Genome map Supplementary Figure 1 Epsilon 15 genome

- Slides: 8

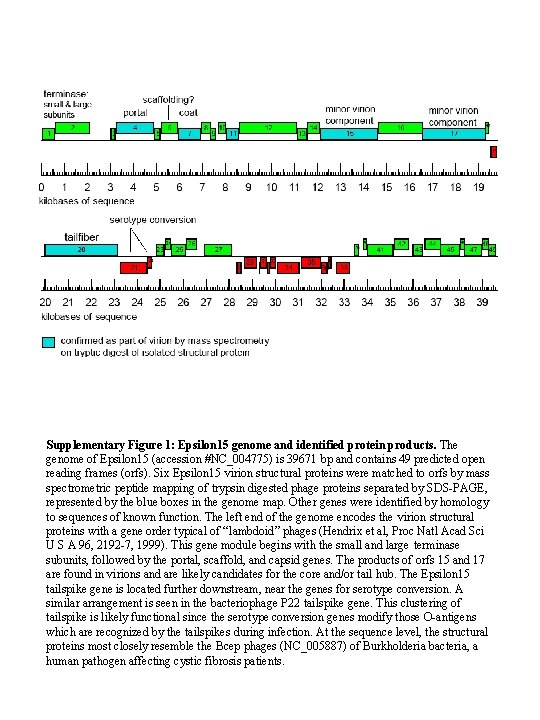
Genome map Supplementary Figure 1: Epsilon 15 genome and identified protein products. The genome of Epsilon 15 (accession #NC_004775) is 39671 bp and contains 49 predicted open reading frames (orfs). Six Epsilon 15 virion structural proteins were matched to orfs by mass spectrometric peptide mapping of trypsin digested phage proteins separated by SDS-PAGE, represented by the blue boxes in the genome map. Other genes were identified by homology to sequences of known function. The left end of the genome encodes the virion structural proteins with a gene order typical of “lambdoid” phages (Hendrix et al, Proc Natl Acad Sci U S A 96, 2192 -7, 1999). This gene module begins with the small and large terminase subunits, followed by the portal, scaffold, and capsid genes. The products of orfs 15 and 17 are found in virions and are likely candidates for the core and/or tail hub. The Epsilon 15 tailspike gene is located further downstream, near the genes for serotype conversion. A similar arrangement is seen in the bacteriophage P 22 tailspike gene. This clustering of tailspike is likely functional since the serotype conversion genes modify those O-antigens which are recognized by the tailspikes during infection. At the sequence level, the structural proteins most closely resemble the Bcep phages (NC_005887) of Burkholderia bacteria, a human pathogen affecting cystic fibrosis patients.

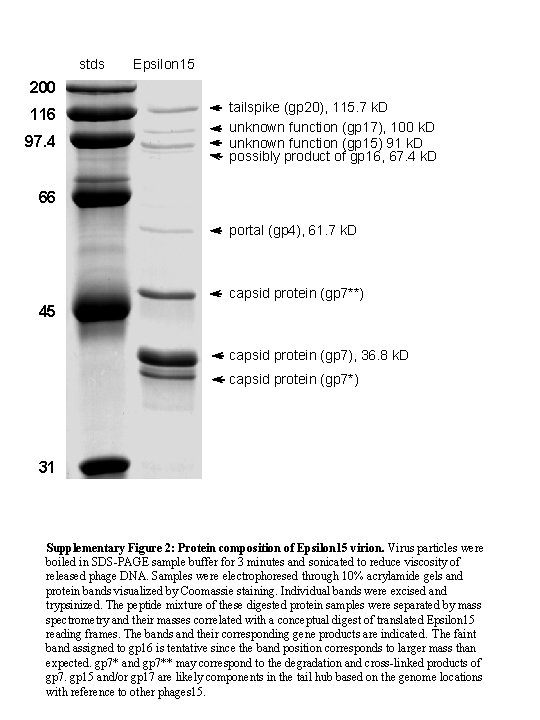
stds Epsilon 15 200 116 97. 4 tailspike (gp 20), 115. 7 k. D unknown function (gp 17), 100 k. D unknown function (gp 15) 91 k. D possibly product of gp 16, 67. 4 k. D 66 portal (gp 4), 61. 7 k. D capsid protein (gp 7**) 45 capsid protein (gp 7), 36. 8 k. D capsid protein (gp 7*) 31 Supplementary Figure 2: Protein composition of Epsilon 15 virion. Virus particles were boiled in SDS-PAGE sample buffer for 3 minutes and sonicated to reduce viscosity of released phage DNA. Samples were electrophoresed through 10% acrylamide gels and protein bands visualized by Coomassie staining. Individual bands were excised and trypsinized. The peptide mixture of these digested protein samples were separated by mass spectrometry and their masses correlated with a conceptual digest of translated Epsilon 15 reading frames. The bands and their corresponding gene products are indicated. The faint band assigned to gp 16 is tentative since the band position corresponds to larger mass than expected. gp 7* and gp 7** may correspond to the degradation and cross-linked products of gp 7. gp 15 and/or gp 17 are likely components in the tail hub based on the genome locations with reference to other phages 15.

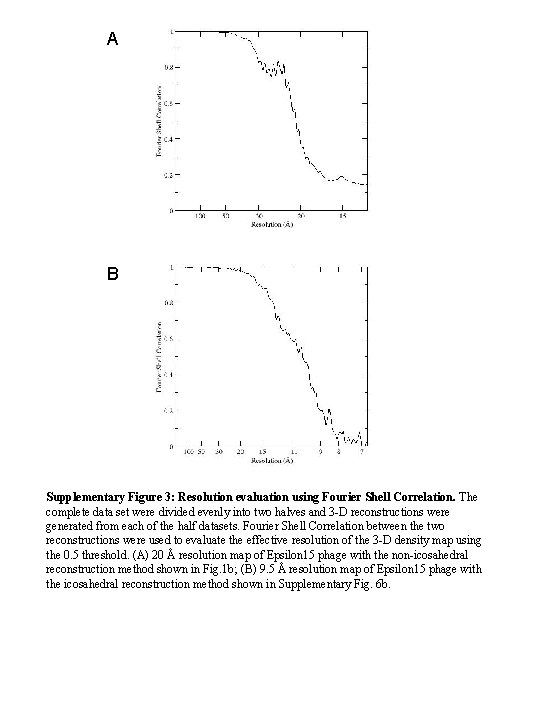
A B Supplementary Figure 3: Resolution evaluation using Fourier Shell Correlation. The complete data set were divided evenly into two halves and 3 -D reconstructions were generated from each of the half datasets. Fourier Shell Correlation between the two reconstructions were used to evaluate the effective resolution of the 3 -D density map using the 0. 5 threshold. (A) 20 Å resolution map of Epsilon 15 phage with the non-icosahedral reconstruction method shown in Fig. 1 b; (B) 9. 5 Å resolution map of Epsilon 15 phage with the icosahedral reconstruction method shown in Supplementary Fig. 6 b.

1 2 3 4 5 6 Side Top * Supplementary Figure 4: Conformation variations of the tailspikes. Shown are the side and top views of individual tailspikes. Top view of the tailspikes exhibits 3 -fold symmetry. Structures of most tailspikes except tailspike 3 (the central knob and one of the petals close to the label *) are very similar.

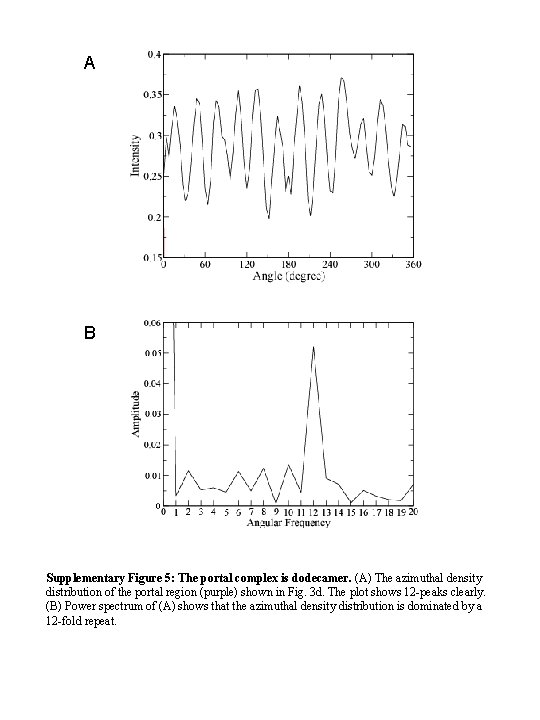
A B Supplementary Figure 5: The portal complex is dodecamer. (A) The azimuthal density distribution of the portal region (purple) shown in Fig. 3 d. The plot shows 12 -peaks clearly. (B) Power spectrum of (A) shows that the azimuthal density distribution is dominated by a 12 -fold repeat.

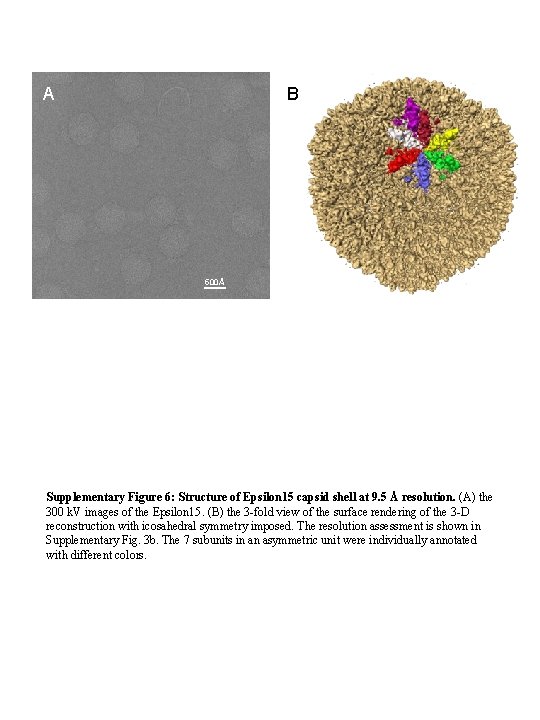
A B 500Å Supplementary Figure 6: Structure of Epsilon 15 capsid shell at 9. 5 Å resolution. (A) the 300 k. V images of the Epsilon 15. (B) the 3 -fold view of the surface rendering of the 3 -D reconstruction with icosahedral symmetry imposed. The resolution assessment is shown in Supplementary Fig. 3 b. The 7 subunits in an asymmetric unit were individually annotated with different colors.

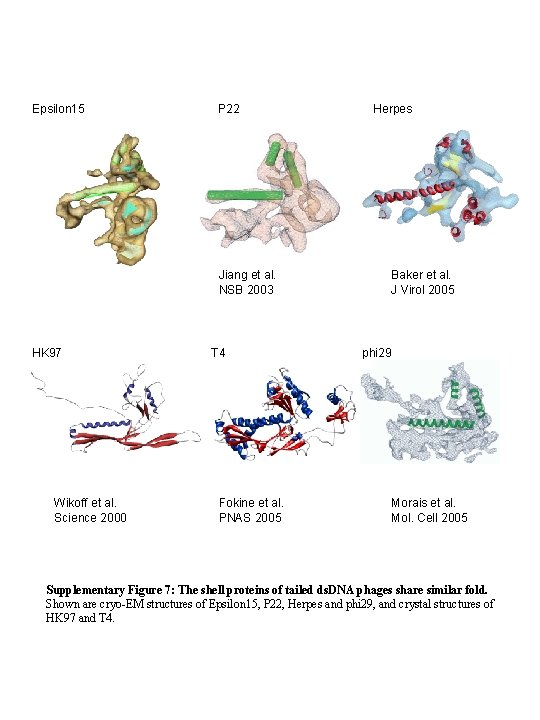
Epsilon 15 P 22 Jiang et al. NSB 2003 HK 97 Wikoff et al. Science 2000 T 4 Fokine et al. PNAS 2005 Herpes Baker et al. J Virol 2005 phi 29 Morais et al. Mol. Cell 2005 Supplementary Figure 7: The shell proteins of tailed ds. DNA phages share similar fold. Shown are cryo-EM structures of Epsilon 15, P 22, Herpes and phi 29, and crystal structures of HK 97 and T 4.

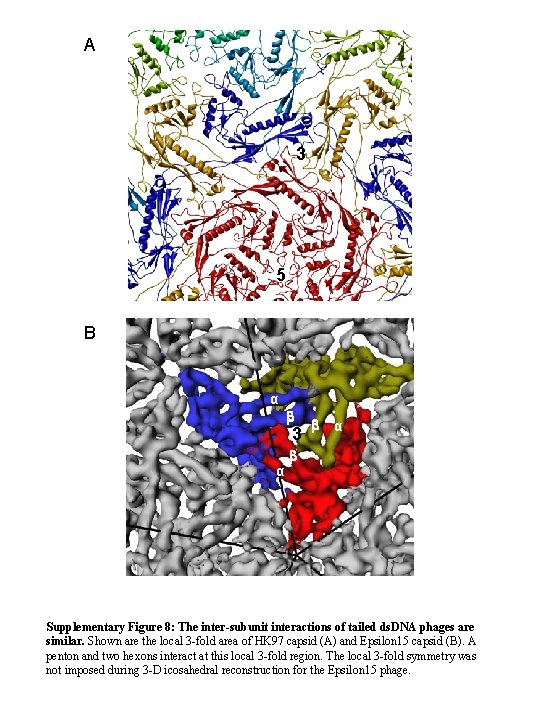
A 3 5 B α b 3 α b Supplementary Figure 8: The inter-subunit interactions of tailed ds. DNA phages are similar. Shown are the local 3 -fold area of HK 97 capsid (A) and Epsilon 15 capsid (B). A penton and two hexons interact at this local 3 -fold region. The local 3 -fold symmetry was not imposed during 3 -D icosahedral reconstruction for the Epsilon 15 phage.