Genomes Organism Estimated size in bases Estimated gene
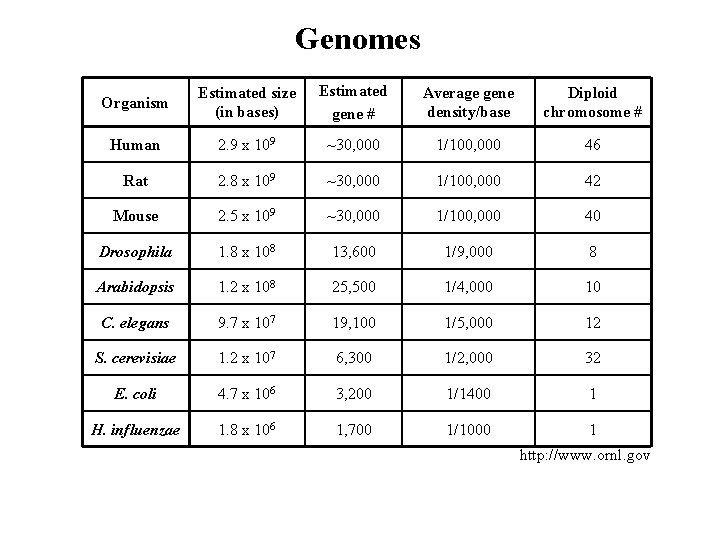
Genomes Organism Estimated size (in bases) Estimated gene # Average gene density/base Diploid chromosome # Human 2. 9 x 109 ~30, 000 1/100, 000 46 Rat 2. 8 x 109 ~30, 000 1/100, 000 42 Mouse 2. 5 x 109 ~30, 000 1/100, 000 40 Drosophila 1. 8 x 108 13, 600 1/9, 000 8 Arabidopsis 1. 2 x 108 25, 500 1/4, 000 10 C. elegans 9. 7 x 107 19, 100 1/5, 000 12 S. cerevisiae 1. 2 x 107 6, 300 1/2, 000 32 E. coli 4. 7 x 106 3, 200 1/1400 1 H. influenzae 1. 8 x 106 1, 700 1/1000 1 http: //www. ornl. gov
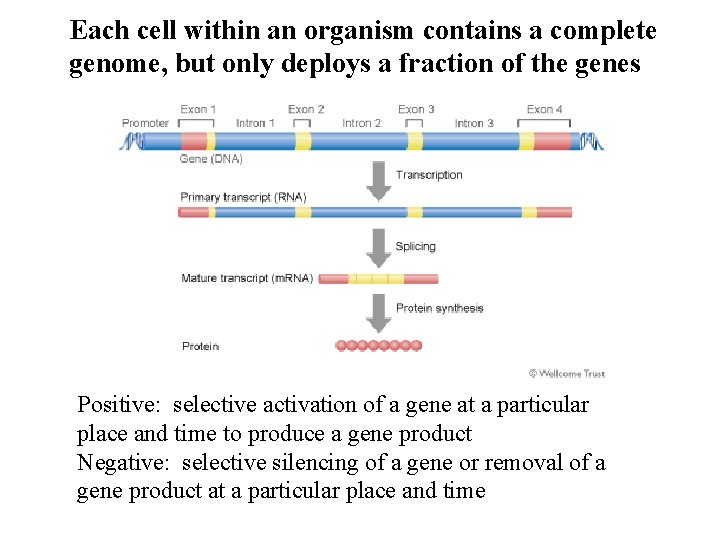
Each cell within an organism contains a complete genome, but only deploys a fraction of the genes Positive: selective activation of a gene at a particular place and time to produce a gene product Negative: selective silencing of a gene or removal of a gene product at a particular place and time
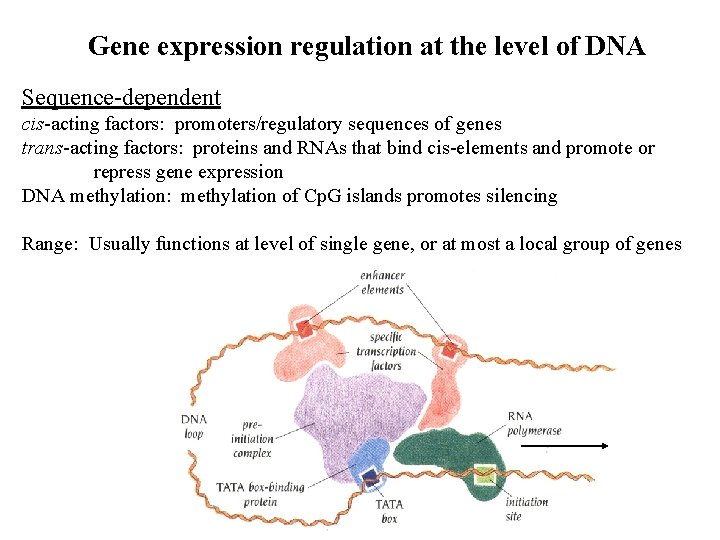
Gene expression regulation at the level of DNA Sequence-dependent cis-acting factors: promoters/regulatory sequences of genes trans-acting factors: proteins and RNAs that bind cis-elements and promote or repress gene expression DNA methylation: methylation of Cp. G islands promotes silencing Range: Usually functions at level of single gene, or at most a local group of genes
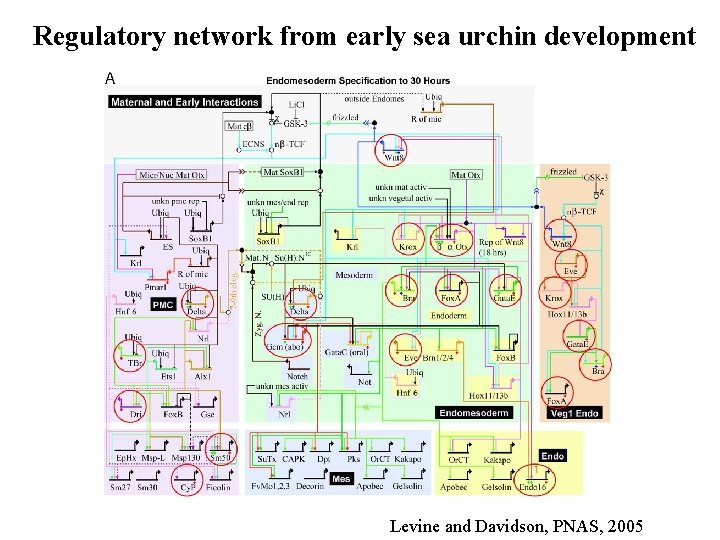
Regulatory network from early sea urchin development Levine and Davidson, PNAS, 2005
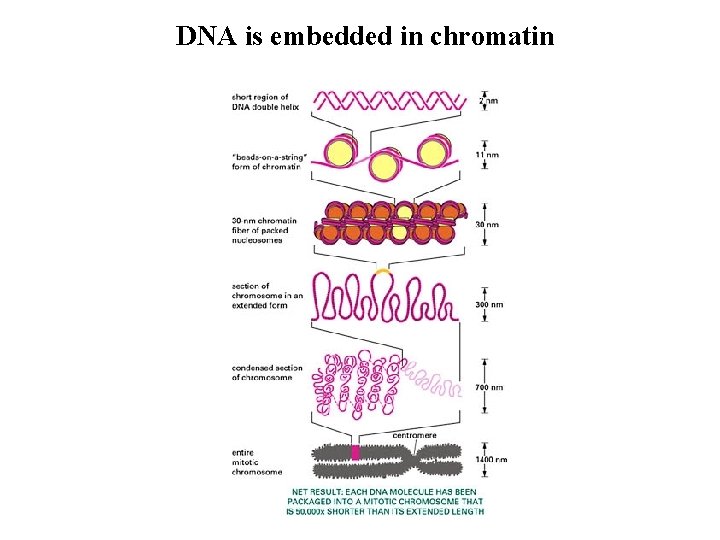
DNA is embedded in chromatin
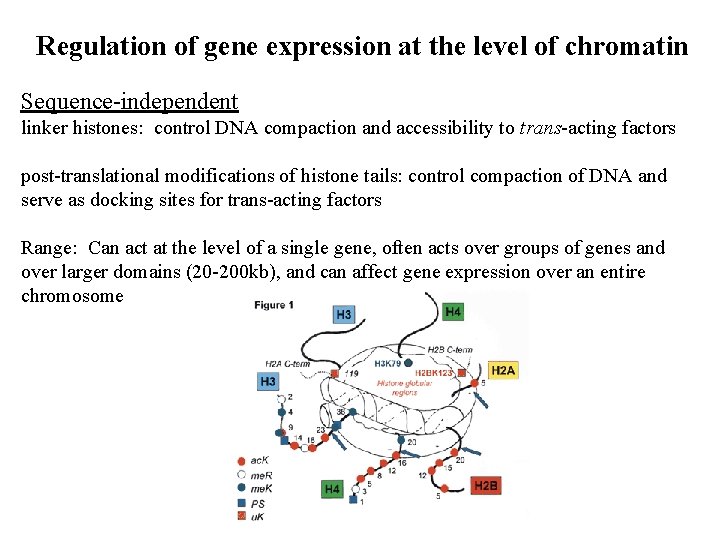
Regulation of gene expression at the level of chromatin Sequence-independent linker histones: control DNA compaction and accessibility to trans-acting factors post-translational modifications of histone tails: control compaction of DNA and serve as docking sites for trans-acting factors Range: Can act at the level of a single gene, often acts over groups of genes and over larger domains (20 -200 kb), and can affect gene expression over an entire chromosome
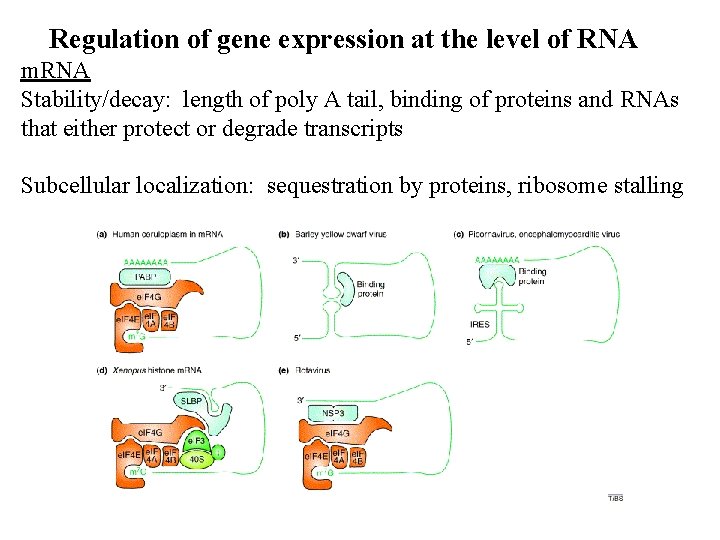
Regulation of gene expression at the level of RNA m. RNA Stability/decay: length of poly A tail, binding of proteins and RNAs that either protect or degrade transcripts Subcellular localization: sequestration by proteins, ribosome stalling
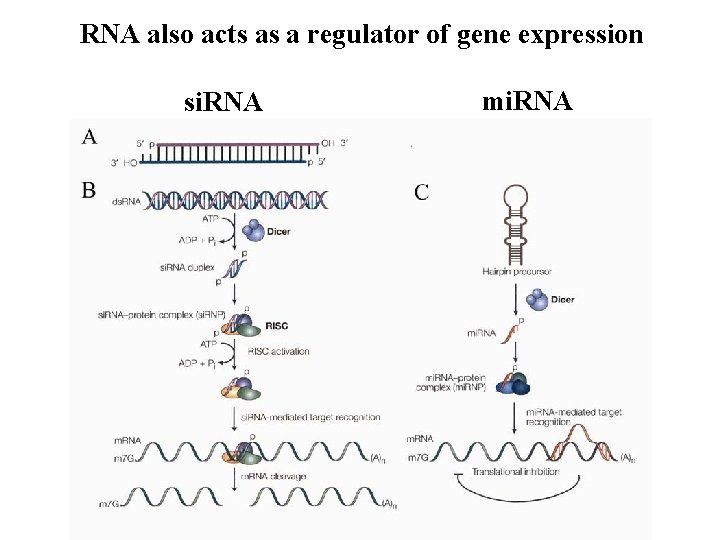
RNA also acts as a regulator of gene expression si. RNA mi. RNA
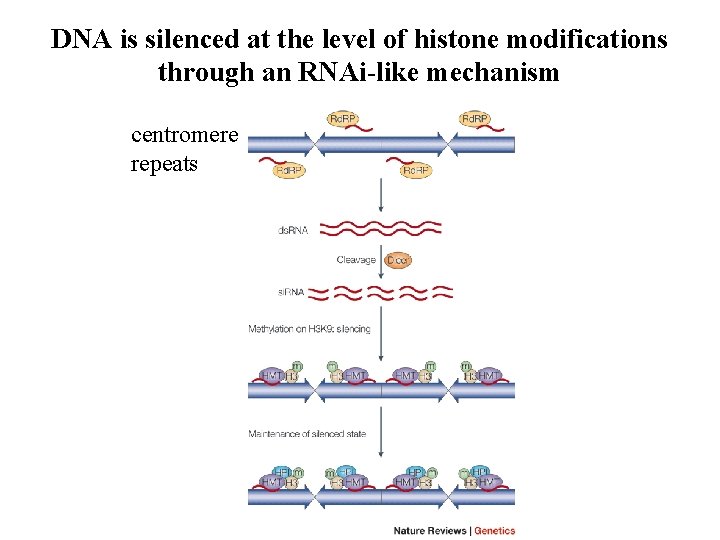
DNA is silenced at the level of histone modifications through an RNAi-like mechanism centromere repeats
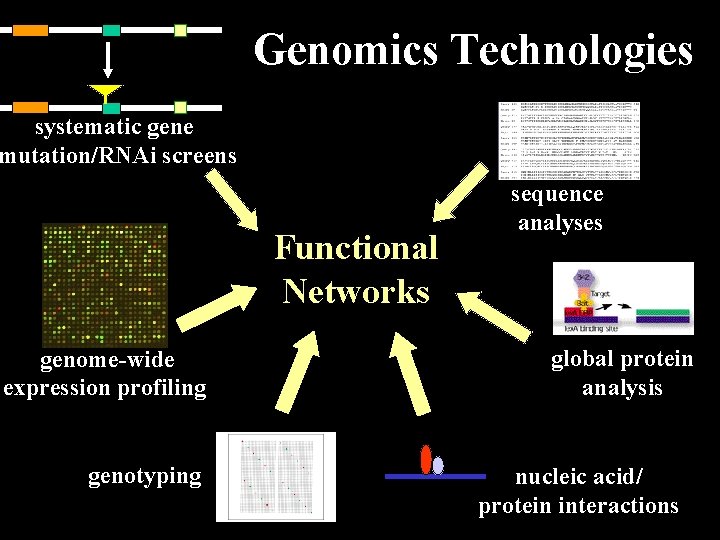
Genomics Technologies systematic gene mutation/RNAi screens Functional Networks genome-wide expression profiling genotyping sequence analyses global protein analysis nucleic acid/ protein interactions
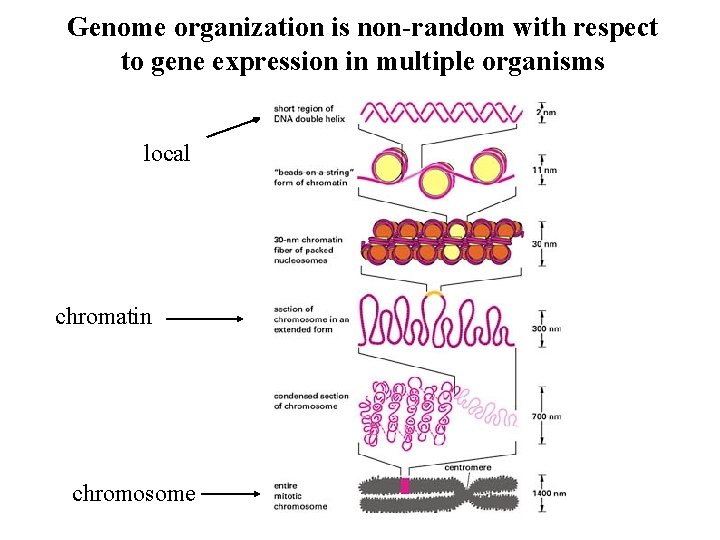
Genome organization is non-random with respect to gene expression in multiple organisms local chromatin chromosome
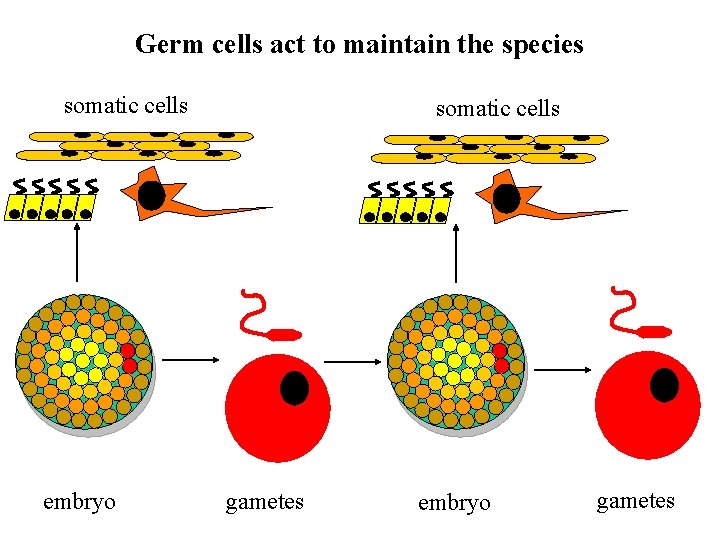
Germ cells act to maintain the species somatic cells embryo somatic cells gametes embryo gametes
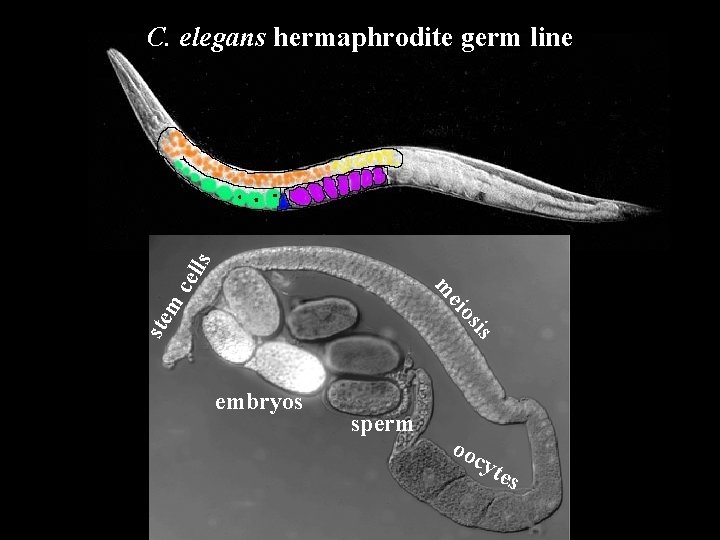
sis ste m eio m cel ls C. elegans hermaphrodite germ line embryos sperm oo cyt es
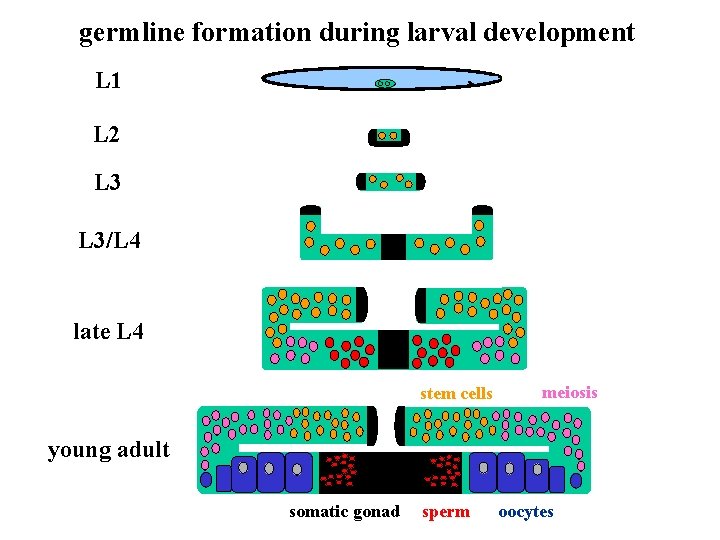
germline formation during larval development L 1 L 2 L 3/L 4 late L 4 stem cells meiosis young adult somatic gonad sperm oocytes
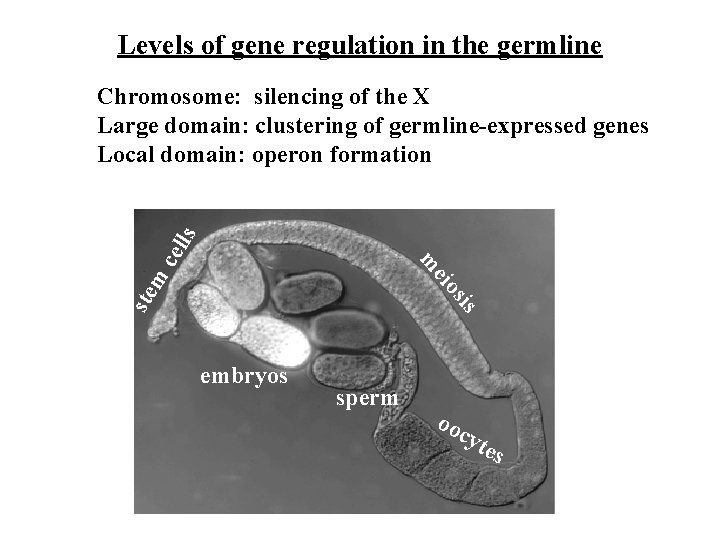
Levels of gene regulation in the germline sis ste m eio m cel ls Chromosome: silencing of the X Large domain: clustering of germline-expressed genes Local domain: operon formation embryos sperm oo cyt es
C. elegans DNA microarrays ~20, 000 genes in the worm genome ~18, 000 genes on the array
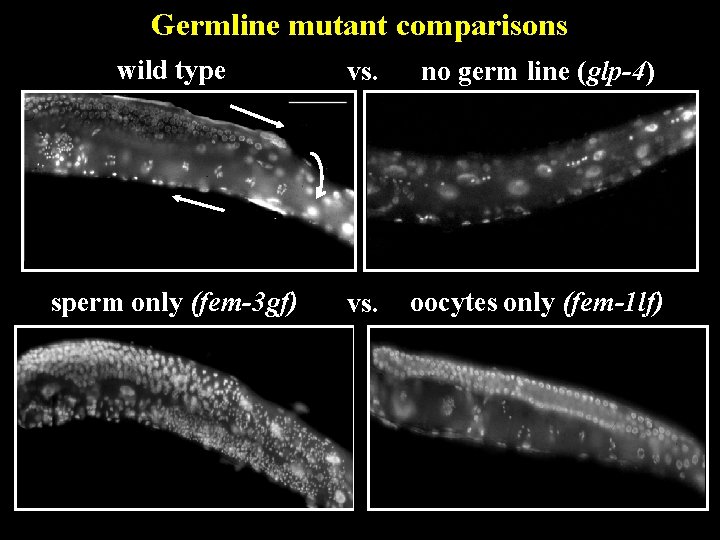
Germline mutant comparisons wild type vs. no germ line (glp-4) sperm only (fem-3 gf) vs. oocytes only (fem-1 lf)
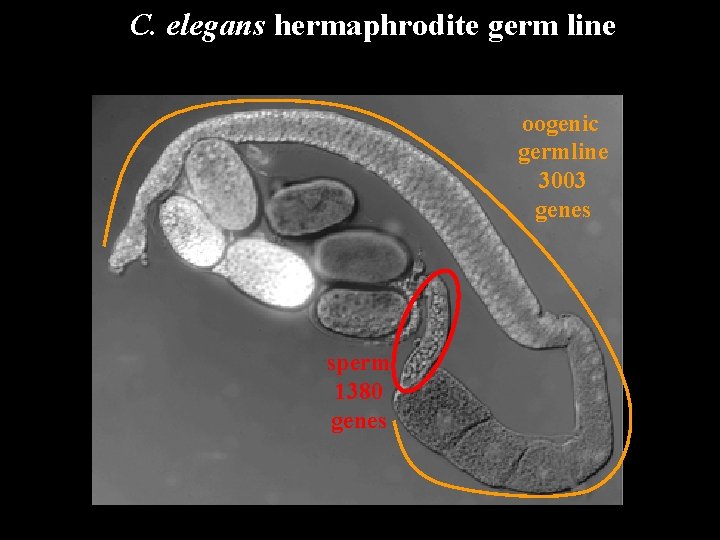
C. elegans hermaphrodite germ line oogenic germline 3003 genes sperm 1380 genes
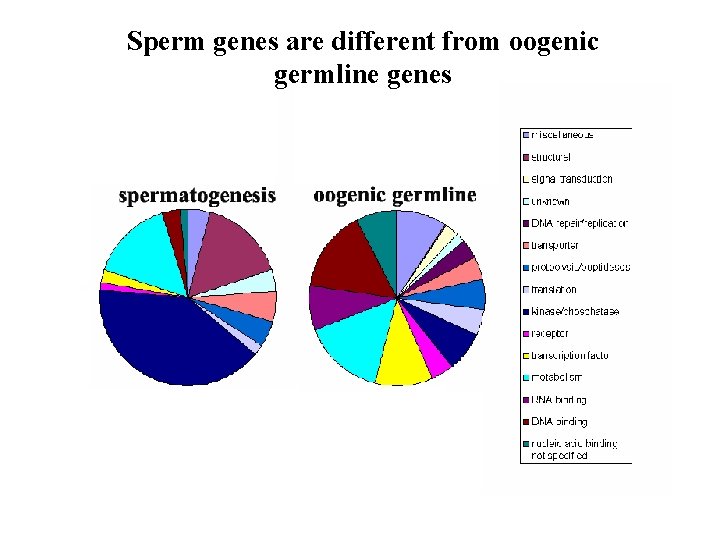
Sperm genes are different from oogenic germline genes
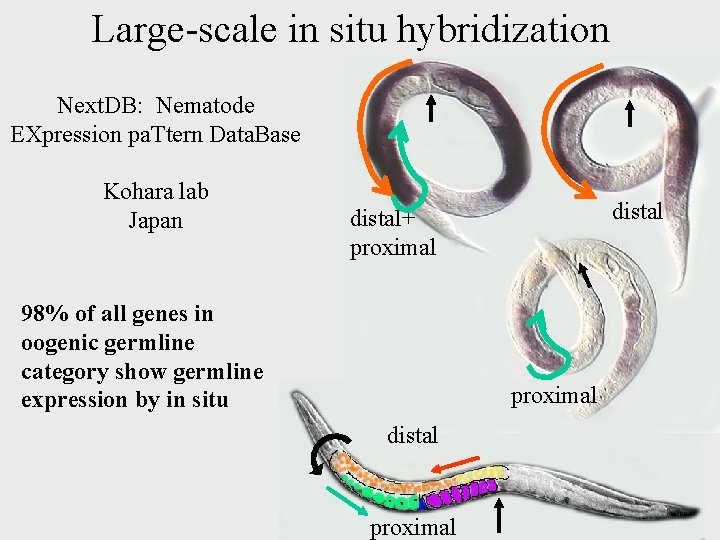
Large-scale in situ hybridization Next. DB: Nematode EXpression pa. Ttern Data. Base Kohara lab Japan distal+ proximal 98% of all genes in oogenic germline category show germline expression by in situ proximal distal proximal
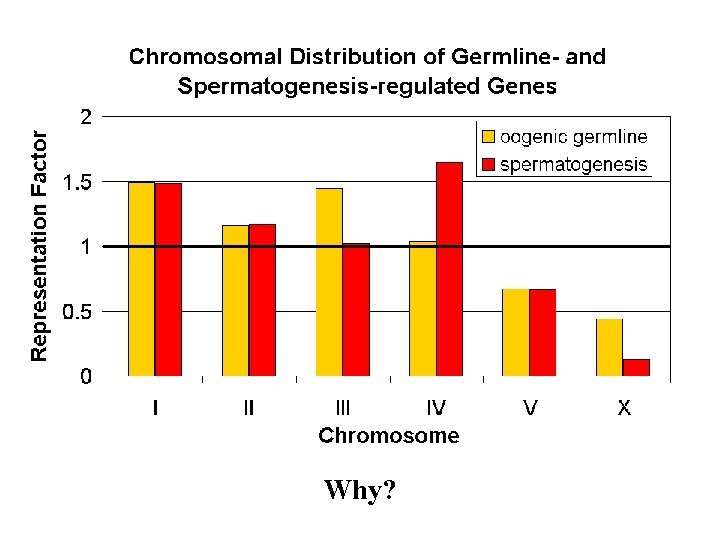
Why?
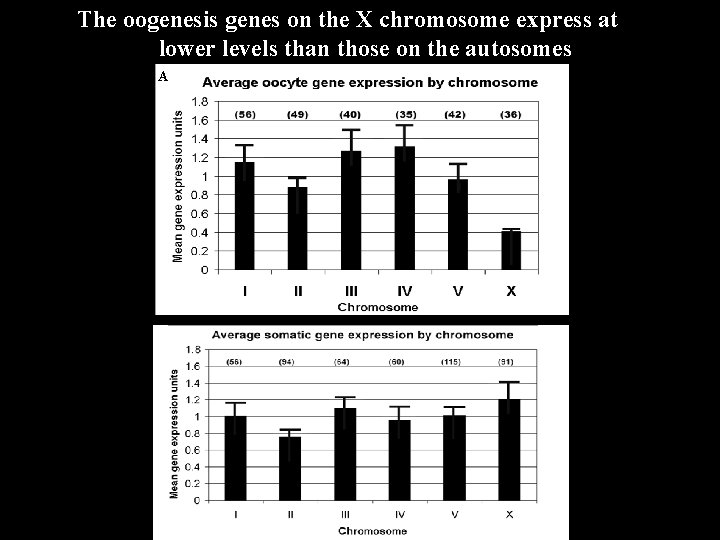
The oogenesis genes on the X chromosome express at lower levels than those on the autosomes A B
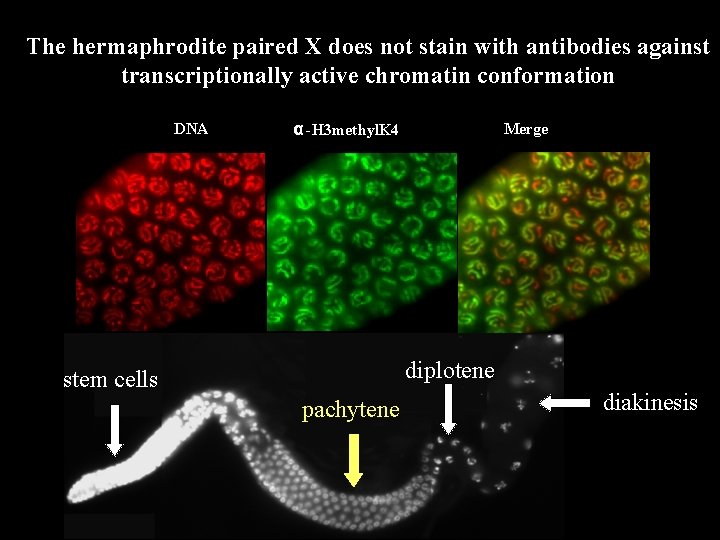
The hermaphrodite paired X does not stain with antibodies against transcriptionally active chromatin conformation DNA α -H 3 methyl. K 4 Merge diplotene stem cells pachytene diakinesis
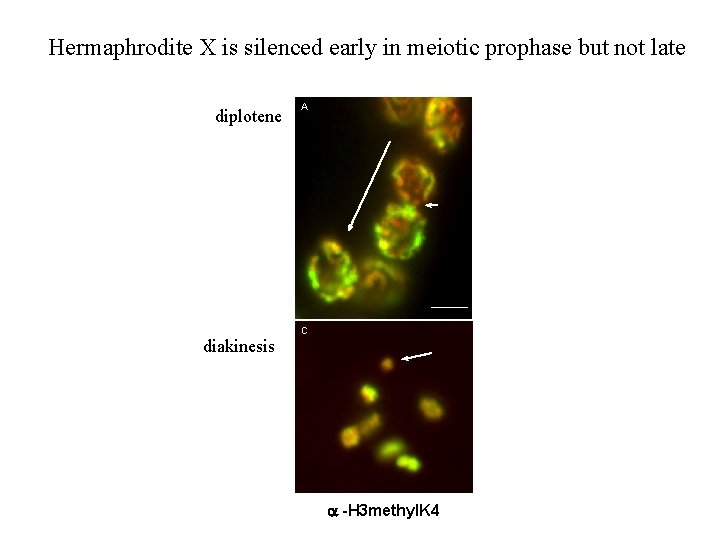
Hermaphrodite X is silenced early in meiotic prophase but not late diplotene A diakines inactive B Transgene 1 2 5 6 3 4 C D diakinesis -H 3 methyl. K 4 diakines active tra
- Slides: 25