Genomic Measures of Relationship and Inbreeding Paul Van

Genomic Measures of Relationship and Inbreeding Paul Van. Raden Animal Improvement Programs Laboratory, USDA Agricultural Research Service, Beltsville, MD, USA Paul. Van. Raden@ars. usda. gov 2007
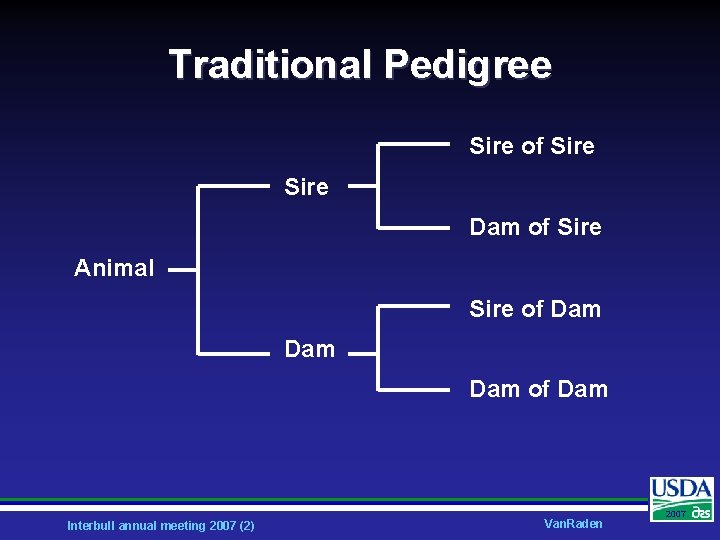
Traditional Pedigree Sire of Sire Dam of Sire Animal Sire of Dam Dam of Dam Interbull annual meeting 2007 (2) Van. Raden 2007
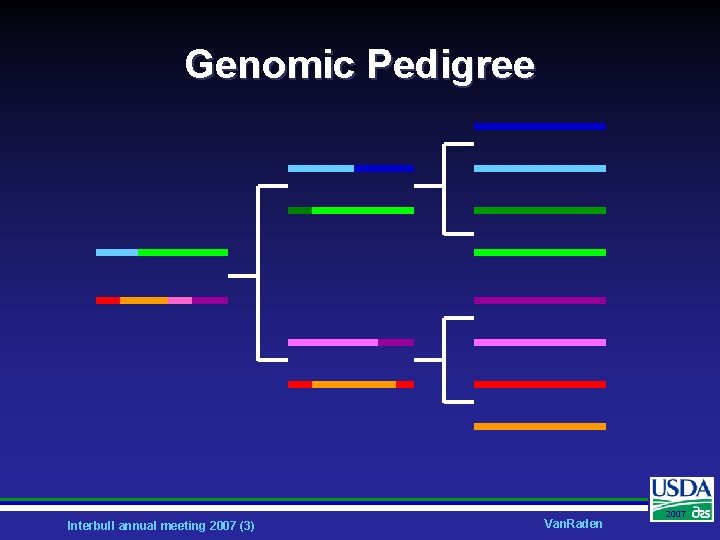
Genomic Pedigree Interbull annual meeting 2007 (3) Van. Raden 2007
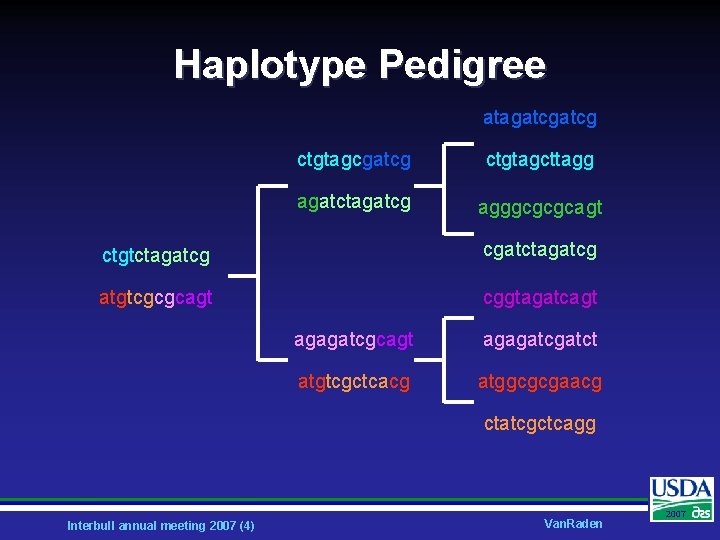
Haplotype Pedigree atagatcg ctgtagcgatcg ctgtagcttagg agatctagatcg agggcgcgcagt ctgtctagatcg cgatctagatcg atgtcgcgcagt cggtagatcagt agagatcgatct atgtcgctcacg atggcgcgaacg ctatcgctcagg Interbull annual meeting 2007 (4) Van. Raden 2007

Genotype Pedigree Count number of second allele 121101011110 11120200 101121101111 122221121111 101101111102 011111012011 121120011010 0 = homozygous for first allele (alphabetically) 1 = heterozygous 2 = homozygous for second allele (alphabetically) Interbull annual meeting 2007 (5) Van. Raden 2007

How Related are Relatives? Ø Example: Full sibs • • Ø are expected to share 50% of their DNA on average may actually share 45% or 55% of their DNA because each inherits a different mixture of chromosome segments from the two parents. Combine genotype and pedigree data to determine exact fractions Interbull annual meeting 2007 (6) Van. Raden 2007
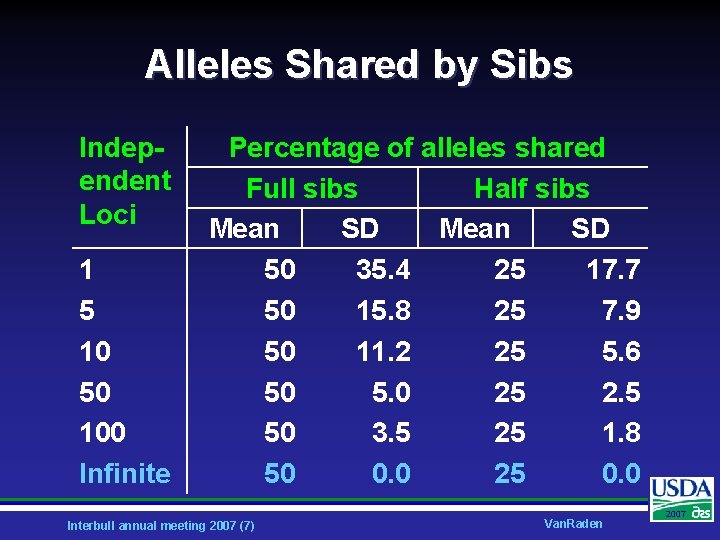
Alleles Shared by Sibs Independent Loci 1 5 10 50 100 Infinite Percentage of alleles shared Full sibs Half sibs Mean SD 50 35. 4 25 17. 7 50 15. 8 25 7. 9 50 11. 2 25 5. 6 50 5. 0 25 2. 5 50 3. 5 25 1. 8 50 0. 0 25 0. 0 Interbull annual meeting 2007 (7) Van. Raden 2007

Relationship Matrices Ø Measures of genetic similarity • • • A = Expected % genes identical by descent from pedigree (Wright, 1922) G = Actual % of DNA shared (using genotype data) T = % genes shared that affect a given trait (using genotype and phenotype) Interbull annual meeting 2007 (8) Van. Raden 2007

Computing Relationships Ø Construct G from marker incidence matrix M minus allele frequencies pj • • Ø M = markers (j) inherited by animals (i) P contains frequency of second allele Z = M – P (elements of Z are –pj or 1 -pj) G = Z Z’ / [2 ∑ pj(1 -pj)] Construct T using similar math, but all QTL that affect a trait not observable Interbull annual meeting 2007 (9) Van. Raden 2007

Linear Model Equations Ø BLUP equations for marker effects, then sum to get EBV • • Ø Selection index equations for EBV • • Ø u^ = Z [Z’R-1 Z + I k]-1 Z’R-1(y – Xb) k = var(u) / var(m) = 2 ∑ pj(1 -pj) u^ = Z Z’ [Z Z’ + R]-1 (y – Xb) R has diagonals = (1 / Reliability) - 1 Equivalent model from Garrick (2007) • u^ = [(Z Z’)-1 + R-1]-1 R-1 (y – Xb) Interbull annual meeting 2007 (10) Van. Raden 2007
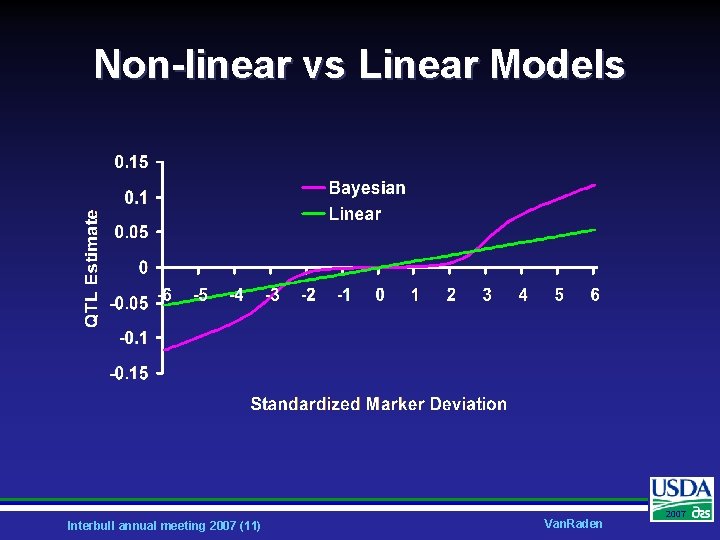
Non-linear vs Linear Models Interbull annual meeting 2007 (11) Van. Raden 2007
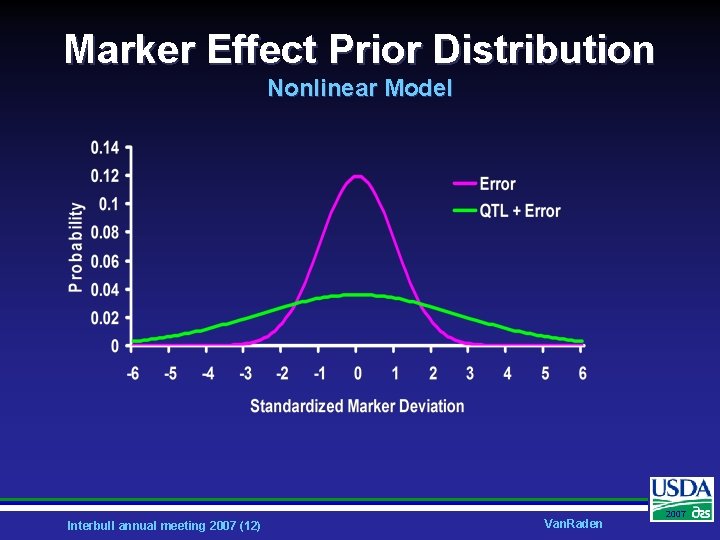
Marker Effect Prior Distribution Nonlinear Model Interbull annual meeting 2007 (12) Van. Raden 2007
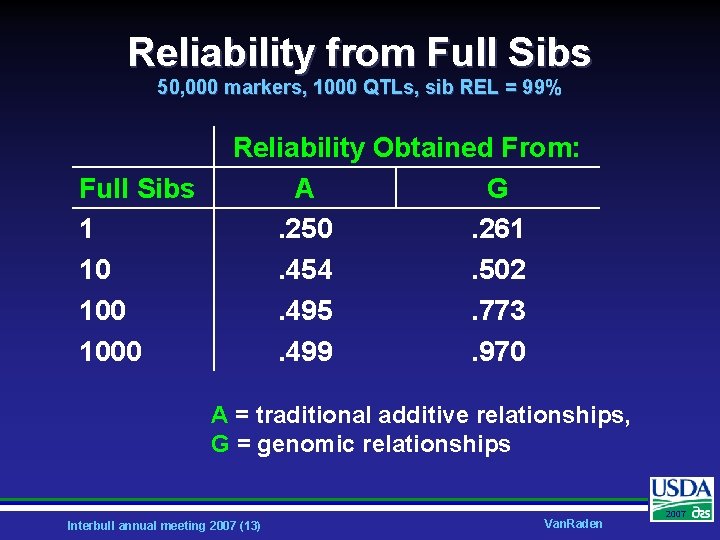
Reliability from Full Sibs 50, 000 markers, 1000 QTLs, sib REL = 99% Full Sibs 1 10 1000 Reliability Obtained From: A G. 250. 261. 454. 502. 495. 773. 499. 970 A = traditional additive relationships, G = genomic relationships Interbull annual meeting 2007 (13) Van. Raden 2007

Reliability from Genotyping Ø Daughter equivalents • • • Ø DETotal = DEPA + DEProg + DEYD + DEG is additional DE from genotype Reliability = DEtotal / (DETotal + k) Gains in reliability • • • DEG could be about 15 for Net Merit More for traits with low heritability Less for traits with high heritability Interbull annual meeting 2007 (14) Van. Raden 2007

Conclusions Ø Relationships can be defined as: • • • Ø Ø A = expected genes in common G = actual fraction of DNA in common T = QTL alleles in common for a trait Full sibs share 50% ± 3. 5% of DNA Genomic (G) or non-linear models can better approximate QTL relationships (T) and increase reliability as compared to traditional relationships (A) Interbull annual meeting 2007 (15) Van. Raden 2007
- Slides: 15