Genome Characterization Assemblysequencing Assigned reading Ch 9 BIO

Genome Characterization Assembly/sequencing Assigned reading: Ch 9 BIO 520 Bioinformatics Jim Lund
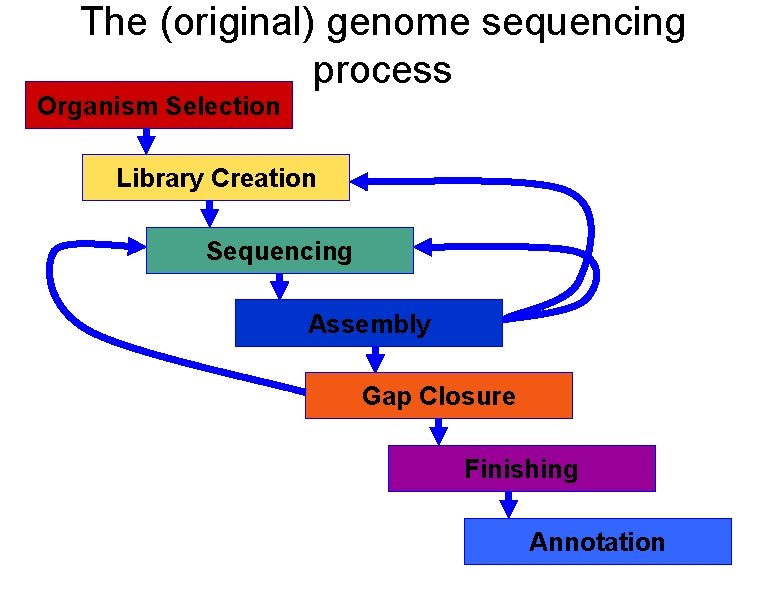
The (original) genome sequencing process Organism Selection Library Creation Sequencing Assembly Gap Closure Finishing Annotation
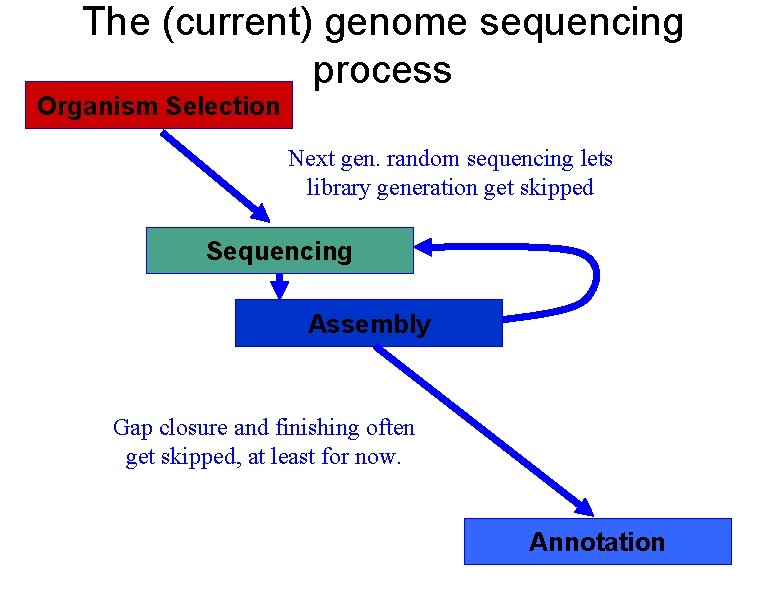
The (current) genome sequencing process Organism Selection Next gen. random sequencing lets library generation get skipped Sequencing Assembly Gap closure and finishing often get skipped, at least for now. Annotation
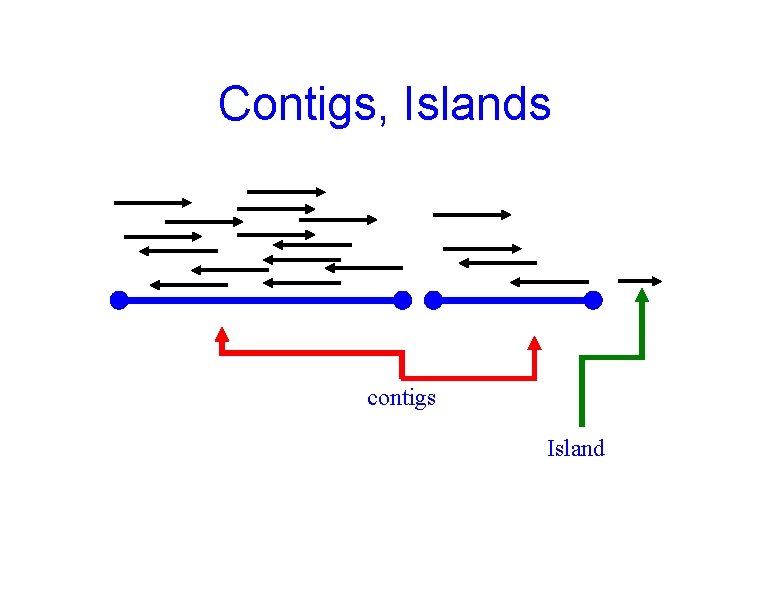
Contigs, Islands contigs Island

Assembly pipeline 1. Sequence reads. 2. Phred: base calling. 3. crossmatch: screen out vector, E. coli sequence. 4. Phrap: assemble contigs. 5. Consed: view assembly, correct problems. 6. Finishing.

Assembly Methods • • Strip out vector (or contaminant) Mask known repeats Trim off unreliable data Find Matches (n seq x n seq comparisons) – how long (what ktuple [10 common]) – how perfect (reliability index) – where to look? (ends only vs entire)

Assembly Programs • PHRAP FAMILY – phred/phrap/consed/cross_match – Developed by Phil Green, U of Wash. • Other assemblers – phrap, kangaroo, phrapo, – CAP, TIGRAssembler, . . . http: //www. phrap. org/

Assembly • Phred -reads DNA sequencing trace files, calls bases, and assigns a quality value to each called base. – The quality value is a log-transformed error probability, specifically: Q = -10 log 10( Pe ) – Q = quality value, Pe = error probability. – Q= 20 -> 1% chance of miscall, Q= 30 -> 0. 1% chance of miscall. • Phrap -assembles shotgun DNA sequence data. • Consed/Autofinish -view, edit, and finish sequence assemblies created with phrap. – Allows the user to pick primers and templates – Suggests additional sequencing reactions – Suggest digests and forward/reverse pair information to check accuracy of assembly.

Poisson statistics for sequencing completion L=read length N=#reads G=genome size =e-L(N)/G P 0 Coverage 1 = 1 -fold = 1 X 1 % not sequenced 37 3 5 8 0. 03 10 0. 005 50 < 1 e 20 E. coli 15 kb H. sapiens 900 kb
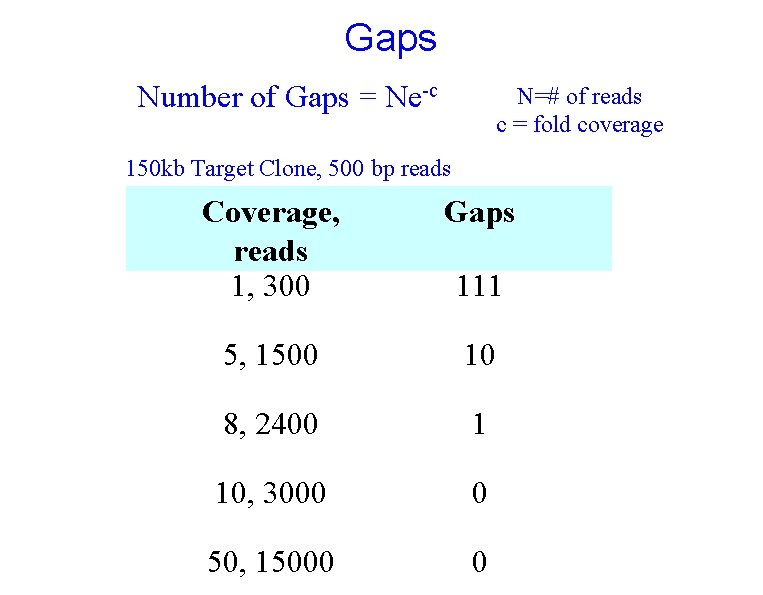
Gaps Number of Gaps = Ne-c N=# of reads c = fold coverage 150 kb Target Clone, 500 bp reads Coverage, reads 1, 300 Gaps 5, 1500 10 8, 2400 1 10, 3000 0 50, 15000 0 111
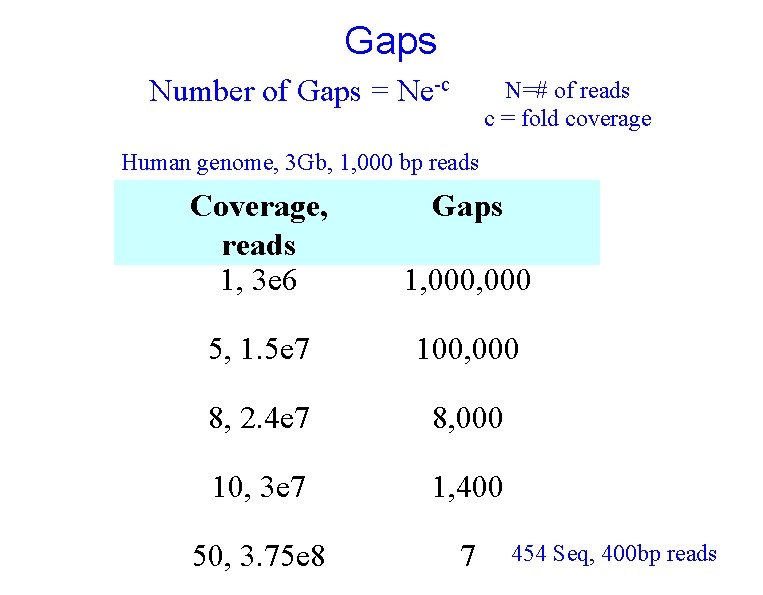
Gaps Number of Gaps = Ne-c N=# of reads c = fold coverage Human genome, 3 Gb, 1, 000 bp reads Coverage, reads 1, 3 e 6 Gaps 1, 000 5, 1. 5 e 7 100, 000 8, 2. 4 e 7 8, 000 10, 3 e 7 1, 400 50, 3. 75 e 8 7 454 Seq, 400 bp reads
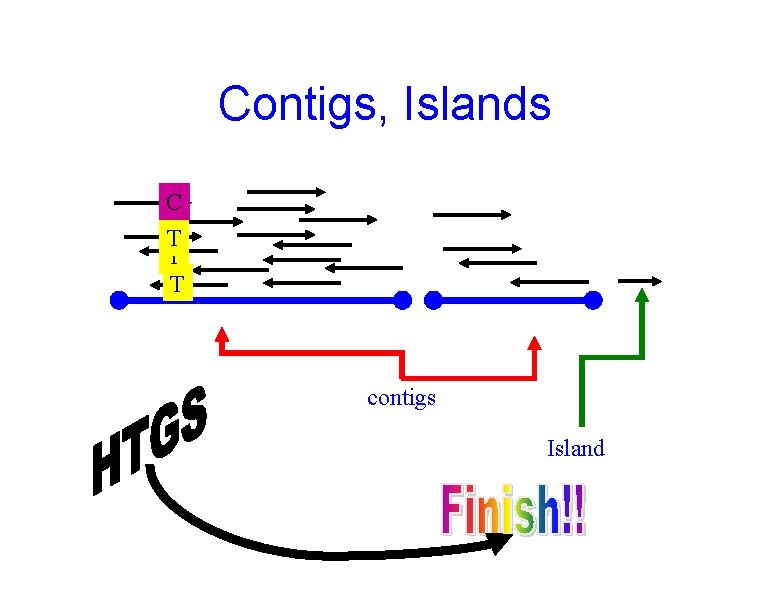
Contigs, Islands C T T T contigs Island

Finishing • GOALS – >95% coverage on BOTH strands – every base covered 3 X – resolve ambiguities • Finish when random no longer productive (~8 X range)

Sequence finishing. How? • Identify gaps, ambiguities – Captured gaps: gaps is contained in a clone • Extend from end of contigs – Resequencing, new chemistry. – Specific primers – Subcloning and sequencing. • Uncaptured gaps. – New specific primers – PCR across gap, sequence PCR product. • Resolve ambiguities – Consensus or resequence • Specific primers, different chemistry

Large clone sequencing process • Phase 1: Unfinished, may be unordered/unoriented contigs, with gaps. • Phase 2: Unfinished, fully oriented and ordered sequence, may contain gaps and low quality sequences • Phase 3: Finished, no gaps.

Genome assembly after initial contigs are made • Order clones/contig sequences: – Sequence overlaps. • Clone/contig end sequences. – Clone fingerprints. – Anchor using other maps • Sequence based markers on genetic or physical maps. • Conserved synteny to other genomes. • Easiest when re-sequencing, e. g, another human genome!

Process Control • LIMS – Laboratory information management system • AIMS – Analysis information management system
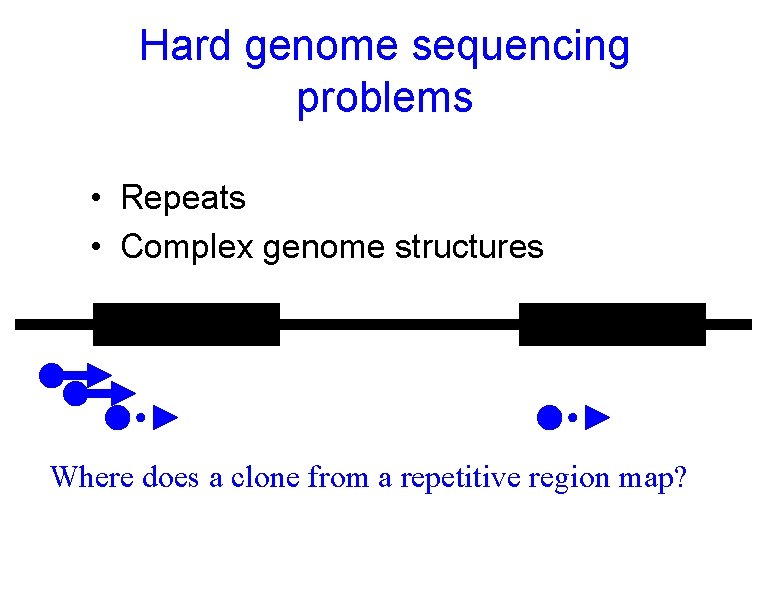
Hard genome sequencing problems • Repeats • Complex genome structures Where does a clone from a repetitive region map?
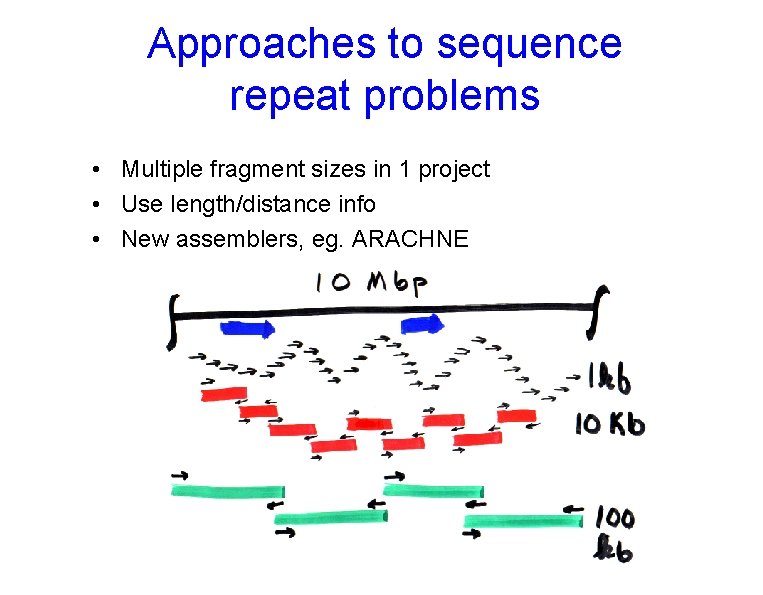
Approaches to sequence repeat problems • Multiple fragment sizes in 1 project • Use length/distance info • New assemblers, eg. ARACHNE
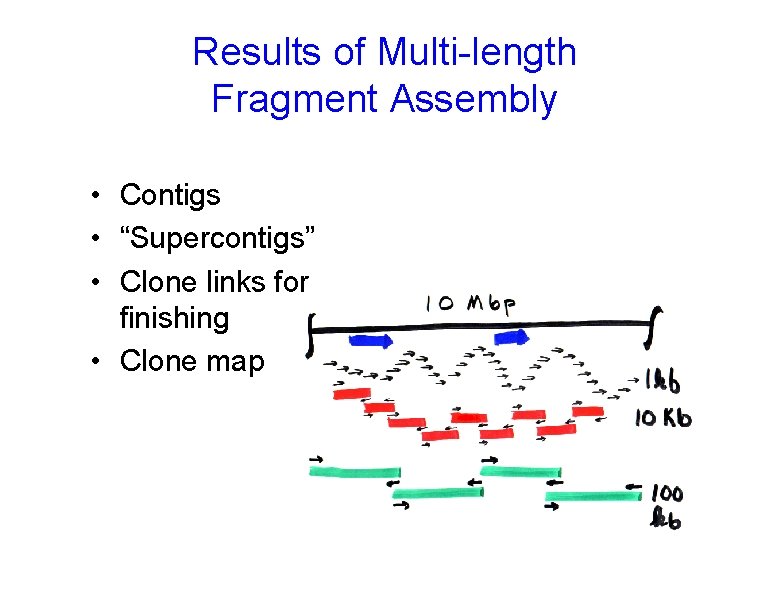
Results of Multi-length Fragment Assembly • Contigs • “Supercontigs” • Clone links for finishing • Clone map

DOE Joint Genome Institute (JGI) Prokaryote Finishing Standards • All low-quality areas (<Q 30) are reviewed and resequenced. • The final error rate must be less than 0. 2 per 10 Kb. • No single-clone coverage is permitted (minimum of 2 x depth everywhere). • Single-stranded regions are manually inspected and quantified. • All positions where an aligned high-quality read (>Q 29) disagrees with the consensus base are checked. • All strings of xxxx are resolved in the final sequence. • All repeats are verified. • The ends of final contigs (chromosomes, plasmids) are checked • The final assembly is given a manual QC check.

Completed genomes 23 complete, 329 in assembly, in progress 389 Arabidopsis thaliana Caenorhabditis elegans Candida glabrata Cryptococcus neoformans Cyanidioschyzon merolae Debaryomyces hansenii Drosophila melanogaster Encephalitozoon cuniculi Entamoeba histolytica Plants Animal s Eremothecium gossypii Homo sapiens Kluyveromyces lactis Leishmzania major Friedlin Mus musculus Oryza sativa Saccharomyces cerevisiae Schizosaccharomyces pombe Trypanosoma cruzi Yarrowia lipolytica Protists Fungi http: //www. ncbi. nlm. nih. gov/genomes/leuks. cgi

Genomes Complete • Eukaryotes--23 complete, 329 in assembly, in progress 389 – – – Human, mouse, rat, zebrafish, Homo sapiens neanderthalensis Drosophila, Anopheles, Caenorhabditis Arabadopsis, oat, corn, barley, rice, tomato Saccharomyces, Schizosaccharomyces, Magnaportha, Cryptococcus, Candida… – Encephalitozoon cuniculi, Guillardia theta – Toxoplasma, Plasmodium – And many more…

Eubacteria and Archaea genomes • 608 Bacteria and 48 Archaea completed • Comprehensive Microbial Resource – http: //pathema. tigr. org/tigrscripts/CMR/Cmr. Home. Page. cgi • Joint Genome Institute – http: //www. jgi. doe. gov/genome-projects/ – 2065 genome projects underway or completed! • NCBI Genomes

Genome Centers • Joint Genome Institute (DOE) • Whitehead Institute (MIT) • TIGR • Washington University (St. Louis) • Celera • Sanger Institute (the other UK) • RIKEN (Japan) • Beijing Genomics Institute (China) • Max Planck (Germany) …

Where do you find Genomic data? • NCBI – Entrez (by clone, by Refseq) – Genome (view and search map) • Genome center sites • Organism genome project sites • Annotations projects – UCSC Genome Browser, – Ensembl Genome Browser
Arabidopsis http: //mips. helmholtz-muenchen. de/plant/athal/index. jsp

C. elegans (nematode) http: //wormbase. org
- Slides: 28