Genomic identification of Structural RNAs using phyloSCFGs Jakob

Genomic identification of Structural RNAs using phylo-SCFGs Jakob Skou Pedersen Bioinformatics Center, University of Copenhagen Ref: Knudsen B & Hein J. 1999

Structural RNA identification problem Genome nc. RNA protein-coding gene Structural RNA: any transcribed region with functional structure. Such as: • independently transcribed nc. RNAs • nc. RNAs co-transcribed with protein-coding genes • cis-regulatory elements within protein-coding genes

Highly diverse set histone 3’UTR stem loop mi. RNA t. RNA RNase P Xist (~20 kb long) Little single sequence signal: • Lack of common nucleotide biases • Lack of common sequence motifs • Free energies of structures generally insignificant Figures from: rfam (http: //www. sanger. ac. uk/Software/Rfam/), Cell, Vol. 116, 281– 297, January 23, 2004 , & Ng et al. EMBO reports 8, 1,

Evolutionary signal Signal Unprecedented comparative data Structure functionally important Primary sequence subs tolerated From: Margulies et al. , Genome Reseach, 2007.

Introduction to phylogenetic models Based on: Continuous time Markov chain acting on Substitution rates of Markov chain branches of phylogenetic tree Captures: • Nucleotide biases • Patterns of substitution • Evolutionary sequence correlations • Correlated changes (multi nucleotide models) Transition prob. : Felsenstein 81: Refs. : Jukes TH & Cantor CR, 1969, in Mammalian protein metabolism & Felsenstein J. 1981, Journal of Molecular Evolution. Alignment column

Evo. Fold Phylogenetic models Di-nucleotide model • • Single-nucleotide model 16 x 16 rate matrix • 4 x 4 rate matrix Learned from data • Marginal average of di- Favors pairing di-nucs nucleotide matrix Slow substitution rate • Fast substitution rate Ref: Yang Z, Nielsen R, Hasegawa M,
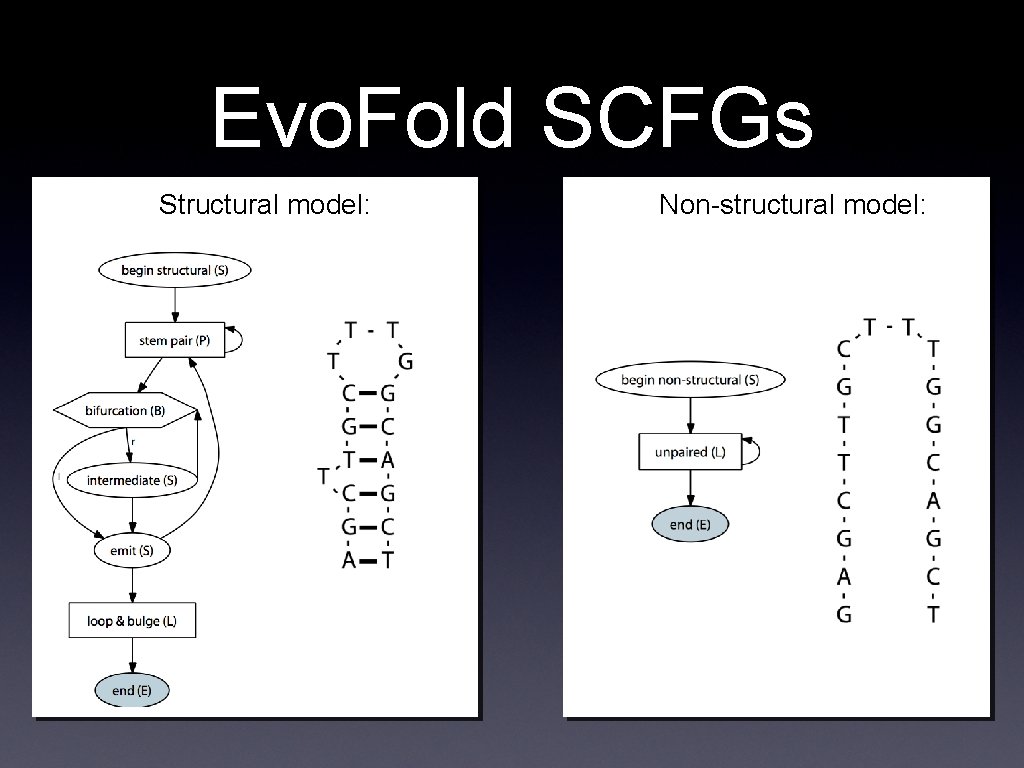
Evo. Fold SCFGs Structural model: Non-structural model:
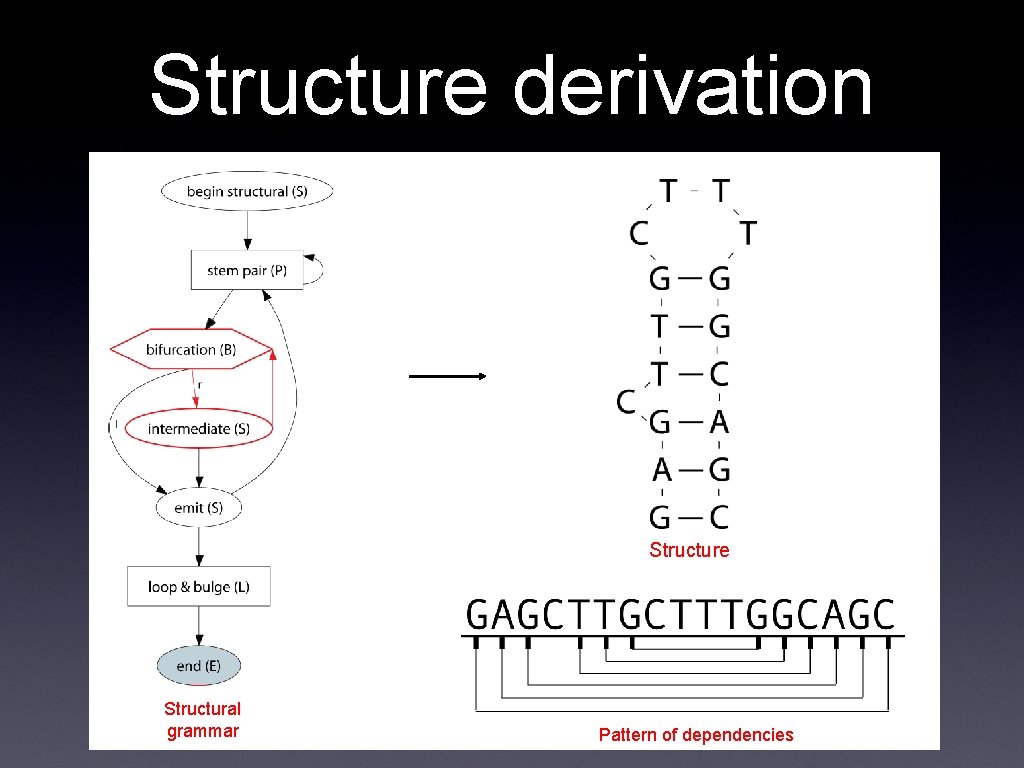
Structure derivation Structure Structural grammar Pattern of dependencies
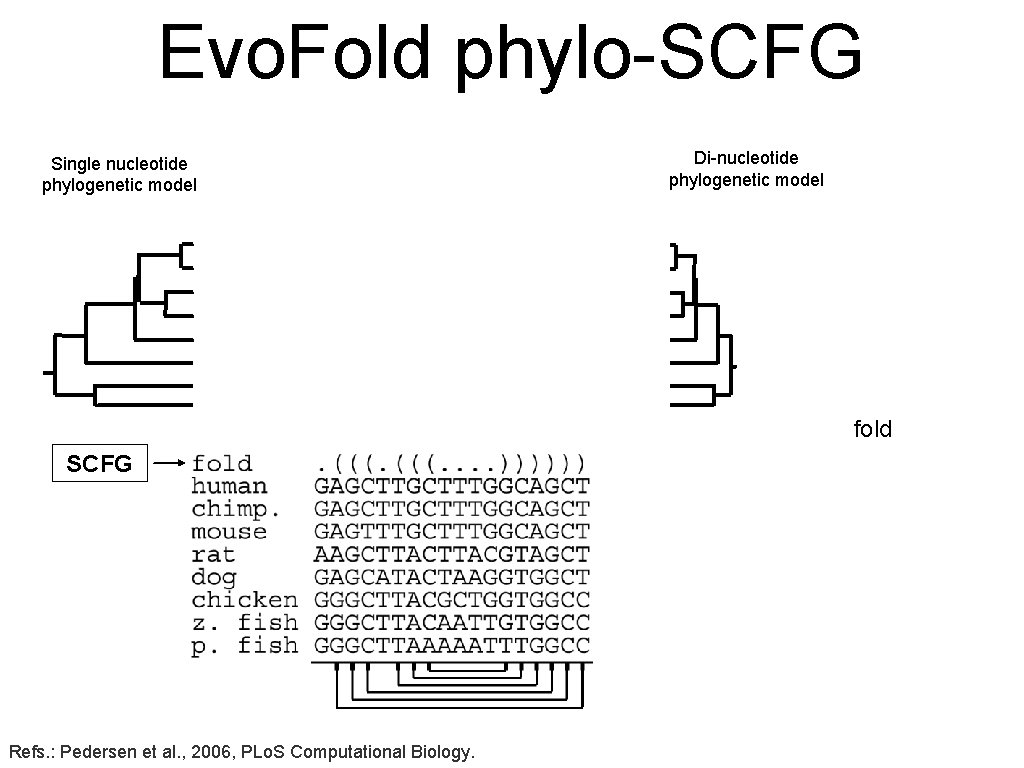
Evo. Fold phylo-SCFG Single nucleotide phylogenetic model Di-nucleotide phylogenetic model fold SCFG Refs. : Pedersen et al. , 2006, PLo. S Computational Biology.
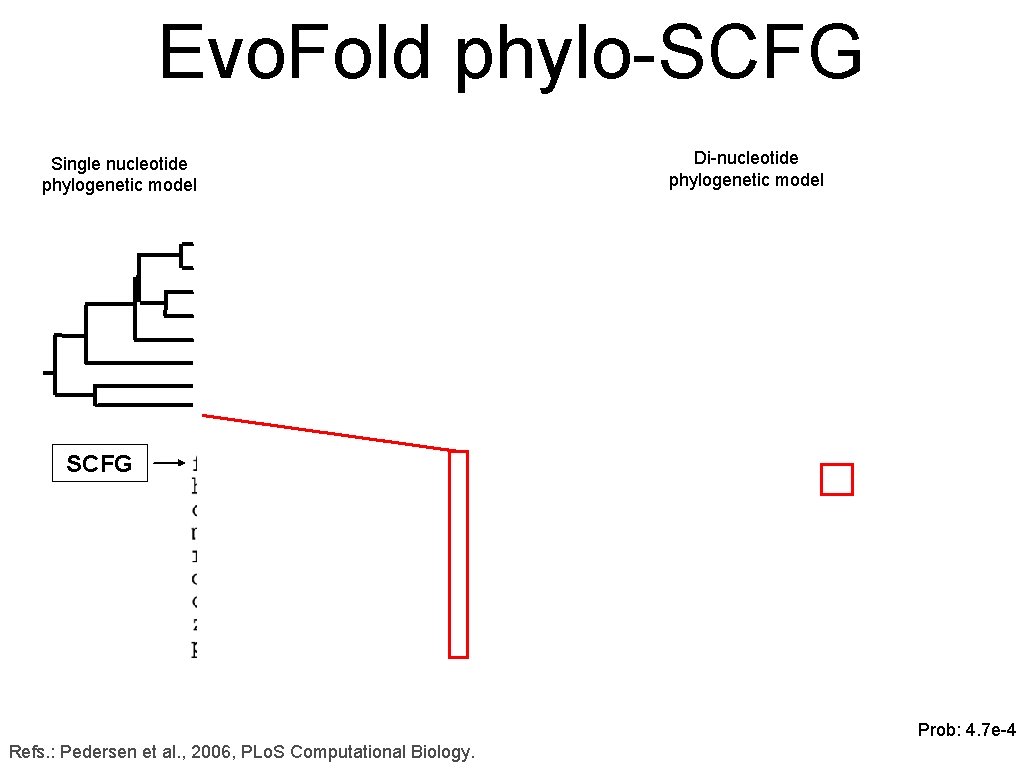
Evo. Fold phylo-SCFG Single nucleotide phylogenetic model Di-nucleotide phylogenetic model fold SCFG Prob: 4. 7 e-4 Refs. : Pedersen et al. , 2006, PLo. S Computational Biology.
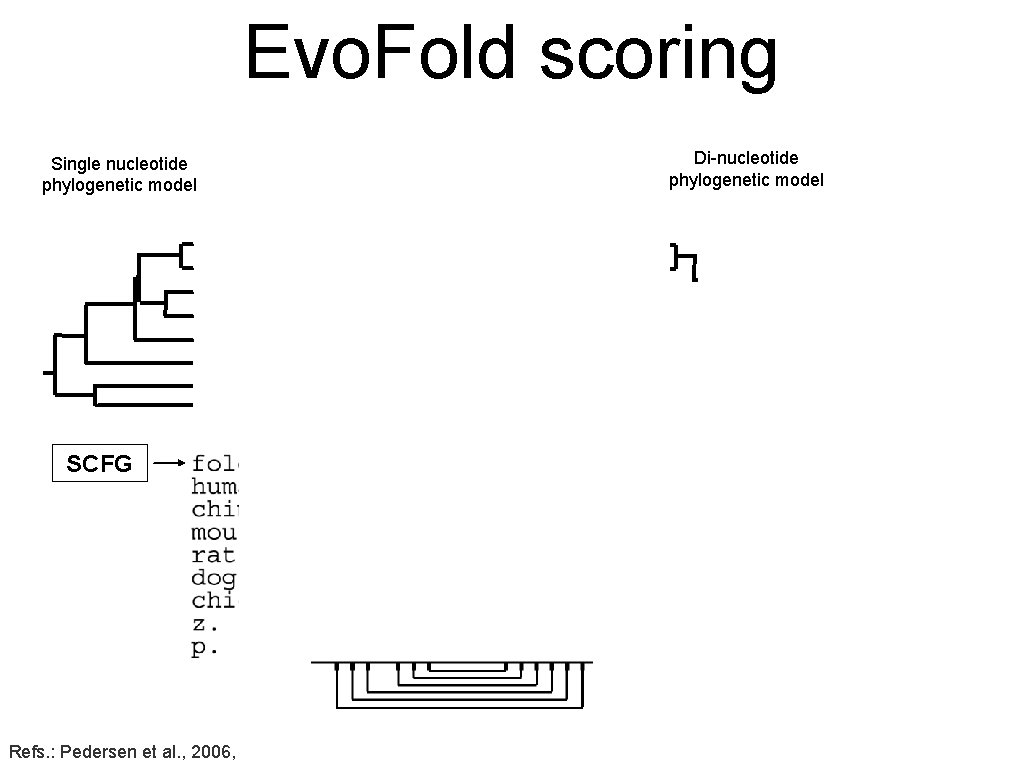
Evo. Fold scoring Single nucleotide phylogenetic model Di-nucleotide phylogenetic model fold SCFG Score: 11. 4 Refs. : Pedersen et al. , 2006, PLo. S Computational Biology.

Algorithms and training Algorithms: Traditional SCFG algorithms (CYK and insideoutside) combined with Felsenstein 81. Complexities: For an n long alignment with m sequences: Time: Space: Training of Evo. Fold: Data: Rfam structures mapped onto genomic alignments Method: EM with quasi-newton optimization

Genomic screens in vertebrates & drosophilids Input: conserved segments Output: sub-folds sub-fold ……(((((((. . ))). (((…. ))))))). ………………(((((…. ))))). . Performance Sensitivity: 43% Overall FDR: ~70% Refs. : Pedersen et al. , 2006, PLo. S Computational Biology & Stark A, Lin MF, Kheradpour P, and Pedersen JS, et al. 2007

Experimentally studied vertebrate cases HAR 1 Expression in developing neocortex Editing in Mouse Brain GABRA 3 A-to-I RNA editing substrate I/M site A Genomic c. DNA Refs: Pollard, K. S. et al. 2006 Nature & Johan Ohlson, Jakob S. Pedersen, David Haussler, and Marie Öhman. 2007 RNA.
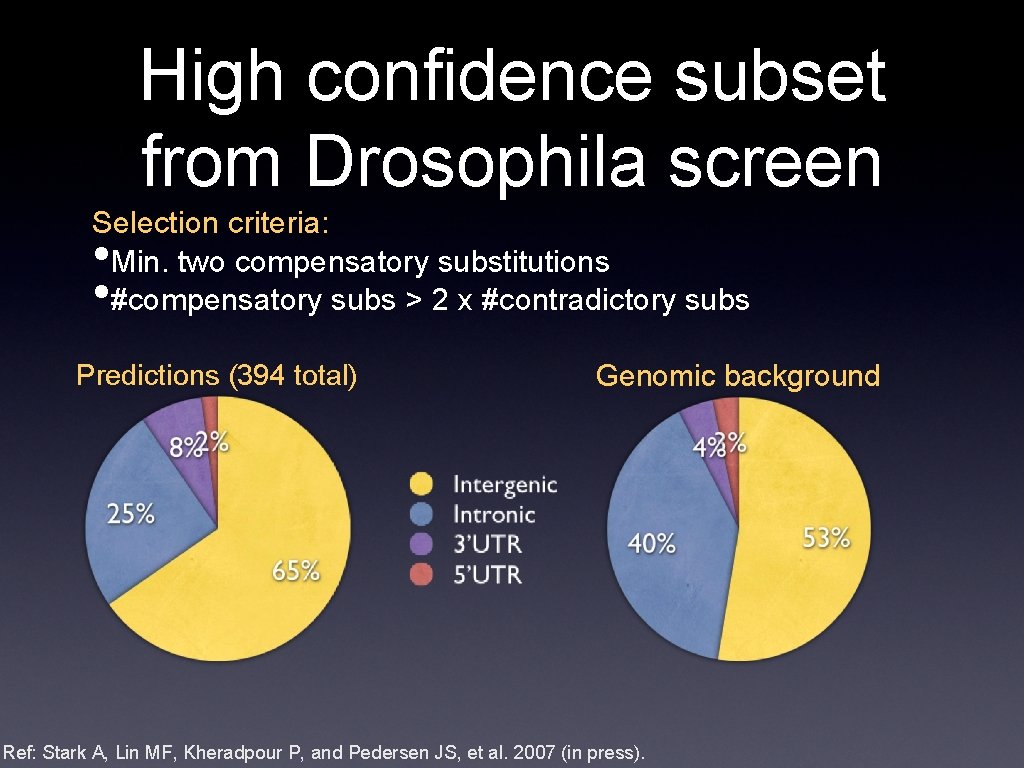
High confidence subset from Drosophila screen Selection criteria: Min. two compensatory substitutions #compensatory subs > 2 x #contradictory subs • • Predictions (394 total) Genomic background Ref: Stark A, Lin MF, Kheradpour P, and Pedersen JS, et al. 2007 (in press).

UTR structures High fraction of UTR predictions on transcribed strand (5’UTR: 80% & 3’UTR: 75%) Significant enrichment of genes regulatory roles. Gene involved in biogenesis and assembly of the ribosome (by homology to RPL 24) Ref: Stark A, Lin MF, Kheradpour P, and Pedersen JS, et al. 2007 (in press).

Acknowledgments Evo. Fold: Gill Bejerano (Stanford), Adam Siepel (Cornell), Kate Rosenbloom (UCSC), Kerstin Lindblad-Toh (Broad), Eric S. Lander (Broad), Jim Kent (UCSC), Webb Miller (Penn State), and David Haussler (UCSC) HAR 1 study: Katherine S. Pollard (UCSC), Sofie R. Salama (UCSC), Nelle Lambert (ULB), Marie-Alexandra Lambot (ULB), Sandra Coppens (ULB), Sol Katzman (UCSC), Bryan King (UCSC), Courtney Onodera (USCS), Adam Siepel (Cornell), Andrew D. Kern (UCSC), Colette Dehay (Lyon), Haller Igel (UCSC), Manuel Ares, Jr (UCSC), Pierre Vanderhaeghen (ULB) GABRA 3 study: Johan Ohlson, Marie Ohman (Stockholm University), and David Haussler (UCSC) Fly screen: Manolis Kellis (MIT), David Haussler (UCSC), Drosophila Sequencing and Analysis Consortium

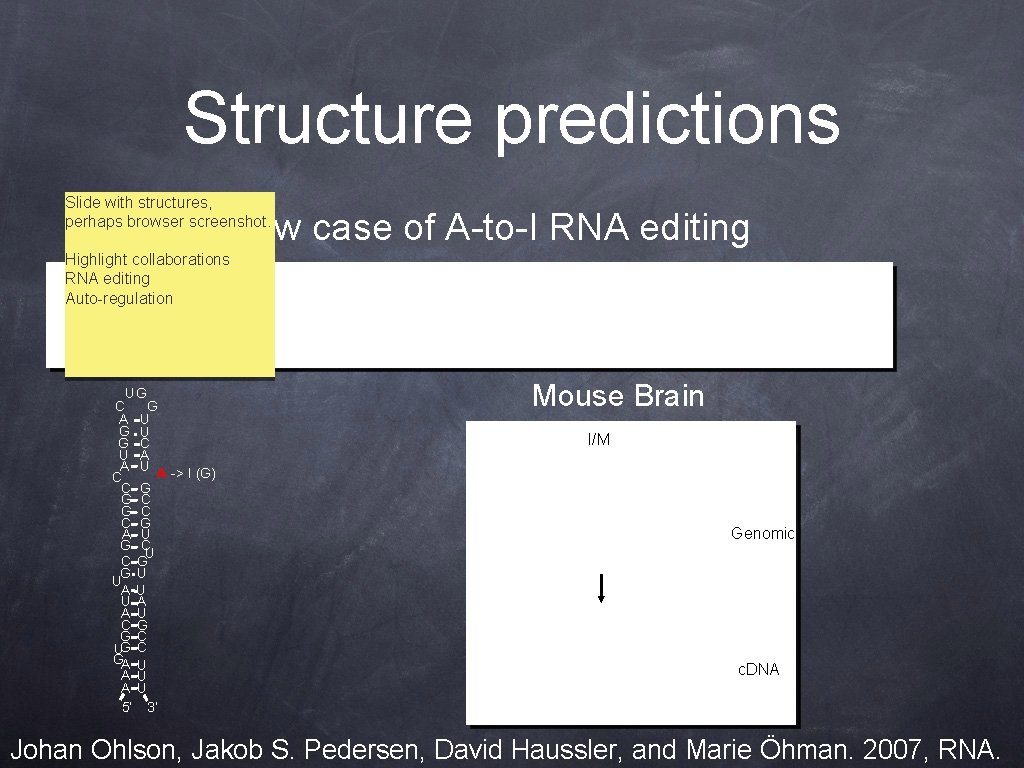
Structure predictions Slide with structures, perhaps browser screenshot. New case of A-to-I RNA editing Highlight collaborations RNA editing Auto-regulation UG C G A U G • U G C U A A U A -> I (G) C C G G C C G A U G C U C G G U U • A U U A A U C G G C UG C GA U A U 5’ 3’ Mouse Brain I/M A Genomic c. DNA Johan Ohlson, Jakob S. Pedersen, David Haussler, and Marie Öhman. 2007, RNA.

Intronic hairpin in RDL flanked by Ato-I edited exons
Spen function: Transcription co-factor and involved in neuronal cell fate, survival, and axon guidance. It has three RNA recognition motifs.

Staufen hairpin similar to known localization element in Orb Stau Orb Transport and Localization Sequence (TLS) Cohen et al, RNA, 2005.

Hairpin in highly expressed intergenic region

Structure can be extended RNAfold str. Evo. Fold str. Alignment & full structure

- Slides: 25