Genome Wide Searches for RNA Secondary Structure Motifs






- Slides: 22

Genome Wide Searches for RNA Secondary Structure Motifs Russell S. Hamilton Davis Lab Wellcome Trust Centre for Cell Biology Drosophila melanogaster

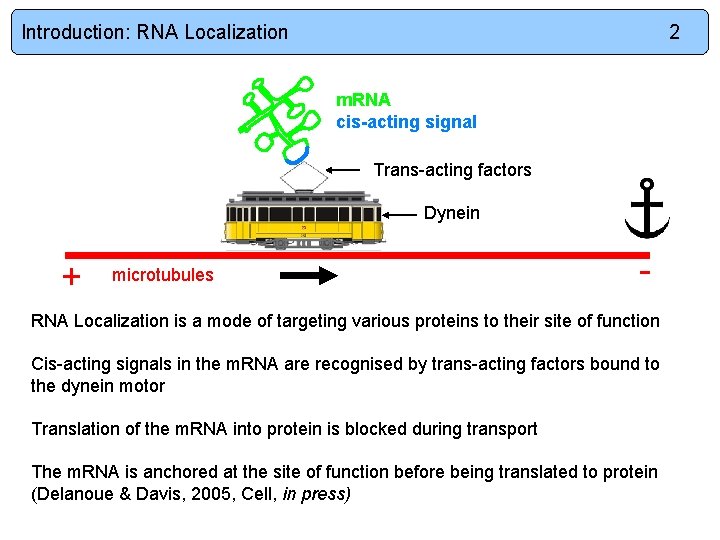
Introduction: RNA Localization 2 m. RNA cis-acting signal Trans-acting factors Dynein + microtubules - RNA Localization is a mode of targeting various proteins to their site of function Cis-acting signals in the m. RNA are recognised by trans-acting factors bound to the dynein motor Translation of the m. RNA into protein is blocked during transport The m. RNA is anchored at the site of function before being translated to protein (Delanoue & Davis, 2005, Cell, in press)

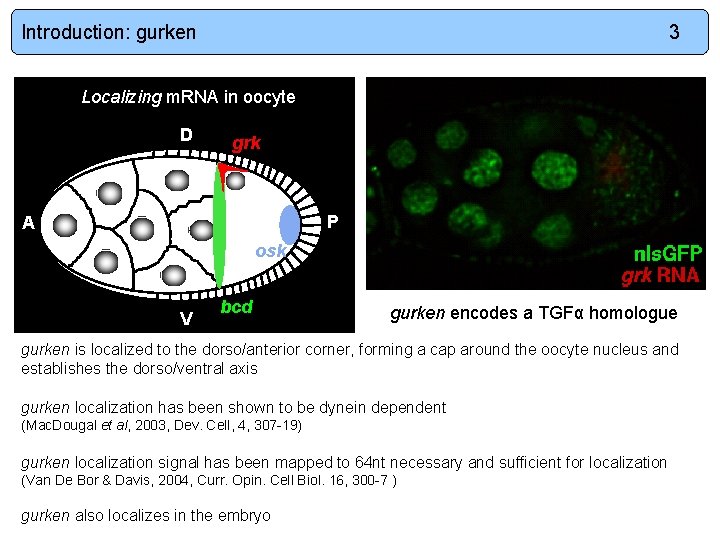
Introduction: gurken 3 Localizing m. RNA in oocyte D grk P A osk V bcd gurken encodes a TGFα homologue gurken is localized to the dorso/anterior corner, forming a cap around the oocyte nucleus and establishes the dorso/ventral axis gurken localization has been shown to be dynein dependent (Mac. Dougal et al, 2003, Dev. Cell, 4, 307 -19) gurken localization signal has been mapped to 64 nt necessary and sufficient for localization (Van De Bor & Davis, 2004, Curr. Opin. Cell Biol. 16, 300 -7 ) gurken also localizes in the embryo

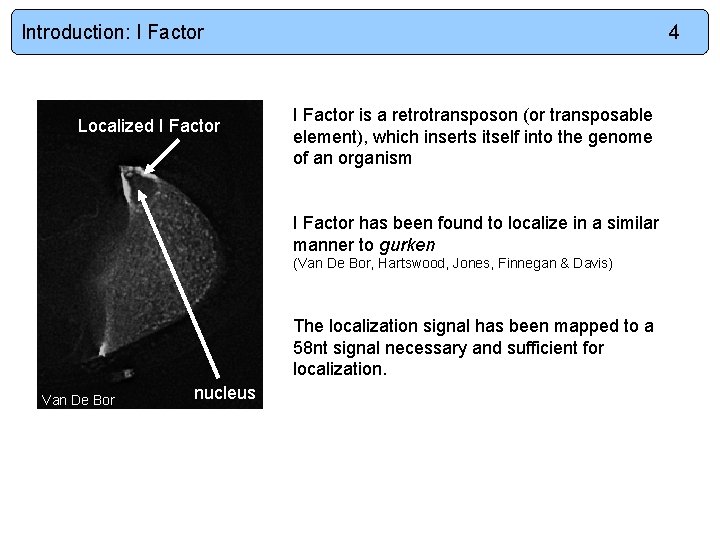
Introduction: I Factor Localized I Factor 4 I Factor is a retrotransposon (or transposable element), which inserts itself into the genome of an organism I Factor has been found to localize in a similar manner to gurken (Van De Bor, Hartswood, Jones, Finnegan & Davis) The localization signal has been mapped to a 58 nt signal necessary and sufficient for localization. Van De Bor nucleus

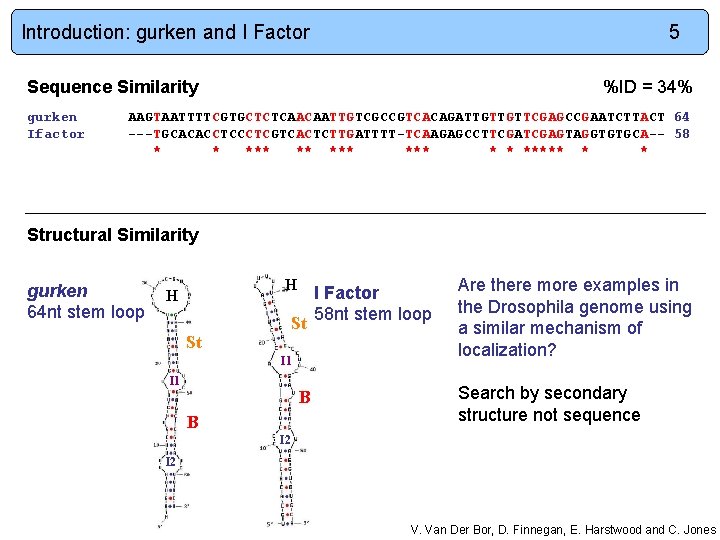
Introduction: gurken and I Factor 5 Sequence Similarity gurken Ifactor %ID = 34% AAGTAATTTTCGTGCTCTCAACAATTGTCGCCGTCACAGATTGTTGTTCGAGCCGAATCTTACT 64 ---TGCACACCTCGTCACTCTTGATTTT-TCAAGAGCCTTCGAGTAGGTGTGCA-- 58 * * *** *** * * ***** * * Structural Similarity gurken 64 nt stem loop H H I Factor St 58 nt stem loop St I 1 B B Are there more examples in the Drosophila genome using a similar mechanism of localization? Search by secondary structure not sequence I 2 V. Van Der Bor, D. Finnegan, E. Harstwood and C. Jones

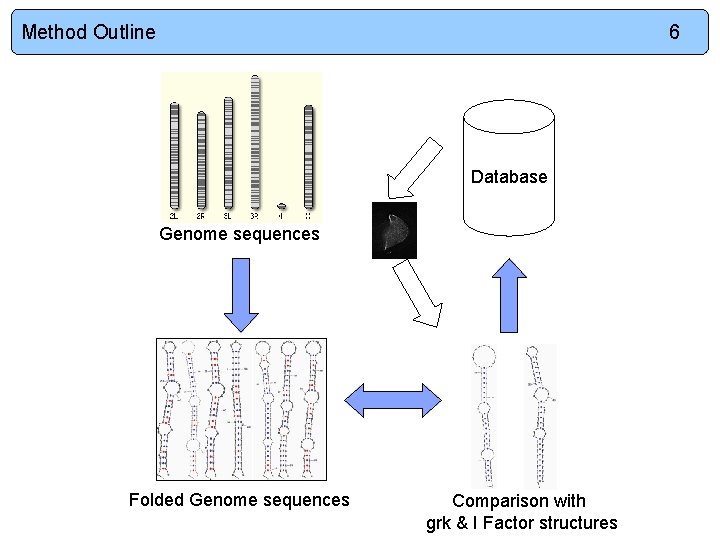
Method Outline 6 Database Genome sequences Folded Genome sequences Comparison with grk & I Factor structures

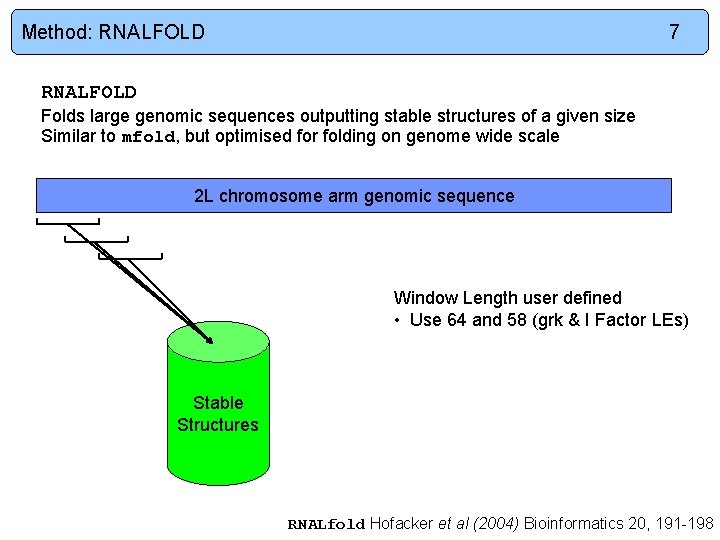
Method: RNALFOLD 7 RNALFOLD Folds large genomic sequences outputting stable structures of a given size Similar to mfold, but optimised for folding on genome wide scale 2 L chromosome arm genomic sequence Window Length user defined • Use 64 and 58 (grk & I Factor LEs) Stable Structures RNALfold Hofacker et al (2004) Bioinformatics 20, 191 -198

Method: RNAdistance & RNAforester 8 RNAdistance & RNAforester • Structures represented in bracket format Minimal representation maintaining all structural characteristics • Structures then aligned (not by sequence) with the query structure e. g. gurken LE • Scores can be weighted by sequence length and total number of base pairs . . (((((. . . ))))). . -(-(((-. . ))))-. RNAdistance Global Structure Comparison Hofacker (1994) Monatsh. Chem. 125, 167 -188 RNAforester Local Structure Comparison Hochsmann (2003) Proc. Comp. Sys. Bioinf. (CSB 2003) Matches = + score Mismatches = - score ( = base pair. = unpaired base - = gap

Method: RNAMotif 9 Flexible secondary structure definition and searching algorithm Two step process Step 1. Create a structure description Step 2. Use the description to find matching structures in a sequence database Uses Mfold (and pknots) for secondary structure predictions Output can be ranked by thermodynamic stability User Defined Scoring Based on if/then/else statements e. g. if loop has 6 -8 bases then score += 10 else score -= 10 Algorithm Summary Description converted to a tree structure Sequence being matched, has secondary structure converted to tree structure Then the matching can occur. Macke, T. J. et al (2001) Nucl. , Acids. , Res. , 29, 4724 -4735

Method: RNAMotif 10 Define base pairings allowed (in addition to Watson-Crick) Define stems, loops, and bulges • Including number of nucleotides • Setting a range 0 -N means it can either be present or not Can also put in sequence constraints Including tolerated mismatches Can search for pseudoknots, triplexes & quadruplexes Very flexible method of describing secondary structures

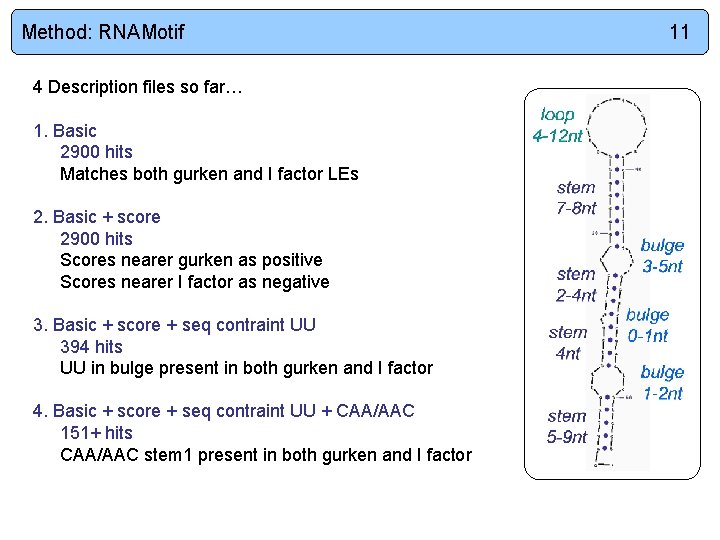
Method: RNAMotif 4 Description files so far… 1. Basic 2900 hits Matches both gurken and I factor LEs 2. Basic + score 2900 hits Scores nearer gurken as positive Scores nearer I factor as negative 3. Basic + score + seq contraint UU 394 hits UU in bulge present in both gurken and I factor 4. Basic + score + seq contraint UU + CAA/AAC 151+ hits CAA/AAC stem 1 present in both gurken and I factor 11

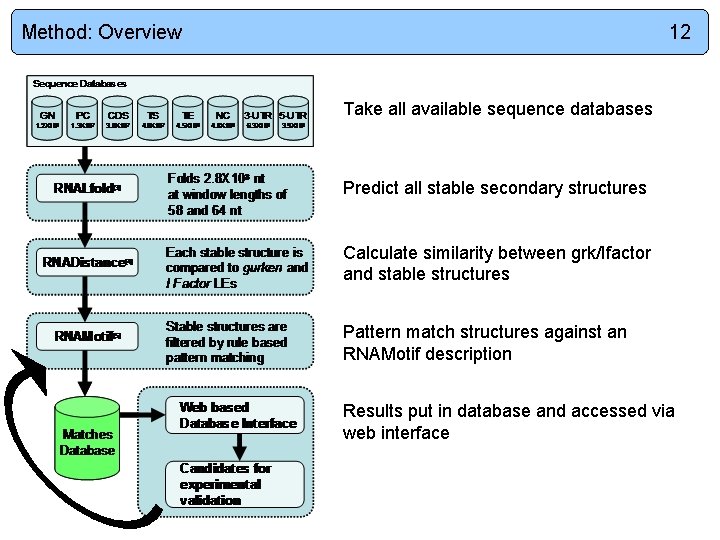
Method: Overview 12 Take all available sequence databases Predict all stable secondary structures Calculate similarity between grk/Ifactor and stable structures Pattern match structures against an RNAMotif description Results put in database and accessed via web interface

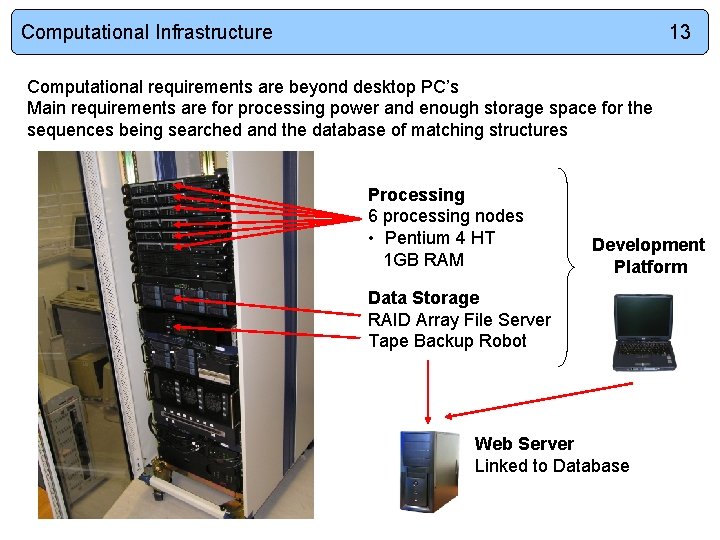
Computational Infrastructure 13 Computational requirements are beyond desktop PC’s Main requirements are for processing power and enough storage space for the sequences being searched and the database of matching structures Processing 6 processing nodes • Pentium 4 HT 1 GB RAM Development Platform Data Storage RAID Array File Server Tape Backup Robot Web Server Linked to Database

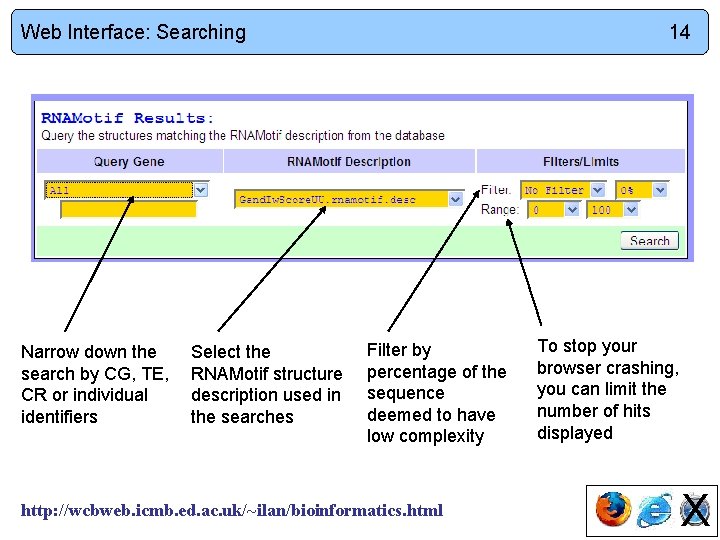
Web Interface: Searching Narrow down the search by CG, TE, CR or individual identifiers Select the RNAMotif structure description used in the searches 14 Filter by percentage of the sequence deemed to have low complexity http: //wcbweb. icmb. ed. ac. uk/~ilan/bioinformatics. html To stop your browser crashing, you can limit the number of hits displayed X

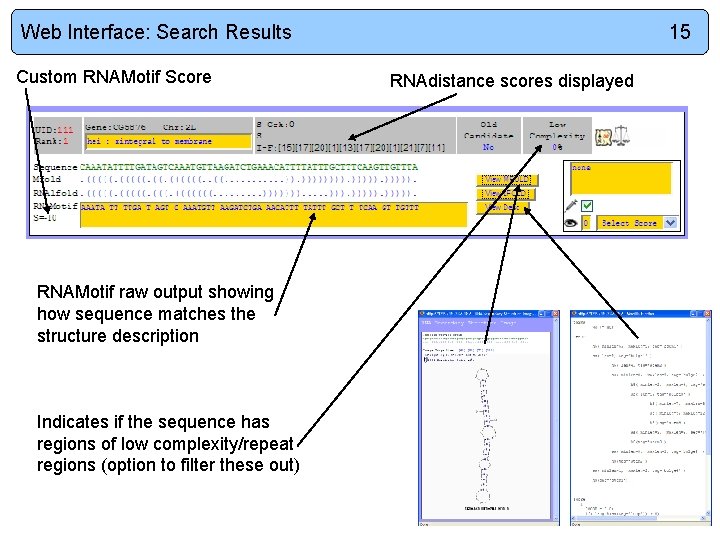
Web Interface: Search Results Custom RNAMotif Score RNAMotif raw output showing how sequence matches the structure description Indicates if the sequence has regions of low complexity/repeat regions (option to filter these out) 15 RNAdistance scores displayed

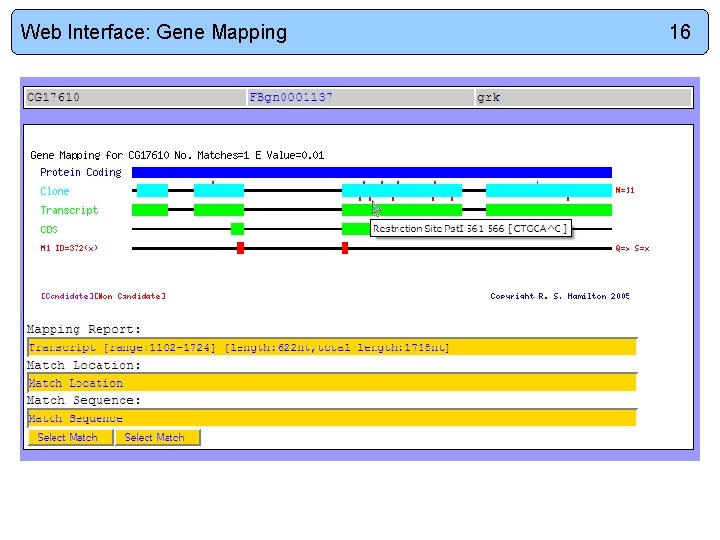
Web Interface: Gene Mapping 16

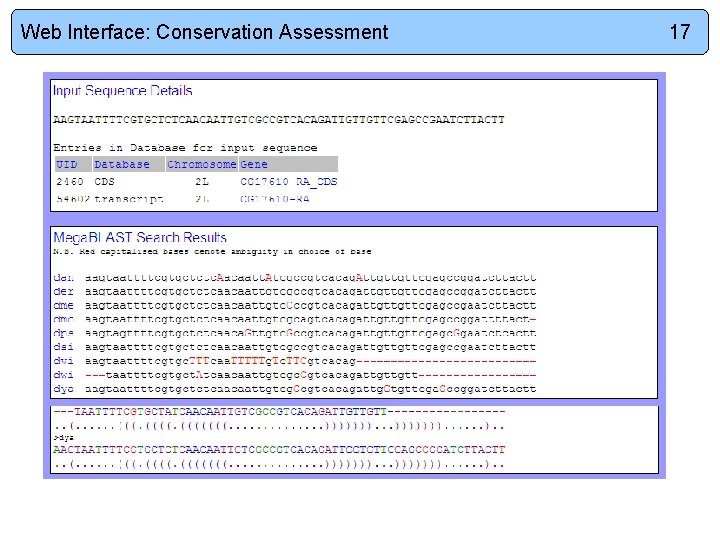
Web Interface: Conservation Assessment 17

Results: Candidate Injections 18 We are currently in the process of injecting candidates from the database into oocytes and embryos to determine if the RNA is localized. Results of candidate injections are stored in the database There have been suggestions that up to 20% of Drosophila genes may localize in the oocyte and/or embryo So we want to show that our method is able to enrich for localizing genes

Future Work: Expanding Searches Depending of the success of the experimental localization assays… Expand the searches to: • Other Drosophilid genomes 12 will be sequenced in the near future • Mammalian genomes (particularly human) Will require considerable computational power Search for LINE/SINE elements in human (transposon equivalents) Develop the web interface to enable real time searches to be performed on genes/genomes of interest • Requires massive computational power… 19

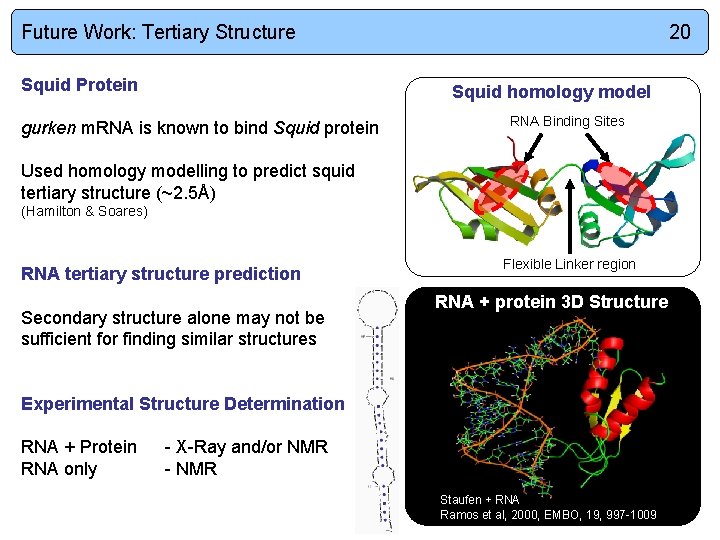
Future Work: Tertiary Structure Squid Protein 20 Squid homology model gurken m. RNA is known to bind Squid protein RNA Binding Sites Used homology modelling to predict squid tertiary structure (~2. 5Å) (Hamilton & Soares) RNA tertiary structure prediction Secondary structure alone may not be sufficient for finding similar structures Flexible Linker region RNA + protein 3 D Structure Experimental Structure Determination RNA + Protein RNA only - X-Ray and/or NMR - NMR Staufen + RNA Ramos et al, 2000, EMBO, 19, 997 -1009

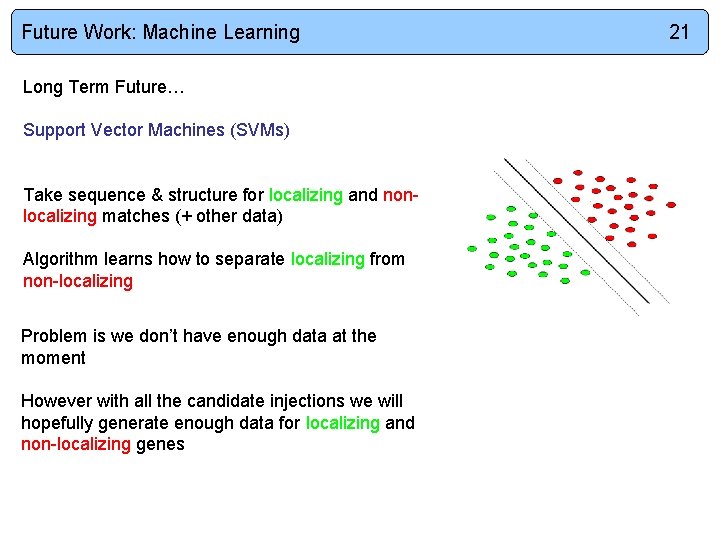
Future Work: Machine Learning Long Term Future… Support Vector Machines (SVMs) Take sequence & structure for localizing and nonlocalizing matches (+ other data) Algorithm learns how to separate localizing from non-localizing Problem is we don’t have enough data at the moment However with all the candidate injections we will hopefully generate enough data for localizing and non-localizing genes 21

Acknowledgements Davis Lab Ilan Davis Veronique Van De Bor Georgia Vendra Hille Tekotte Renald Delanue Carine Meignin Alejandra Clark Isabelle Kos Richard Parton 22 Finnegan Lab David Finnegan Eve Hartswood Cheryl Jones Bioinformatics Discussions Alastair Kerr Systems Administration Paul Taylor Homology Modelling Dinesh Soares Funding Software