Genomic Characterization of Invasive Lobular Breast Carcinoma Michael
Genomic Characterization of Invasive Lobular Breast Carcinoma Michael L. Gatza, Ph. D. TCGA Breast Cancer AWG
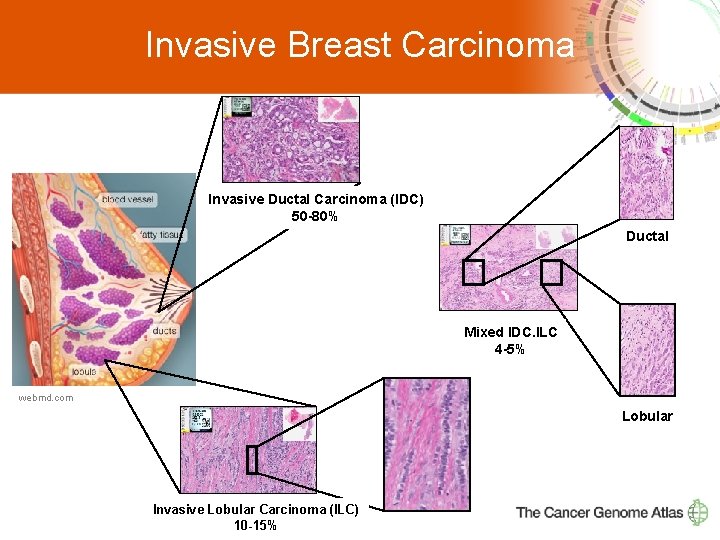
Invasive Breast Carcinoma Invasive Ductal Carcinoma (IDC) 50 -80% Ductal Mixed IDC. ILC 4 -5% webmd. com Lobular 2 Invasive Lobular Carcinoma (ILC) 10 -15%
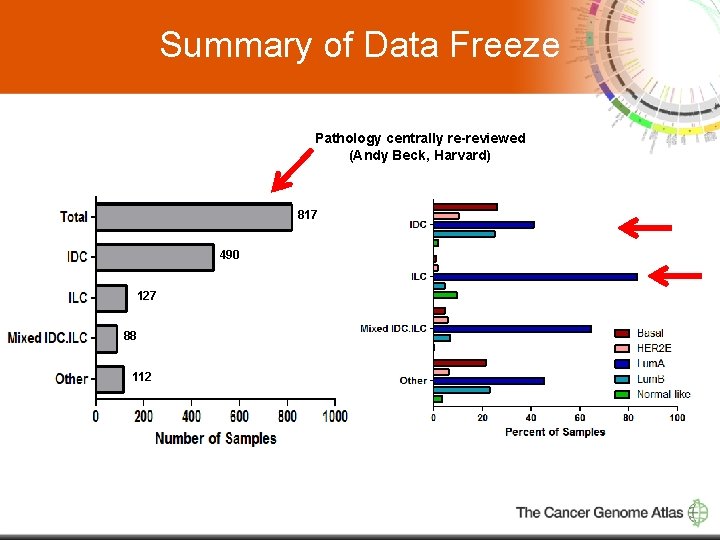
Summary of Data Freeze Pathology centrally re-reviewed (Andy Beck, Harvard) 817 490 127 88 112 3
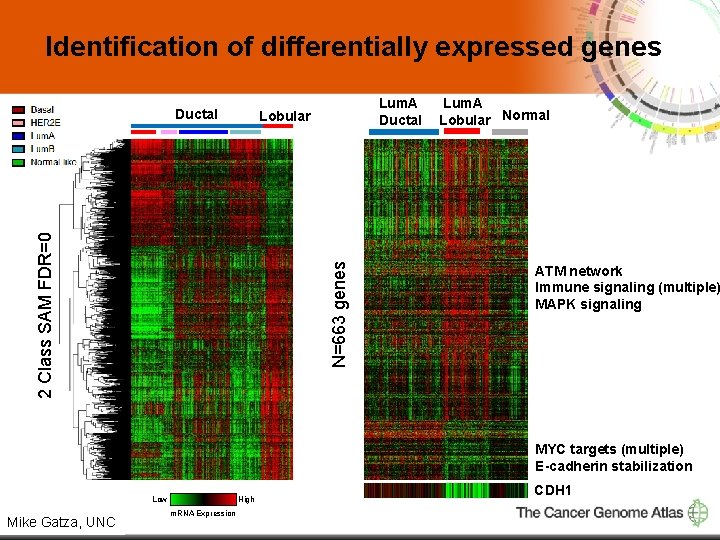
Identification of differentially expressed genes Ductal Lum. A Ductal N=663 genes 2 Class SAM FDR=0 Lobular Lum. A Lobular Normal ATM network Immune signaling (multiple) MAPK signaling MYC targets (multiple) E-cadherin stabilization Low 4 Mike Gatza, UNC High m. RNA Expression CDH 1
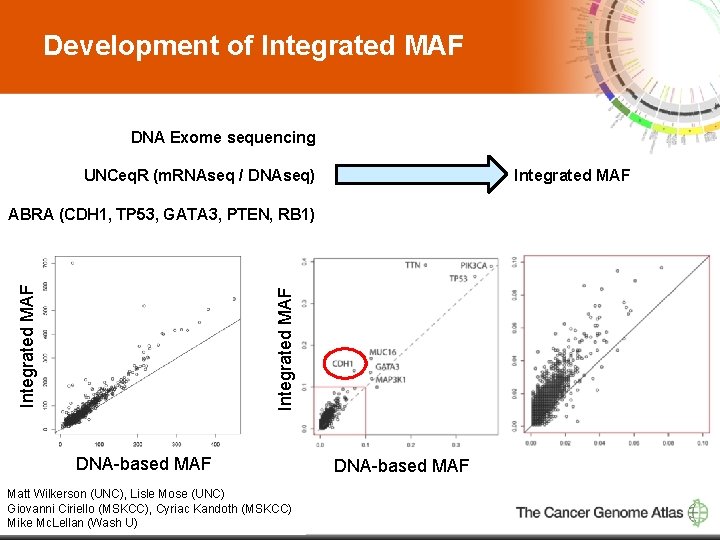
Development of Integrated MAF DNA Exome sequencing Integrated MAF UNCeq. R (m. RNAseq / DNAseq) Integrated MAF ABRA (CDH 1, TP 53, GATA 3, PTEN, RB 1) DNA-based MAF Matt Wilkerson (UNC), Lisle Mose (UNC) Giovanni Ciriello (MSKCC), Cyriac Kandoth (MSKCC) 5 Mike Mc. Lellan (Wash U) DNA-based MAF
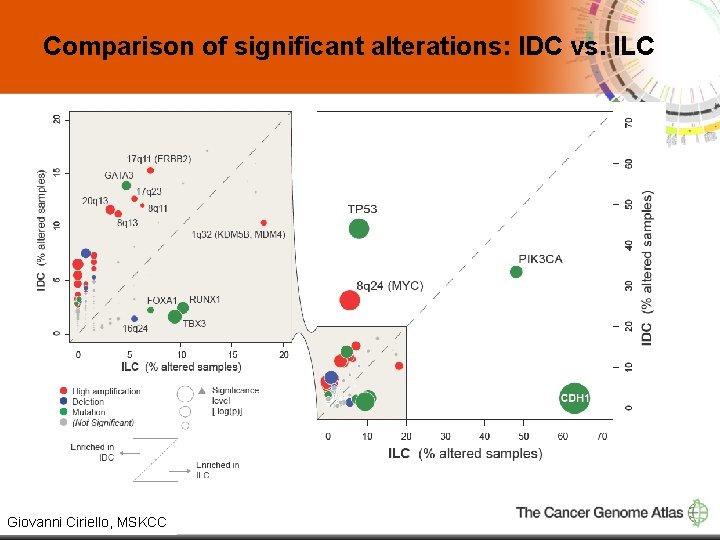
Comparison of significant alterations: IDC vs. ILC 6 Giovanni Ciriello, MSKCC
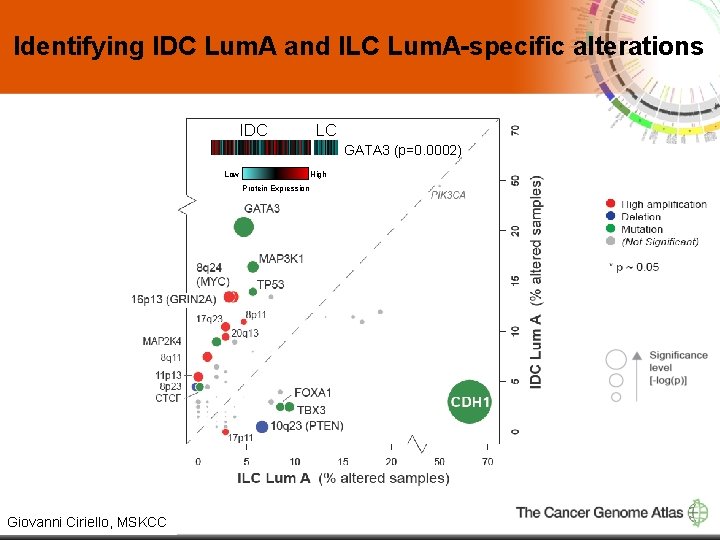
Identifying IDC Lum. A and ILC Lum. A-specific alterations IDC ILC GATA 3 (p=0. 0002) Low High Protein Expression 7 Giovanni Ciriello, MSKCC
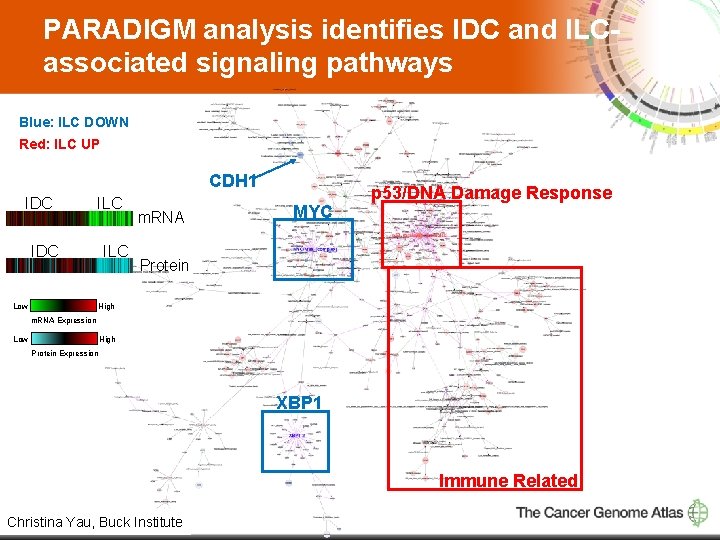
PARADIGM analysis identifies IDC and ILCassociated signaling pathways Blue: ILC DOWN Red: ILC UP CDH 1 IDC ILC IDC Low ILC m. RNA MYC p 53/DNA Damage Response Protein High m. RNA Expression Low High Protein Expression XBP 1 Immune Related 8 Christina Yau, Buck Institute
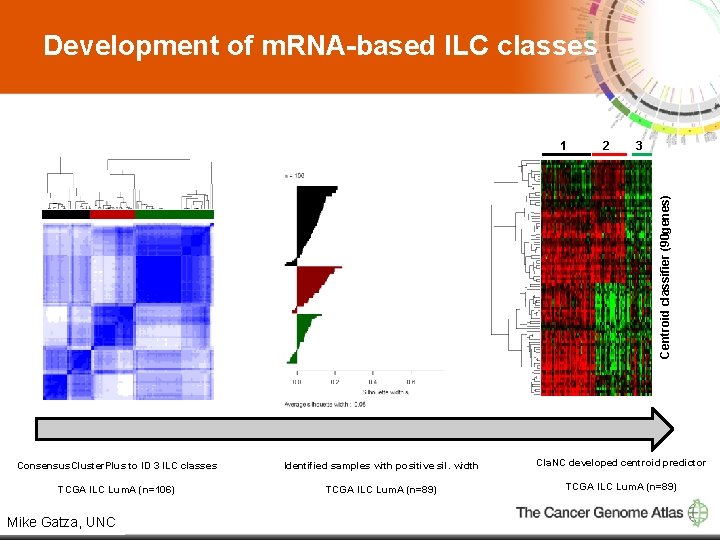
Development of m. RNA-based ILC classes 2 3 Centroid classifier (90 genes) 1 Consensus. Cluster. Plus to ID 3 ILC classes Identified samples with positive sil. width Cla. NC developed centroid predictor TCGA ILC Lum. A (n=106) TCGA ILC Lum. A (n=89) 9 Mike Gatza, UNC
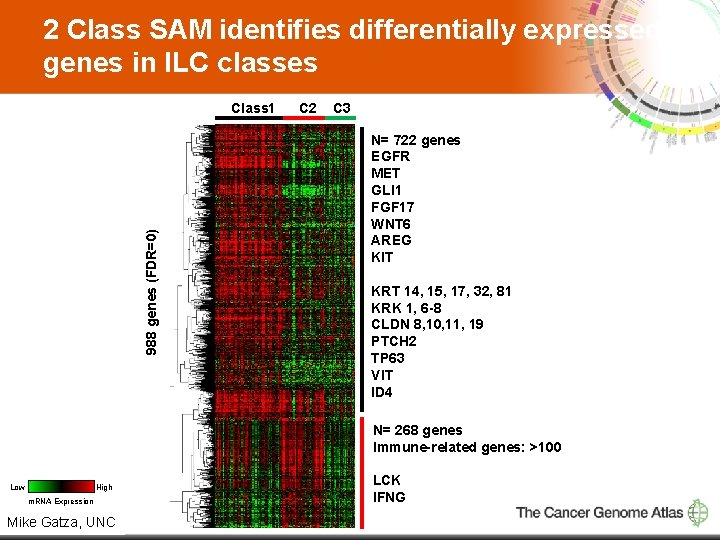
2 Class SAM identifies differentially expressed genes in ILC classes 988 genes (FDR=0) Class 1 C 2 C 3 N= 722 genes EGFR MET GLI 1 FGF 17 WNT 6 AREG KIT KRT 14, 15, 17, 32, 81 KRK 1, 6 -8 CLDN 8, 10, 11, 19 PTCH 2 TP 63 VIT ID 4 N= 268 genes Immune-related genes: >100 Low High m. RNA Expression 10 Mike Gatza, UNC LCK IFNG
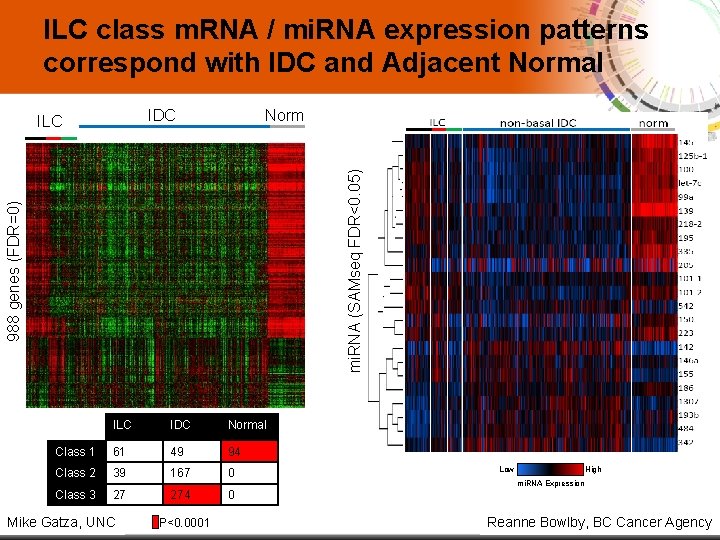
ILC class m. RNA / mi. RNA expression patterns correspond with IDC and Adjacent Normal IDC Norm 988 genes (FDR=0) mi. RNA (SAMseq FDR<0. 05) ILC Low ILC IDC High Normal m. RNA Expression Class 1 61 49 94 Class 2 39 167 0 Class 3 27 274 0 11 Mike Gatza, UNC P<0. 0001 Low High mi. RNA Expression Reanne Bowlby, BC Cancer Agency
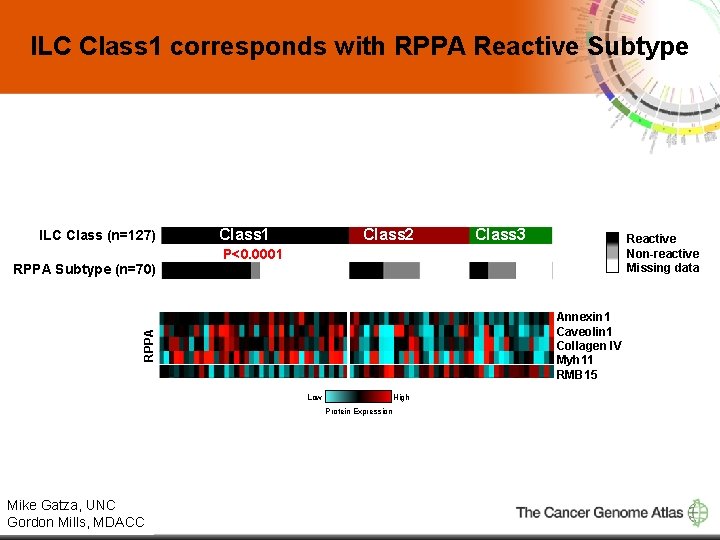
ILC Class 1 corresponds with RPPA Reactive Subtype ILC Class (n=127) Class 2 Reactive Non-reactive Missing data Annexin 1 Caveolin 1 Collagen IV Myh 11 RMB 15 Low High Protein Expression Mike Gatza, UNC 12 Gordon Mills, MDACC Class 3 P<0. 0001 RPPA Subtype (n=70) Class 1
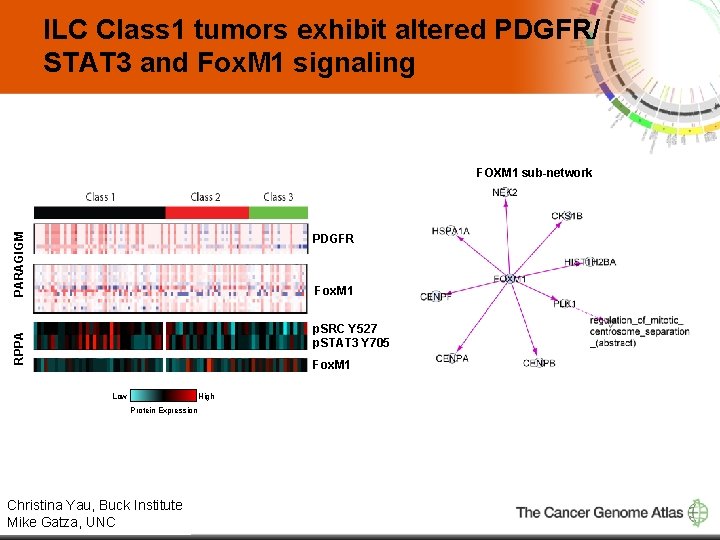
ILC Class 1 tumors exhibit altered PDGFR/ STAT 3 and Fox. M 1 signaling PARAGIGM FOXM 1 sub-network PDGFR Fox. M 1 RPPA p. SRC Y 527 p. STAT 3 Y 705 Fox. M 1 Low High Protein Expression Christina Yau, Buck Institute 13 Mike Gatza, UNC
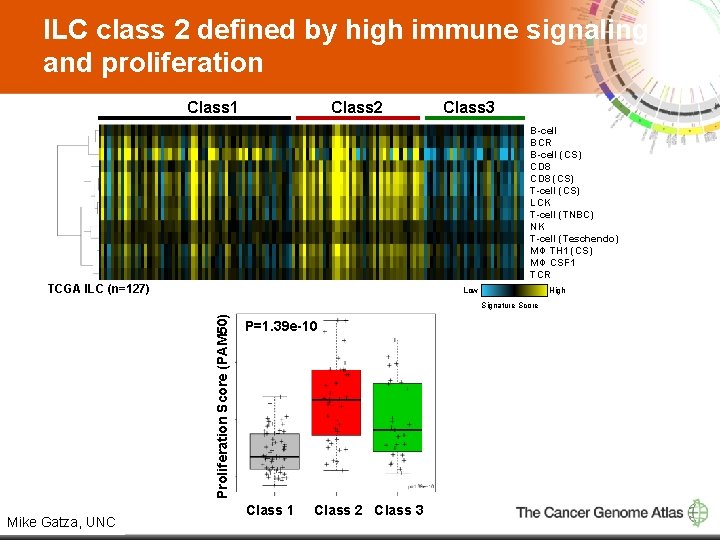
ILC class 2 defined by high immune signaling and proliferation Class 1 Class 2 Class 3 B-cell BCR B-cell (CS) CD 8 (CS) T-cell (CS) LCK T-cell (TNBC) NK T-cell (Teschendo) MΦ TH 1 (CS) MΦ CSF 1 TCR TCGA ILC (n=127) Low High Proliferation Score (PAM 50) Signature Score 14 Mike Gatza, UNC P=1. 39 e-10 Class 1 Class 2 Class 3
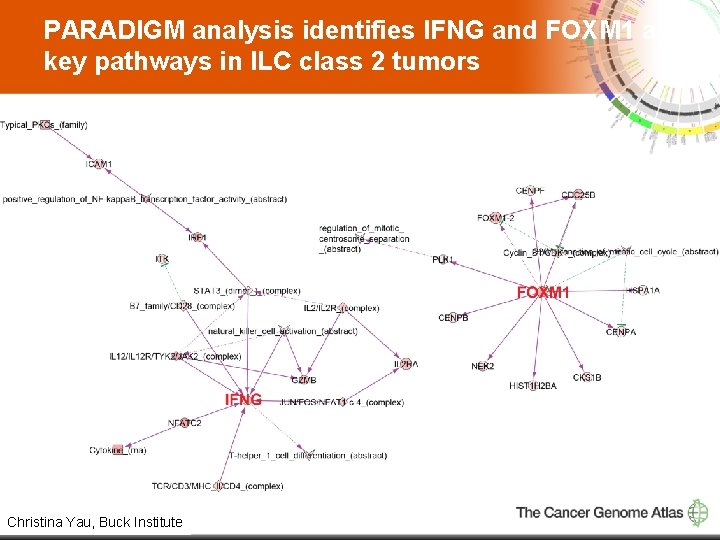
PARADIGM analysis identifies IFNG and FOXM 1 as key pathways in ILC class 2 tumors 15 Christina Yau, Buck Institute

Summary • Developed unique integrated MAF utilizing both DNA exome and m. RNA sequencing • ILC vs. IDC – – • FOXA 1, CDH 1 mutations associated with ILC GATA 3 mutation associated with IDC Altered signaling: CDH 1, Myc, p 53/DNA damage, immune signaling Identified differentially expressed mi. RNA and methylation ILC classes – Class 1 associated with Reactive subtype – Class 2 immune component and highly proliferative 16

TCGA Breast Cancer Analysis Working Group Baylor College of Medicine Chad Creighton Xiaosong Wang British Columbia Cancer Agency Andy Chu Elizabeth Chun Andy Mungall Gordon Robertson Dominik Stoll Broad Institute Andrew Cherniack Greater Poland Cancer Center Maciej Wiznerowicz Harvard Medical School Terrence Wu Yonghong Xiao Institute for Systems Biology Sheila Reynolds Ilya Shmulevich Lawrence Berkeley National Laboratory Paul Spellman Mayo Clinic Jim Ingle 17 The University of Texas MD Anderson Cancer Center Roel Verhaak Rehan Akbani Nancy Shih Gordon Mills Memorial Sloan-Kettering Cancer Center Giovanni Ciriello Niki Schultz Ethan Cerami Arthur Goldberg Caitlin Byrne Anders Jacobsen Tari King Chris Sander Nationwide Children’s Hospital Jay Bowen Julie Gastier-Foster National Cancer Institute Chunhua Yan John Demchok Laura Dillon Margi Sheth Peter Good Jacqueline Palchik Heidi Sofia Kenna Shaw University of California Santa Cruz Buck Institute Chris Benz David Haussler Christina Yau Sam Ng Ted Goldstein Kyle Ellrott Charlie Vaske Josh Stuart Jing Zhu University of California, San Francisco Fred Waldman University of Southern California Peter Laird Swapna Mahurkar Simeen Malik Dan Weisenberger Windber Research Institute Hai Hu Richard Mural University of North Carolina Chuck Perou (co-chair) Katie Hoadley Mike Gatza Joel Parker Xiaping He Michael Iglesia Grace Silva Wei Zhao The Genome Institute at Washington University Matthew Ellis (co-chair) Li Ding Lucinda Fulton Daniel Koboldt Elaine Mardis
- Slides: 17