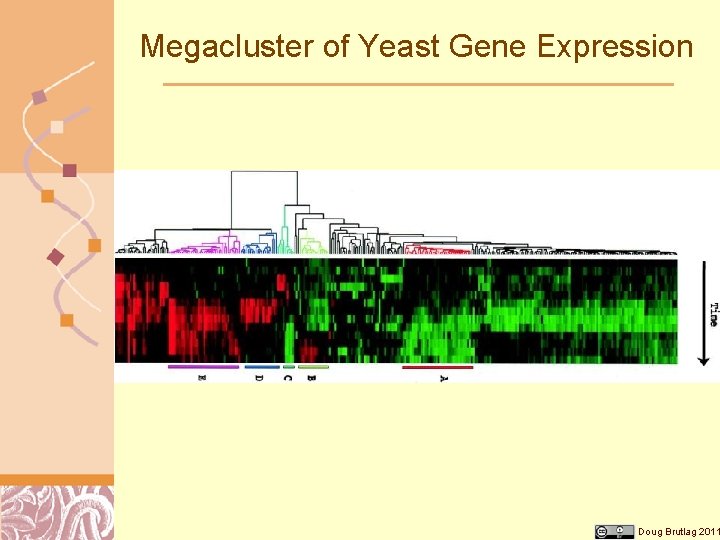
Megacluster of Yeast Gene Expression Doug Brutlag 2011
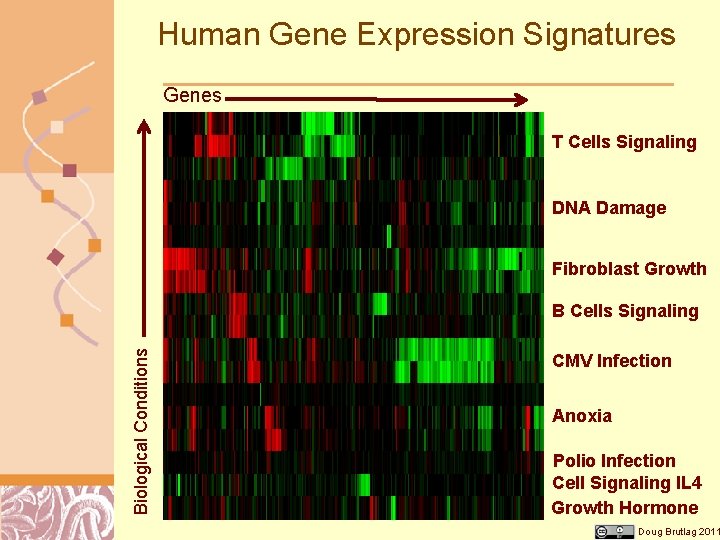
Human Gene Expression Signatures Genes T Cells Signaling DNA Damage Fibroblast Growth Biological Conditions B Cells Signaling CMV Infection Anoxia Polio Infection Cell Signaling IL 4 Growth Hormone Doug Brutlag 2011
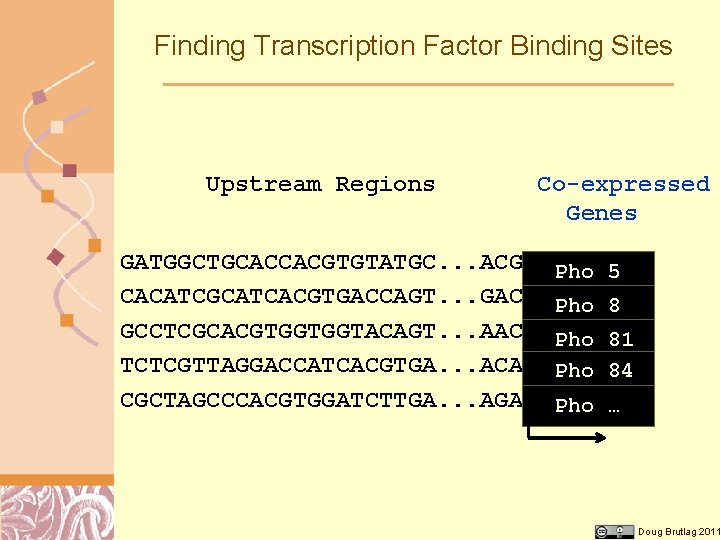
Finding Transcription Factor Binding Sites Upstream Regions Co-expressed Genes GATGGCTGCACCACGTGTATGC. . . ACGATGTCTCGC Pho 5 CACATCGCATCACGTGACCAGT. . . GACATGGACGGC Pho 8 GCCTCGCACGTGGTGGTACAGT. . . AACATGACTAAA Pho 81 TCTCGTTAGGACCATCACGTGA. . . ACAATGAGAGCG Pho 84 CGCTAGCCCACGTGGATCTTGA. . . AGAATGACTGGC Pho … Transcription Start Doug Brutlag 2011

Finding Transcription Factor Binding Sites Upstream Regions Co-expressed Genes GATGGCTGCACCACGTGTATGC. . . ACGATGTCTCGC CACATCGCATCACGTGACCAGT. . . GACATGGACGGC GCCTCGCACGTGGTGGTACAGT. . . AACATGACTAAA TCTCGTTAGGACCATCACGTGA. . . ACAATGAGAGCG CGCTAGCCCACGTGGATCTTGT. . . AGAATGGCCTAT Doug Brutlag 2011
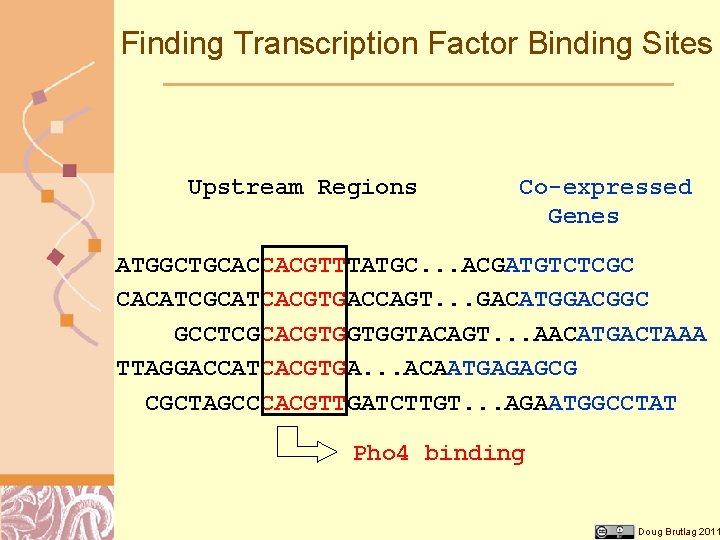
Finding Transcription Factor Binding Sites Upstream Regions Co-expressed Genes ATGGCTGCACCACGTTTATGC. . . ACGATGTCTCGC CACATCGCATCACGTGACCAGT. . . GACATGGACGGC GCCTCGCACGTGGTGGTACAGT. . . AACATGACTAAA TTAGGACCATCACGTGA. . . ACAATGAGAGCG CGCTAGCCCACGTTGATCTTGT. . . AGAATGGCCTAT Pho 4 binding Doug Brutlag 2011

Discovering Transcription Factor Binding Sites is Difficult • • Binding sites are short (5 -15 base pairs) Not highly conserved (as little as 50%) Located in long intergenic regions (>10 kb) Not always present (false positive genes) Doug Brutlag 2011

Three Algorithms • Bio. Prospector – Presented in 2000 – Extends Gibb’s sampling (stochastic method) – For any cluster of sequences • MDscan – Deterministic approach – Enumerative – Very fast – For sequences with some ranking information • Motif. Cut and Motif. Scan – Graph-based – Does not use PSSMs – Novel and sensitive Doug Brutlag 2011
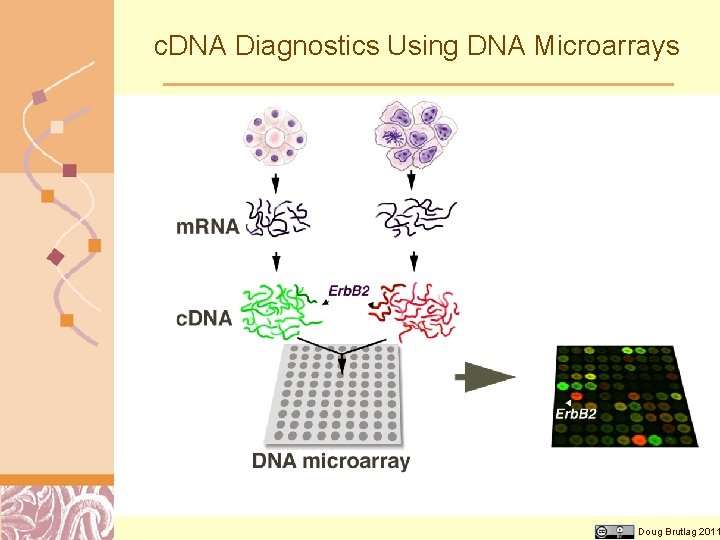
c. DNA Diagnostics Using DNA Microarrays Doug Brutlag 2011
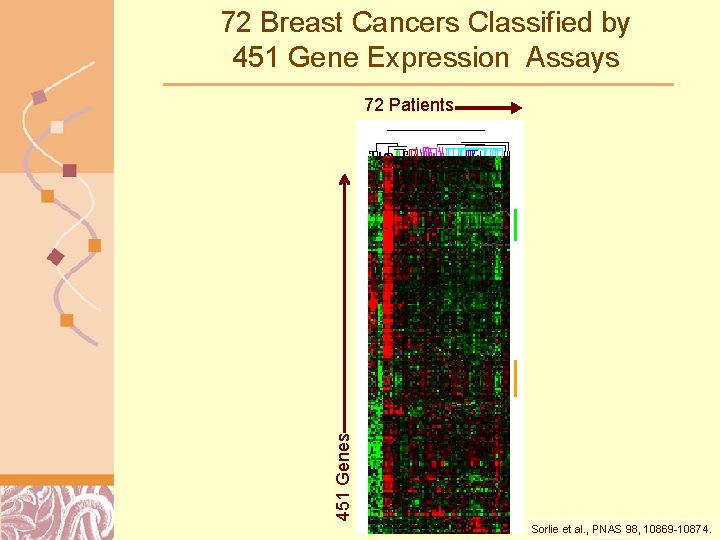
72 Breast Cancers Classified by 451 Gene Expression Assays 451 Genes 72 Patients Sorlie et al. , PNAS 98, Doug 10869 -10874. Brutlag 2011
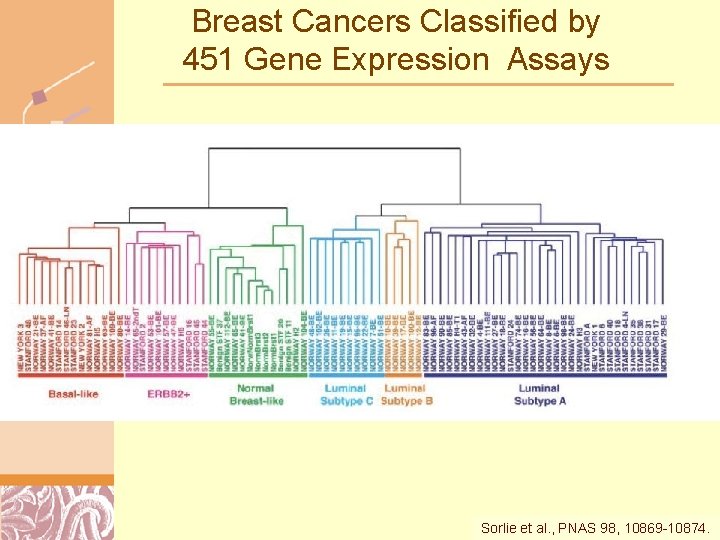
Breast Cancers Classified by 451 Gene Expression Assays Sorlie et al. , PNAS 98, 10869 -10874. Doug Brutlag 2011
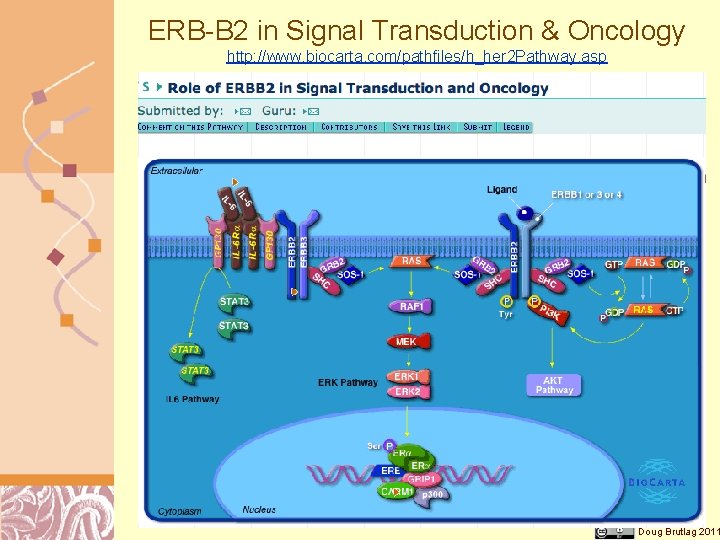
ERB-B 2 in Signal Transduction & Oncology http: //www. biocarta. com/pathfiles/h_her 2 Pathway. asp Doug Brutlag 2011
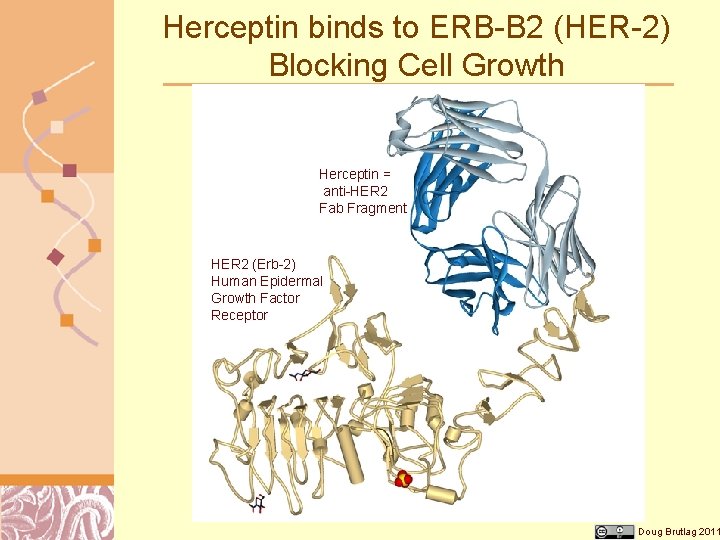
Herceptin binds to ERB-B 2 (HER-2) Blocking Cell Growth Herceptin = anti-HER 2 Fab Fragment HER 2 (Erb-2) Human Epidermal Growth Factor Receptor Doug Brutlag 2011
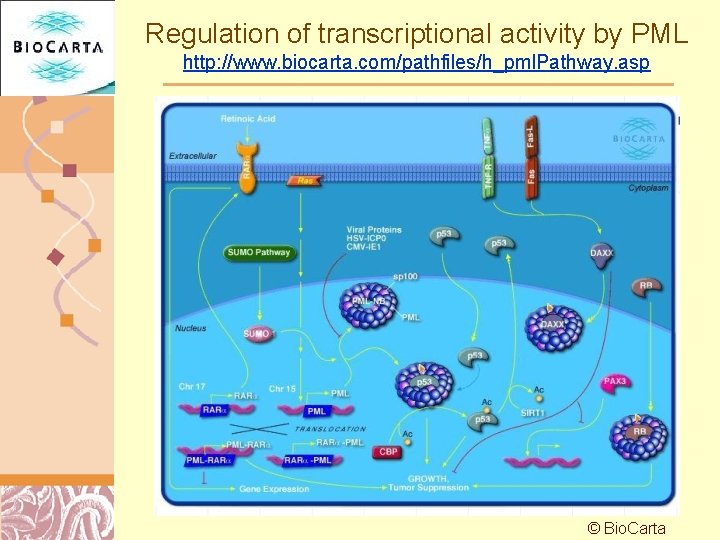
Regulation of transcriptional activity by PML http: //www. biocarta. com/pathfiles/h_pml. Pathway. asp © Bio. Carta Doug Brutlag 2011
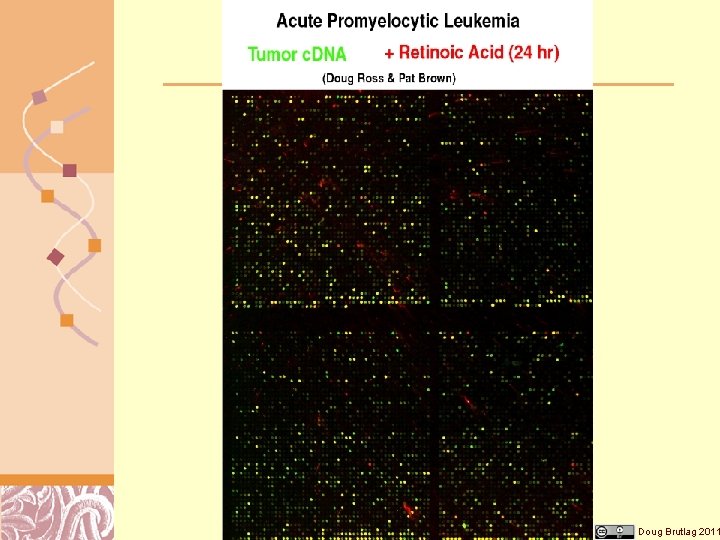
Diagnosis & Gene Expression Doug Brutlag 2011

Path. Work Diagnostics http: //www. pathworkdx. com/ Predicting Site of Origin for Cancers of Unknown Primary © Pathwork Diagnostics Inc. Doug Brutlag 2011

Path. Work Oncology Suite: Site of Origin • 1844 tumors tested one at a time versus all 18 tissues of origin • Retrospective study on well characterized patient samples • Uses Path. Chip (functionally similar to Affymetrix HU-133 A Gene. Chip) – 604 specimens used for training – 636 specimens used for test – 604 specimens in reserve for final validation • Reproducibility from lab to lab • Performance based on sensitivity (> 70%) & accuracy (> 95%) © Pathwork Diagnostics Inc. Doug Brutlag 2011
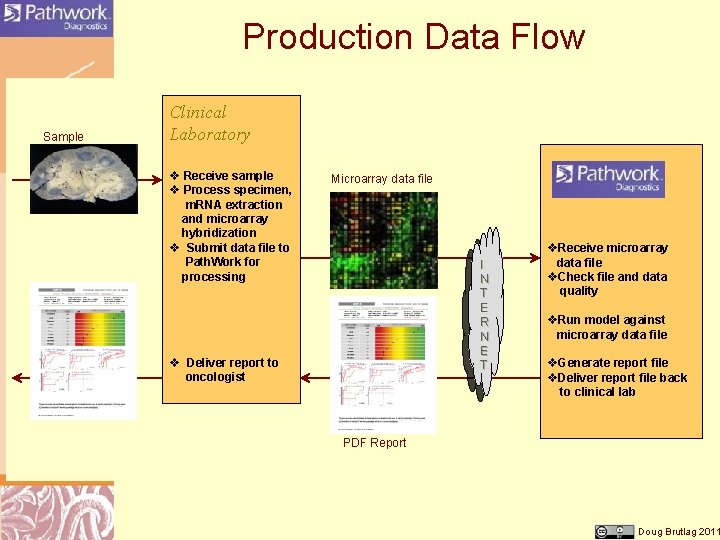
Production Data Flow Sample Clinical Laboratory v Receive sample v Process specimen, m. RNA extraction and microarray hybridization v Submit data file to Path. Work for processing Microarray data file I N T E R N E T v Deliver report to oncologist v. Receive microarray data file v. Check file and data quality v. Run model against microarray data file v. Generate report file v. Deliver report file back to clinical lab PDF Report Doug Brutlag 2011
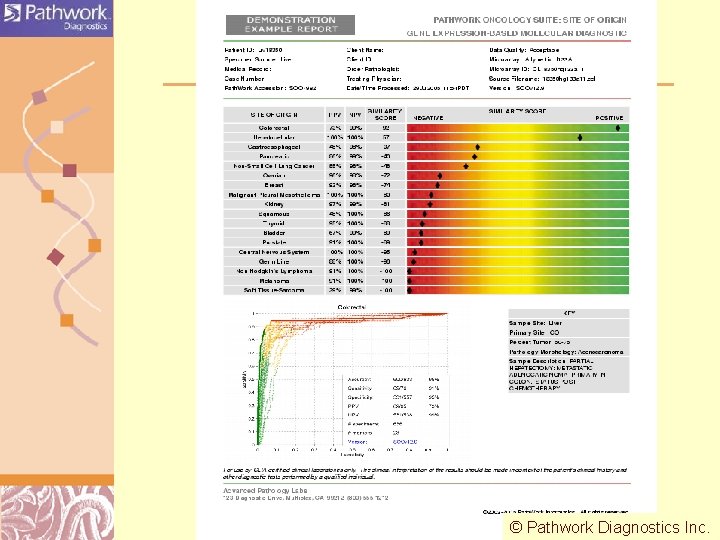
CONFIDENTIAL 43 © Pathwork Diagnostics Inc. Doug Brutlag 2011
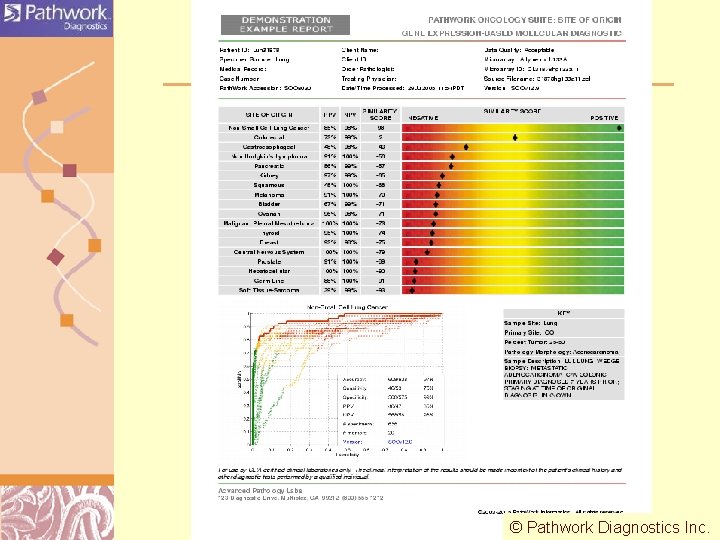
CONFIDENTIAL 44 © Pathwork Diagnostics Inc. Doug Brutlag 2011
- Slides: 44