Geno Mesh A genomewide Me SHbased literature mining

Geno. Mesh: A genome-wide Me. SH-based literature mining system predicts implicit gene-to-gene relationships & networks Yongqun “Oliver” He Unit for Laboratory Animal Medicine Department of Microbiology and Immunology Center for Computational Medicine and Bioinformatics Comprehensive Cancer Center University of Michigan Medical School Ann Arbor, MI 48109

Outline 1. Background 2. Development & evaluation of Geno. Mesh algorithm 3. Geno. Mesh web system and features 4. Usages of Geno. Mesh Reference: Zuoshuang Xiang, Tingting Qin, Zhaohui S. Qin, and Yongqun He. A genome-wide Me. SH-based literature mining system predicts implicit gene-to-gene relationships and networks. BMC Systems Biology. 2013, 7(Suppl 3): S 9. (In. Co. B 2013)
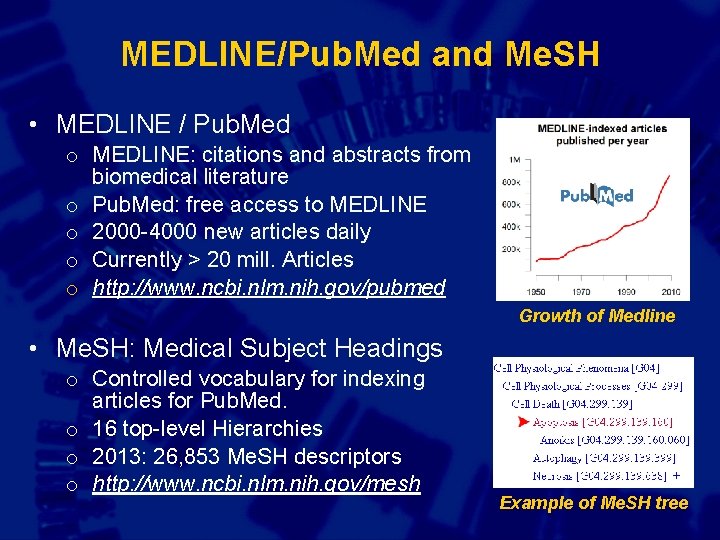
MEDLINE/Pub. Med and Me. SH • MEDLINE / Pub. Med o MEDLINE: citations and abstracts from biomedical literature o Pub. Med: free access to MEDLINE o 2000 -4000 new articles daily o Currently > 20 mill. Articles o http: //www. ncbi. nlm. nih. gov/pubmed Growth of Medline • Me. SH: Medical Subject Headings o Controlled vocabulary for indexing articles for Pub. Med. o 16 top-level Hierarchies o 2013: 26, 853 Me. SH descriptors o http: //www. ncbi. nlm. nih. gov/mesh Example of Me. SH tree

Gene-gene Interaction Literature Mining • Two general strategies: • Gene co-occurrence o o Two genes are related if in the same article Particularly in titles, abstracts, or sentences Example program: Pub. Gene Limitation: unable to predict unknown relations • Infer gene relatedness based on common linkage to keywords (e. g. GO, Me. SH) o o Advantage: predict new gene-gene interactions. Example programs: ARROWSMITH, Me. SHmap No genome-wide Me. SH-based approach reported before Different methods, often not optimized • Geno. Mesh: genome-wide Me. SH-based o E. coli (well studied) and Brucella (less studied)
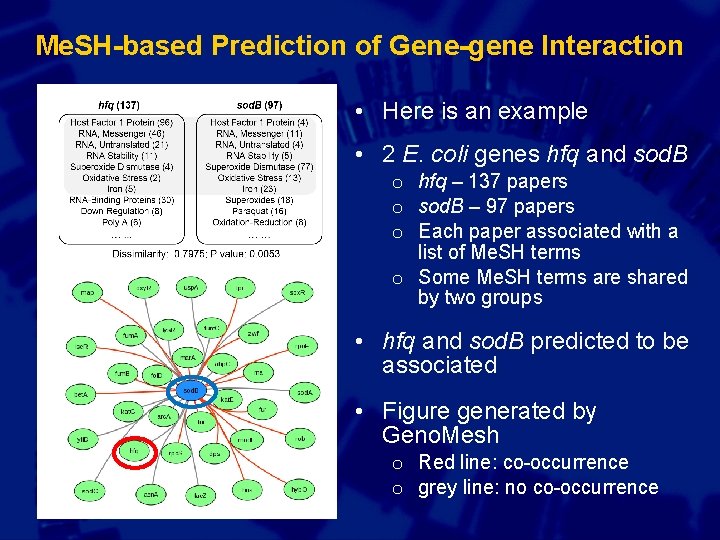
Me. SH-based Prediction of Gene-gene Interaction • Here is an example • 2 E. coli genes hfq and sod. B o hfq – 137 papers o sod. B – 97 papers o Each paper associated with a list of Me. SH terms o Some Me. SH terms are shared by two groups • hfq and sod. B predicted to be associated • Figure generated by Geno. Mesh o Red line: co-occurrence o grey line: no co-occurrence

Geno. Mesh algorithm Pipeline E. coli, Brucella Preprocessing • Me. SH term weighted o each term is highly or rarely used • TF-IDF: term frequency-inverse document frequency Gene-article matrix Gene-Me. SH matrix Gene-gene dissimilarity matrix Clustering, network • Six scores tested to measure gene dissimilarity o Cosine coefficient o Euclidean distance o ….
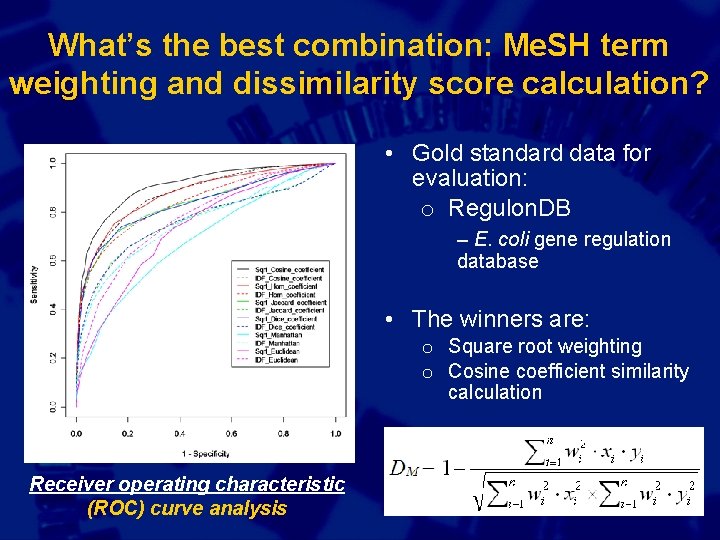
What’s the best combination: Me. SH term weighting and dissimilarity score calculation? • Gold standard data for evaluation: o Regulon. DB – E. coli gene regulation database • The winners are: o Square root weighting o Cosine coefficient similarity calculation Receiver operating characteristic (ROC) curve analysis
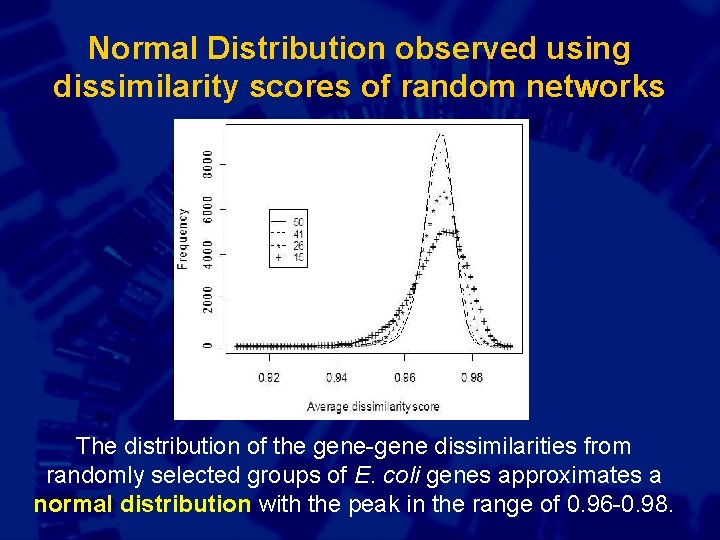
Normal Distribution observed using dissimilarity scores of random networks The distribution of the gene-gene dissimilarities from randomly selected groups of E. coli genes approximates a normal distribution with the peak in the range of 0. 96 -0. 98.

Geno. Mesh able to predict implicit gene-gene interactions Top E. coli 10 gene pairs predicated using literature data before 2004 and verified by literature data afterwards All proven valid
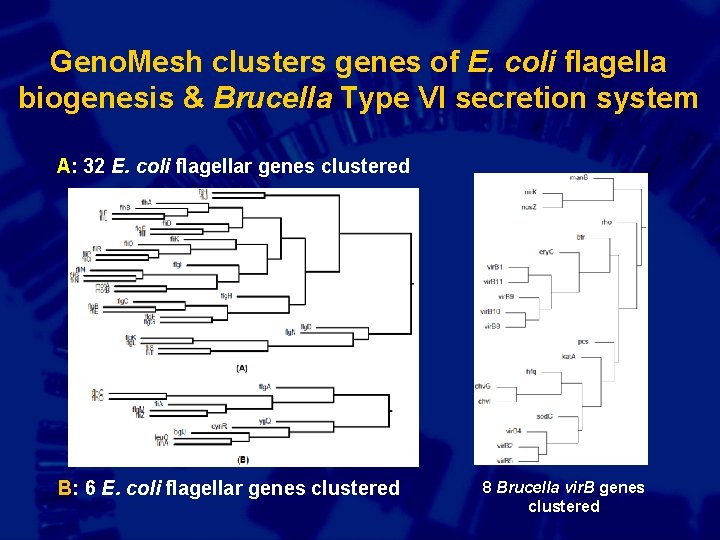
Geno. Mesh clusters genes of E. coli flagella biogenesis & Brucella Type VI secretion system A: 32 E. coli flagellar genes clustered B: 6 E. coli flagellar genes clustered 8 Brucella vir. B genes clustered

Geno. Mesh analysis of 31 E. coli pathways containing at least 10 genes from Eco. Cyc All have significant Z-value and p-value So Geno. Mesh can be used to study gene interaction networks
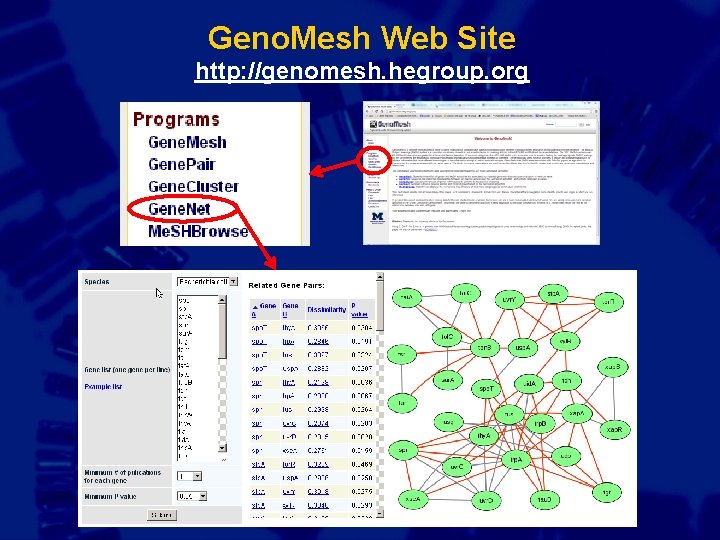
Geno. Mesh Web Site http: //genomesh. hegroup. org

Analysis of the term “Neutrophil Activation” from the Geno. Mesh Me. SHBrowse website This is a Geno. Mesh Me. SHBrowse example

Geno. Mesh predicts new Brucella gene-gene interactions by comparing with homologous E. coli results Homologous E. coli and Brucella genes & associated genes

Summary • The Geno. Mesh genome-wide Me. SH-based literature mining algorithm and web system is generated and evaluated • Geno. Mesh: o predicts implicit gene-gene interactions o clusters genes based on associations o generates gene interaction networks Discussion • More pathogens will be included • Also applicable for human and other eukaryotes • Host-pathogen interactions

Acknowledgements He Lab at the University of Michigan (UM) Ann Arbor, MI, USA • Zuoshuang Xiang • Tingting Qin Emory School of Medicine, Atlanta, GA, USA Zhaohui Steve Qin Funding: • NIH-NIAID Grant 1 R 01 AI 081062 • A pilot grant at the UM Center for Computational Medicine and Biology (CCMB)
- Slides: 16