Genomes and Their Evolution AP Biology Ch 18

Genomes and Their Evolution AP Biology Ch 18 CVHS

Vast stores of Biological Information • Genomics: the study of whole sets of genes and their interactions. • Proteomics: Study of proteins and their functions (more useful) • Bioinformatics: Computation method of storing and analyzing biological information
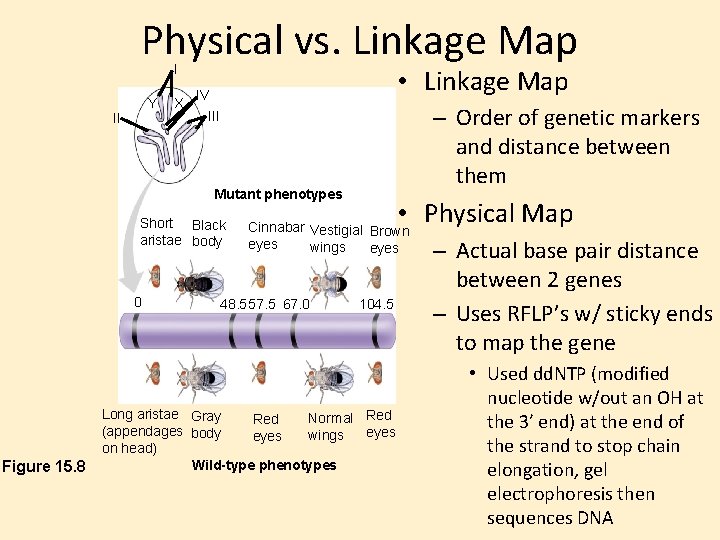
Physical vs. Linkage Map I Y II X • Linkage Map IV – Order of genetic markers and distance between them III Mutant phenotypes Short Black aristae body 0 Figure 15. 8 • Physical Map Cinnabar Vestigial Brown eyes wings eyes 48. 557. 5 67. 0 104. 5 Long aristae Gray Normal Red (appendages body eyes wings eyes on head) Wild-type phenotypes – Actual base pair distance between 2 genes – Uses RFLP’s w/ sticky ends to map the gene • Used dd. NTP (modified nucleotide w/out an OH at the 3’ end) at the end of the strand to stop chain elongation, gel electrophoresis then sequences DNA
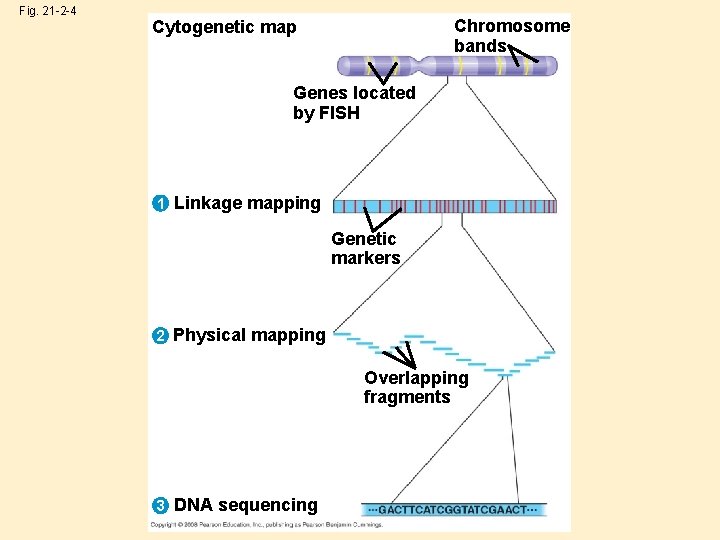
Fig. 21 -2 -4 Chromosome bands Cytogenetic map Genes located by FISH 1 Linkage mapping Genetic markers 2 Physical mapping Overlapping fragments 3 DNA sequencing

Shotgun Sequencing • Cut DNA from multiple copies of a chromosome • Clone the DNA fragments (simply to increase #) • Sequence each fragment using dd. NTP • Computer uses sticky ends and base pairing rules to order the DNA
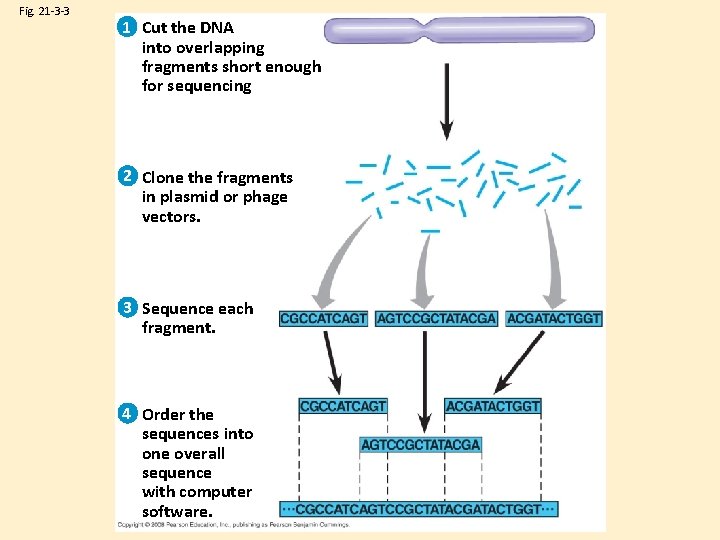
Fig. 21 -3 -3 1 Cut the DNA into overlapping fragments short enough for sequencing 2 Clone the fragments in plasmid or phage vectors. 3 Sequence each fragment. 4 Order the sequences into one overall sequence with computer software.

National Data-Bases & Uses • National Center for Biotechnology Information. mht • NCBI has a program called BLAST that allows anyone to compare DNA sequences. You can go home and do this. • NIH is creating a data-base for comparing normal cells to 3 types of cancer cells – Ovarian – Glioblastoma of the brain – Lung cancer

How do we determine the function of New Genes that are discovered? • 1. Match a known gene, therefore functions similarly • 2. New sequence may be similar to a known gene whose function is unknown & will help us learn the function of the other • 3. Completely unknown: use protein coded for to help determine function
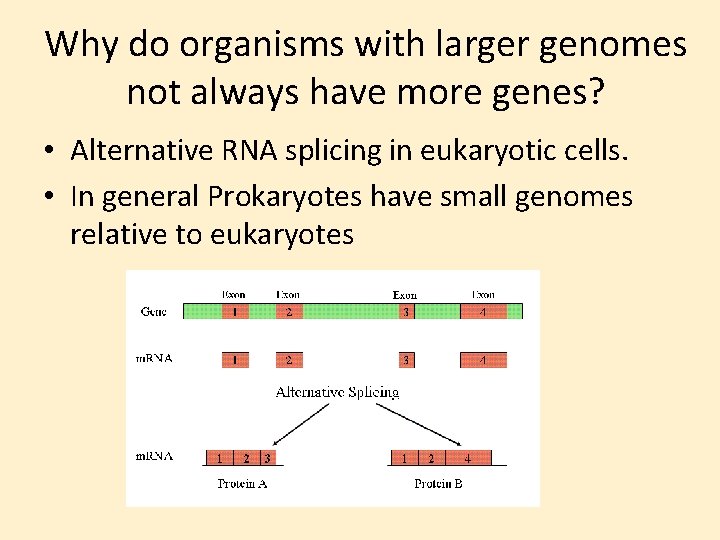
Why do organisms with larger genomes not always have more genes? • Alternative RNA splicing in eukaryotic cells. • In general Prokaryotes have small genomes relative to eukaryotes

Human DNA types
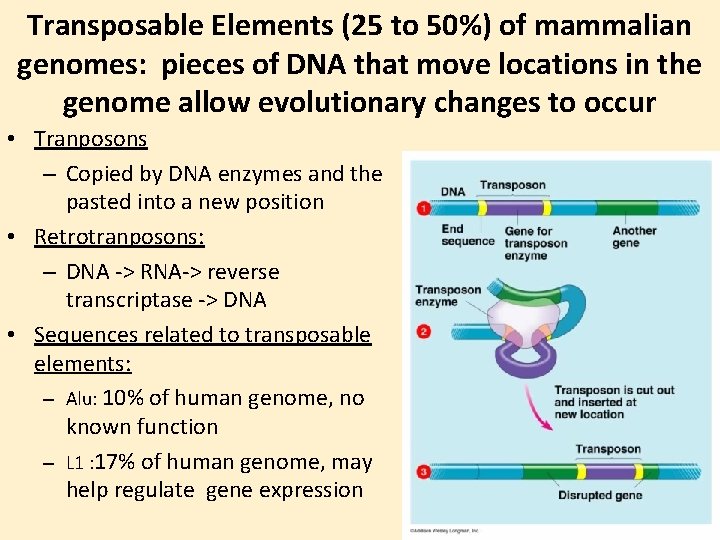
Transposable Elements (25 to 50%) of mammalian genomes: pieces of DNA that move locations in the genome allow evolutionary changes to occur • Tranposons – Copied by DNA enzymes and the pasted into a new position • Retrotranposons: – DNA -> RNA-> reverse transcriptase -> DNA • Sequences related to transposable elements: – Alu: 10% of human genome, no known function – L 1 : 17% of human genome, may help regulate gene expression
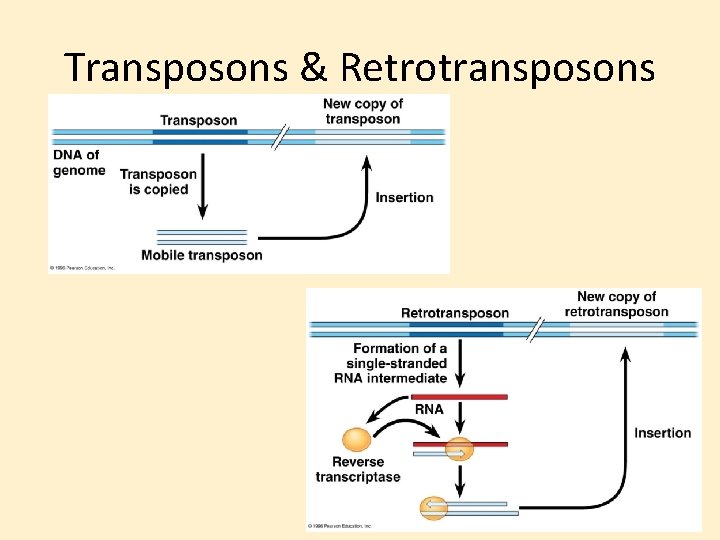
Transposons & Retrotransposons

Simple Sequence DNA & Short Tandem Repeats • Simple Sequence DNA: short sequences that are tandemly repeated • STR: a repetitive unit of DNA w/ 2 to 5 nucleotides • Used in forensic science: the # of STR’s on a particular gene varies greatly between individuals, PCR and Gel Electrophoresis can be used to determine if an individual was present at a crime scene
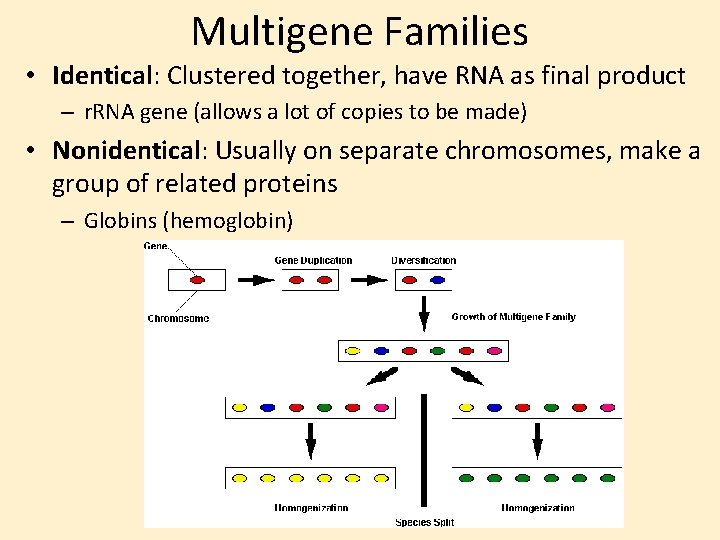
Multigene Families • Identical: Clustered together, have RNA as final product – r. RNA gene (allows a lot of copies to be made) • Nonidentical: Usually on separate chromosomes, make a group of related proteins – Globins (hemoglobin)
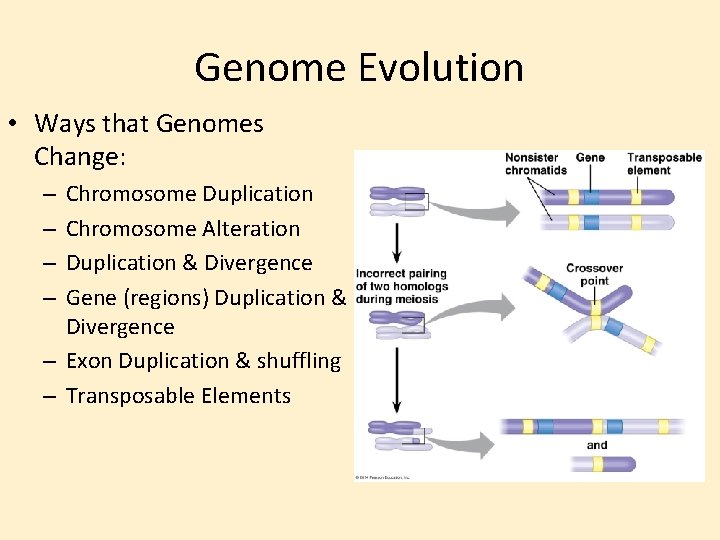
Genome Evolution • Ways that Genomes Change: Chromosome Duplication Chromosome Alteration Duplication & Divergence Gene (regions) Duplication & Divergence – Exon Duplication & shuffling – Transposable Elements – –
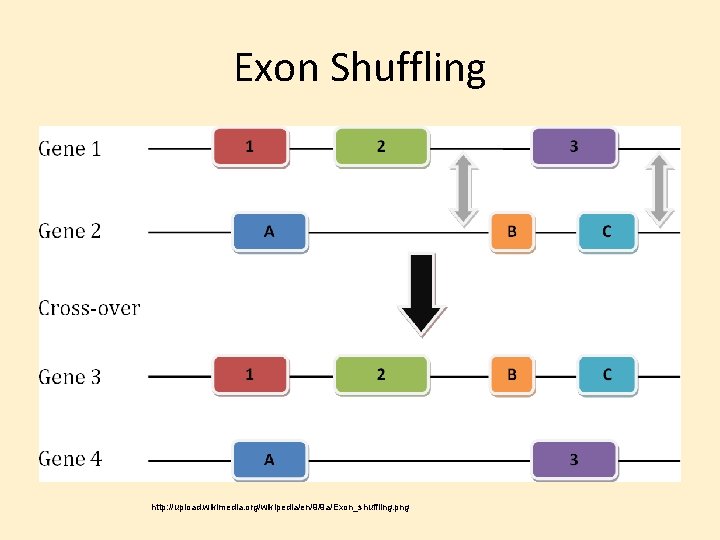
Exon Shuffling http: //upload. wikimedia. org/wikipedia/en/9/9 a/Exon_shuffling. png

Evolutionary Relationships and Genes • Closely related spp. – Look to sequences differences – Repeating sequences – Alu’s, more present in humans than chimps • Distantly related spp. – Look to highly conserved proteins and compare gene sequences • Evo-Devo, compare developmental processes of different organisms • Homeobox (Hox genes): Control anterior (front) & posterior (back) development in fruit flies & mice – same genetic sequences
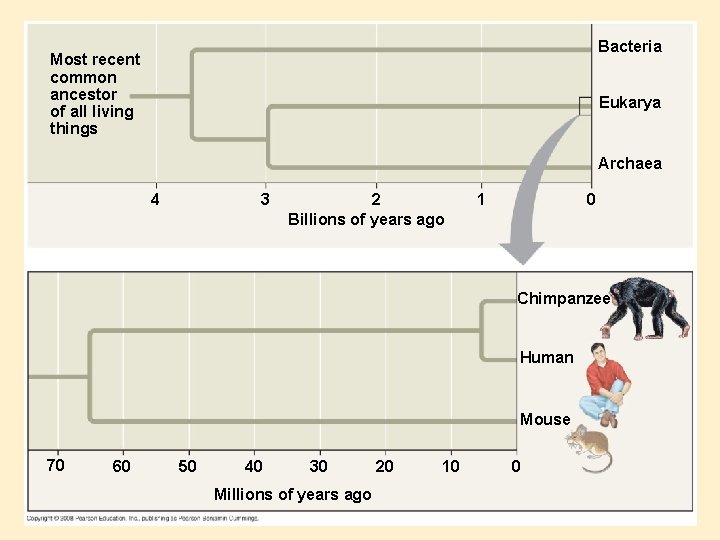
Bacteria Most recent common ancestor of all living things Eukarya Archaea 4 3 2 Billions of years ago 1 0 Chimpanzee Human Mouse 70 60 50 40 30 Millions of years ago 20 10 0
- Slides: 18