Genetics of Diabetes Jan Dorman Ph D University

Genetics of Diabetes Jan Dorman, Ph. D University of Pittsburgh School of Nursing jsd@pitt. edu
Type 1 Diabetes (T 1 D)

Type 1 Diabetes n Caused by the destruction of the pancreatic beta cells – Insulin is no longer produced – Leads to hyperglycemia, ketoacidosis and potentially death if not treated with insulin n Treatment goals for T 1 D – Maintaining near normal levels of blood glucose – Avoidance of long-term complications

Type 1 Diabetes n 2 nd most common chronic childhood disease n Peak age at onset is around puberty – But T 1 D can occur at any age n Incidence is increasing worldwide by ~3% per year – Related to increase in T 2 D?
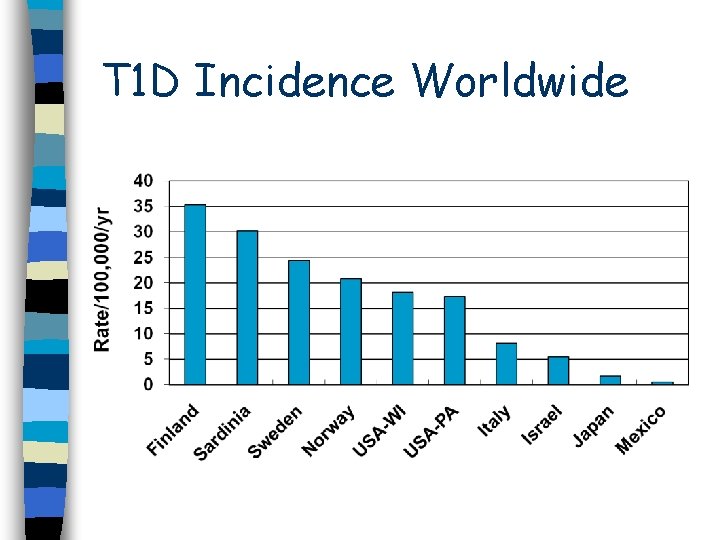
T 1 D Incidence Worldwide

Importance of Environmental Risk Factors in T 1 D n Seasonality at diagnosis n Migrants assume risk of host country n Risk factors from case-control studies – – – Infant/childhood diet Viruses – exposures as early as in utero Hormones Stress Improved hygiene Vitamin D

Importance of Genetic Risk Factors in T 1 D n Concordance in identical twins greater in MZ versus DZ twins n 15 -fold increased risk for 1 st degree relatives – Risk is ~6% through age 30 years – Risk increases in presence of susceptibility genes
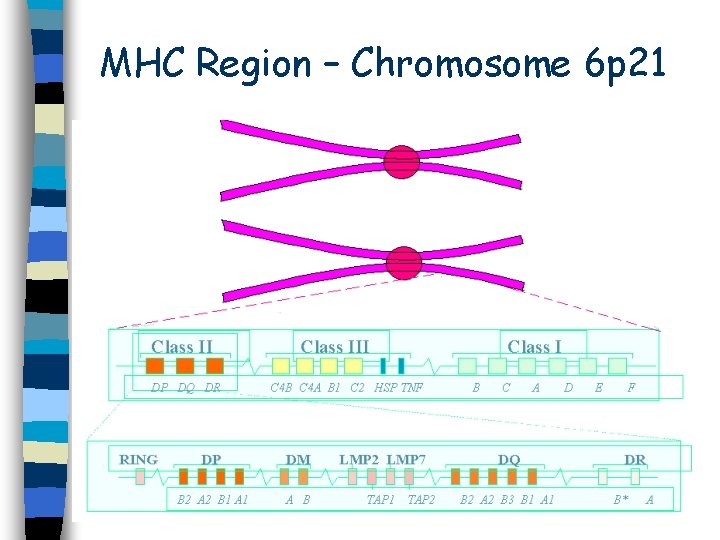
MHC Region – Chromosome 6 p 21

Predisposition to T 1 D is Better Determined by Haplotypes n DRB 1 -DQB 1 haplotypes more accurately determine T 1 D risk n Testing for both genes is more expensive – Most screening is based only on DQA 1 -DQB 1 n High risk T 1 D haplotypes – DQA 1*0501 -DQB 1*0201 – DQA 1*0301 -DQB 1*0302
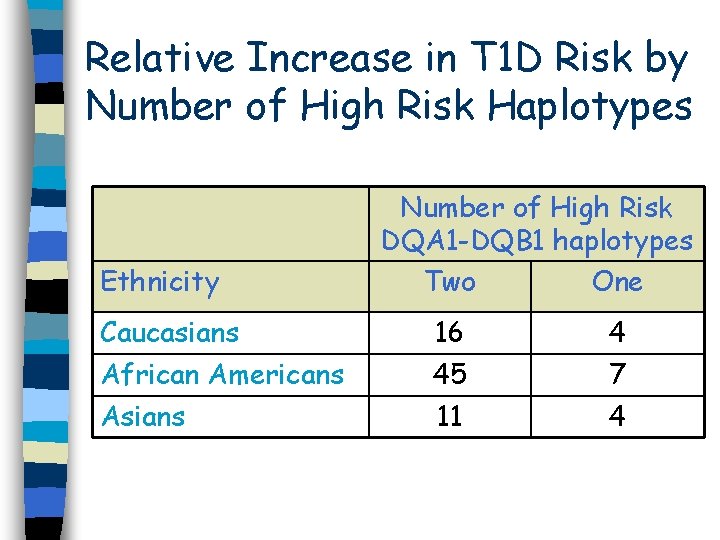
Relative Increase in T 1 D Risk by Number of High Risk Haplotypes Ethnicity Caucasians African Americans Asians Number of High Risk DQA 1 -DQB 1 haplotypes Two One 16 45 11 4 7 4
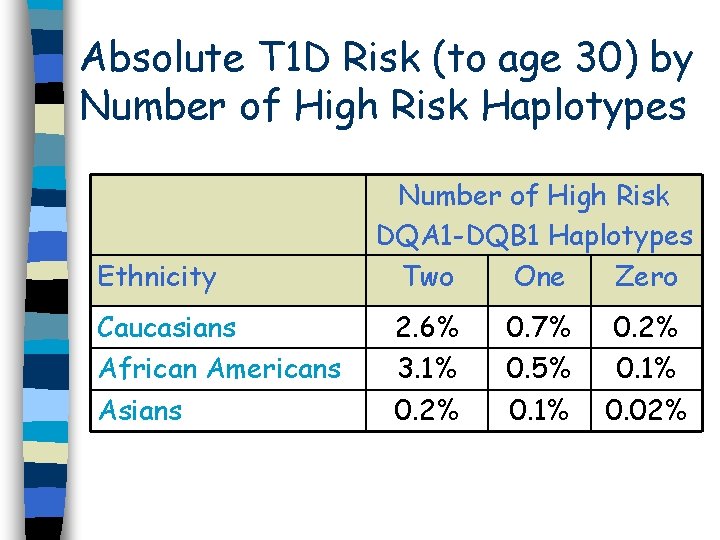
Absolute T 1 D Risk (to age 30) by Number of High Risk Haplotypes Ethnicity Caucasians African Americans Asians Number of High Risk DQA 1 -DQB 1 Haplotypes Two One Zero 2. 6% 3. 1% 0. 2% 0. 7% 0. 5% 0. 1% 0. 2% 0. 1% 0. 02%

Absolute T 1 D Risk for Siblings of Affected Individuals Number of High Risk DQA 1 -DQB 1 Haplotypes Risk of developing T 1 D Two One Zero 25% 8. 3% 1%
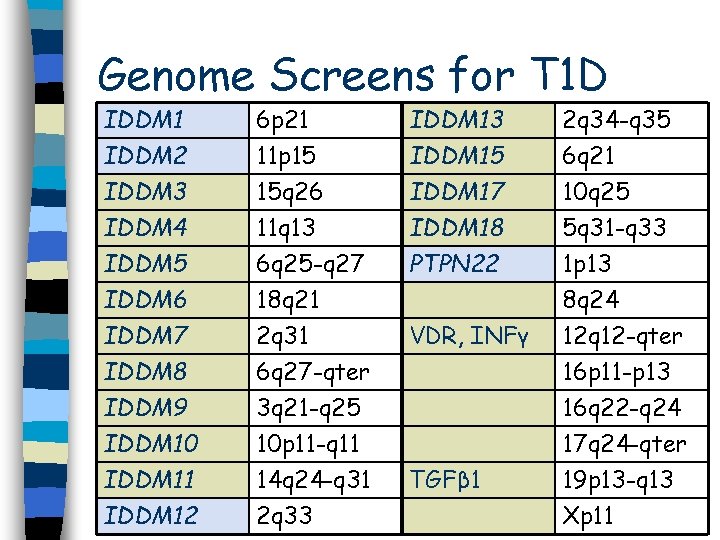
Genome Screens for T 1 D IDDM 1 IDDM 2 IDDM 3 IDDM 4 6 p 21 11 p 15 15 q 26 11 q 13 IDDM 15 IDDM 17 IDDM 18 2 q 34 -q 35 6 q 21 10 q 25 5 q 31 -q 33 IDDM 5 6 q 25 -q 27 PTPN 22 1 p 13 IDDM 6 IDDM 7 IDDM 8 IDDM 9 IDDM 10 IDDM 11 IDDM 12 18 q 21 2 q 31 6 q 27 -qter 3 q 21 -q 25 10 p 11 -q 11 14 q 24 -q 31 2 q 33 VDR, INFγ TGFβ 1 8 q 24 12 q 12 -qter 16 p 11 -p 13 16 q 22 -q 24 17 q 24 -qter 19 p 13 -q 13 Xp 11

IDDM 2 n Insulin (INS) gene n Chromosome 11 p 15, OMIM: 176730 n Variable number of tandem repeats (VNTR) n Class I is associated with lower m. RNA in the thymus – may reduce tolerance to insulin and its precursors – – Class I: 26 -63 repeats Class II: ~80 repeats Class III: 141 -209 repeats Relative increase in risk ~2 -fold with two class I alleles (compared to 0 class I alleles)

IDDM 12 n Cytotoxic T Lymphocyte Associated-4 (CTLA-4) n Chromosome 2 q 33, OMIM: 123890 – ICOS and CD 28 flank n Encodes a T cell receptor that plays are role in T cell apoptosis – A 49 G polymorphism (Thr 17 Ala) – Relative increase in risk ~ 1. 2 n Dysfunction of CTLA-4 is consistent with development of T 1 D

PTPN 22 n Lymphoid specific tyrosine phosphatase (LYP) n Chromosome 1 p 13, OMIM: 600716 n Encodes a LPY that is important in negative T-cell activation and development – C 858 T polymorphism (Arg 620 Trp) – Relative increase in risk ~ 1. 8 n May alter binding of LYP to cytoplasmic tyrosine kinase, which regulates the T-cell receptor signaling kinases

Intervention Trials for T 1 D Study TRIGR DIPP Trial. Net Intervention Avoid CM Insulin (N) Immunosuppressive agents Target /Screen FDR / genetic GP / genetic FDR / antibodies and genetic CM = cows milk, N = nasal, FDR = first degree relatives, GP = general population

Natural History Studies for T 1 D n Conducted in the general population – DAISY - Colorado – PANDA - Florida – TEDDY – US and Europe n Based on newborn genetic screening – Concerns about proper informed consent – Parents are notified of the results by mail – General population at ‘high’ risk (5 -8%) recruited for follow-up >50% of children who will develop T 1 D not eligible

Genetics and Prevention of T 1 D n Type 1 diabetes cannot be prevented n Ethical concerns regarding genetic testing for T 1 D, especially in children n Education programs are need for parents who consent to have their children involved in such studies because risk estimation is – Dependent on genes/autoantibodies used for assessment – Is not sensitive or specific
Type 2 Diabetes (T 2 D)

Type 2 Diabetes n Is group of genetically heterogeneous metabolic disorders that cause glucose intolerance – Involves impaired insulin secretion and insulin action n ~90% of individuals with diabetes have T 2 D n Considerations n Polygenic and multifactorial – May be treated with diet / oral medications / physical activity – T 2 D individuals may be asymptomatic for many years – Associated with long-term complications – Caused by multiple genes that may interact – Caused by genetic and environmental risk factors
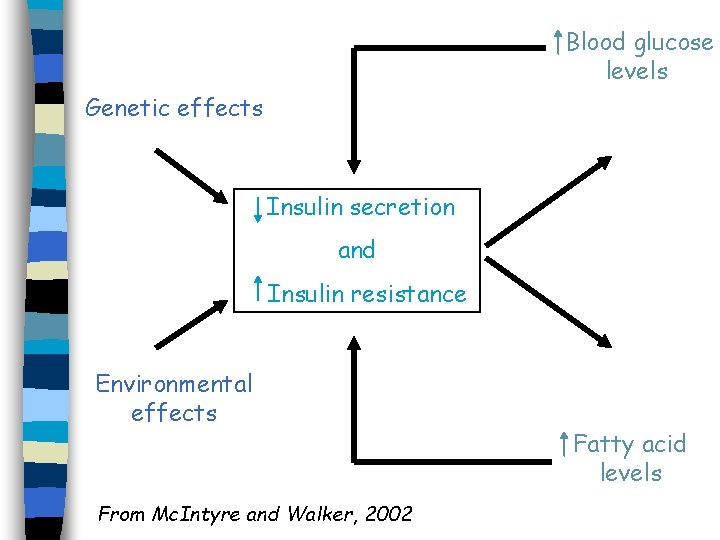
Blood glucose levels Genetic effects Insulin secretion and Insulin resistance Environmental effects From Mc. Intyre and Walker, 2002 Fatty acid levels

Thrifty Genotype n Had a selective advantage n In primitive times, individuals who were ‘metabolically thrifty’ were – Able to store a high proportion of energy as fat when food was plentiful – More likely to survive times of famine n In recent years, most populations have n These changes likely explain the rise in T 2 D worldwide – A continuous supply of calorie-dense processed foods – Reduced physical activity

Revised Classification Criteria for T 2 D n Fasting plasma glucose – > 7. 0 mmol/L – > 126 mg/dl n Random blood glucose – > 11. 1 mmol/L – > 200 mg/dl
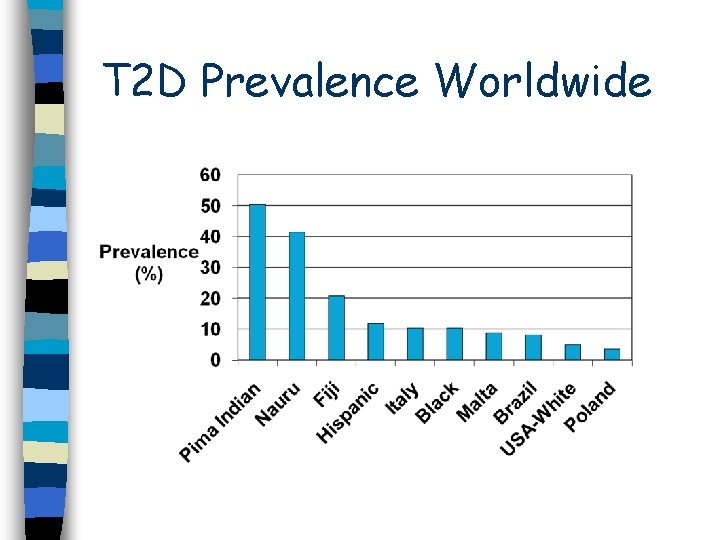
T 2 D Prevalence Worldwide
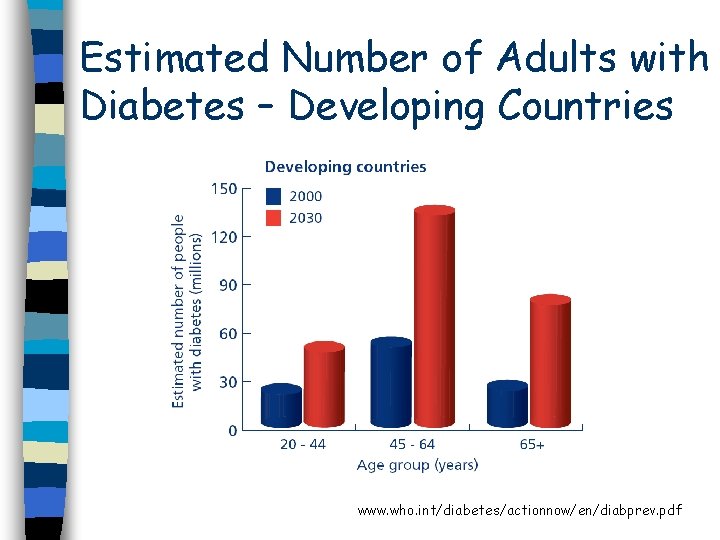
Estimated Number of Adults with Diabetes – Developing Countries www. who. int/diabetes/actionnow/en/diabprev. pdf
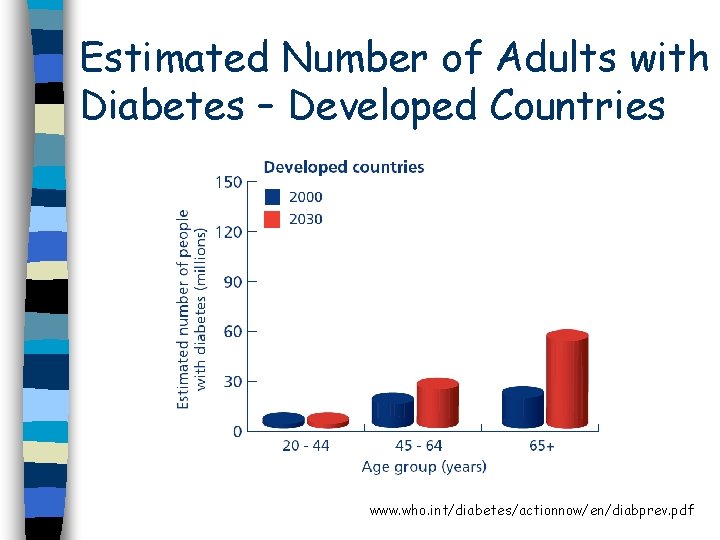
Estimated Number of Adults with Diabetes – Developed Countries www. who. int/diabetes/actionnow/en/diabprev. pdf
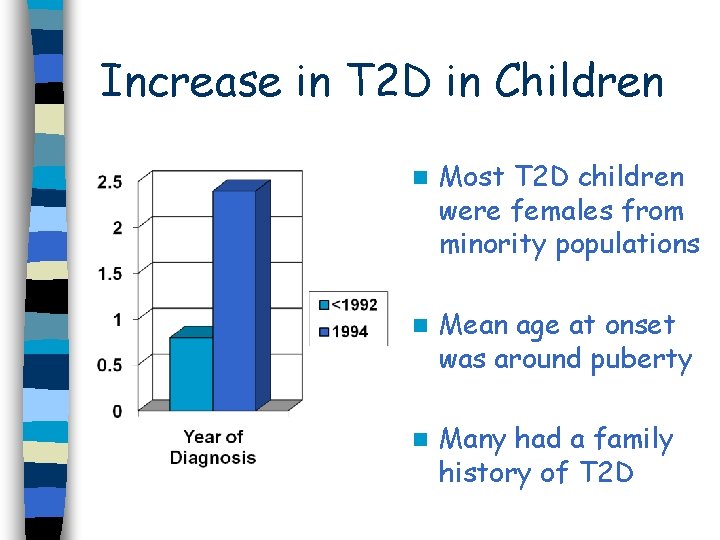
Increase in T 2 D in Children n Most T 2 D children were females from minority populations n Mean age at onset was around puberty n Many had a family history of T 2 D

Environmental Risk Factors in T 2 D n Obesity – Increases risk of developing T 2 D – Defined as: • > 120% of ideal body weight • Body mass index (BMI) > 30 k / m 2 – Likely related to the increase in T 2 D • ~80% newly diagnosed cases due to obesity – Higher association with abdominal or central obesity • Assessed by measuring the waist-to- hip ratio

Environmental Risk Factors in T 2 D n Physical Activity – Increases risk of developing T 2 D – Exercise • Controls weight • Improves glucose and lipid metabolism • Is inversely related to body mass index – Lifestyle interventions decreased risk of progression of impaired glucose tolerance to T 2 D by ~60%

Genetics and T 2 D n Individuals with a positive family history are about 2 -6 times more likely to develop T 2 D than those with a negative family history – Risk ~40% if T 2 D parent; ~80% if 2 T 2 D parents n Higher concordance for MZ versus DZ twins n Has been difficult to find genes for T 2 D – Late age at onset – Polygenic inheritance – Multifactorial inheritance

Finding Genes for T 2 D n Candidates selected because they are involved in – – n Pancreatic beta cell function Insulin action / glucose metabolism Energy intake / expenditure Lipid metabolism Genome wide screens – Nothing is assumed about disease etiology n Genome wide association studies – Current approach based on thousands of cases and controls

Challenges in Finding Genes n Inadequate sample sizes – Multiplex families – Cases and controls Difficult to define the phenotype n Reduced penetrance n – Influence of environmental factors – Gene-gene interactions Variable age at onset n Failure to replicate findings n Genes identified have small effects n

CAPN 10 – NIDDM 1 n Chromosome 2 q 37. 3 (OMIM 601283) – Encodes an intracellular calcium-dependent cytoplasmic protease that is ubiquitously expressed • May modulate activity of enzymes and/or apoptosis – Likely involves insulin secretion and resistance – Stronger influence in Mexican Americans than other ethnic groups • Responsible for ~40% if familial clustering – Genetic variant: A 43 G, Thr 50 Ala, Phe 200 Thr – Estimated relative risk: ~2

PPARγ n Peroxisome proliferator-activated receptor -γ (chromosome 3 p 25, OMIM: 601487) – Transcription factors that play an important role in adipocyte differentiation and function – Is associated with decreased insulin sensitivity – Target for hypoglycemic drugs thiazolidinediones – Genetic variant: Pro 12 Ala, Pro is risk allele (common) – Estimated relative risk = 1 - 3 – Variant is common – May be responsible for ~25% of T 2 D cases

ABCC 8 and KCNJ 11 n ATP-binding cassette, subfamily C member 8 (chromosome 11 p 15. 1, OMIM 600509) n Potassium channel, inwardly rectifying, subfamily J, member 11 (chromosome 11 p 15. 1, OMIM 600937) – ABCC 8 encodes the sulfonylurea receptor (drug target ) – Is coupled to the Kir 6. 2 subunit (encoded by KCNJ 11 – 4. 5 kb apart & near INS ) – Part of the ATP-sensitive potassium channel • Involved in regulating insulin and glucagon • Mutations affect channel’s activity and insulin secretion – Site of action of sulfonylureal drugs – Genetic variants: Ser 1369 Ala & Glu 23 Lys, respectively – Estimated relative risk = 2 – 4

TCF 7 L 2 n Transcription factor 7 -like 2 (chromosome 10 q 25, OMIM 602228) – Related to impaired insulin release of glucagon-like peptide-1 (islet secretagogue), reduced β-cell mass or β-cell dysfunction • Stronger among lean versus obese T 2 D – 10% of individuals are homozygous have 2 -fold increase in risk relative to those with no copy of the variant – Responsive to sulfunynlureals not metformin – Genetic variant: re 7901695 and others in LD – Estimated relative risk ~ 1. 4

GWAS New Loci Identified n FTO – chr 16 q 12 – Fat mass and obesity associated gene – Governs energy balance; gene expression is regulated by feeding and fasting – Estimated relative risk ~ 1. 23 n HHEX/IDE – chr 10 q 23 -24; near TCF 7 L 2 – HHEX - Haematopoietically expressed homeobox • Transcription factor in liver cells – IDE - Insulin degrading enzyme • Has affinity for insulin; inhibits IDE-mediated degradation of other substances – Estimated relative risk ~ 1. 14

GWAS New Loci Identified n CDKAL 1 – chr 6 p 22 – Cyclin-dependent kinase regulatory subunit associated protein 1 -like 1 – Likely plays role in CDK 5 inhibition and decreased insulin secretion – Estimated relative risk ~ 1. 12 n SLC 30 A 8 – chr 8 q 24 – Solute carrier family 30 zinc transporter – May be major autoantigen for T 1 D – Estimated relative risk ~ 1. 12

GWAS New Loci Identified n IGF 2 BP 2 – chr 3 q 28 – Insulin-like growth factor 2 m. RNA binding protein 2 – Regulates IGF 2 translation; stimulates insulin action – Estimated relative risk ~ 1. 17 n CDKN 2 A/B – chr 9 p 21 – Clycin dependent kinase inhibitor 2 A – Plays role in pancreatic development and islet proliferation – Estimated relative risk ~ 1. 2

T 2 D Genes are Drug Targets n PPARγ, ABCC 8 and KCNJ 11 are the targets of drugs used routinely in the treatment of T 2 D – Pharmacogenetic implications – Response to oral agents may be related to one’s genotype – Genetic testing may • Identify individuals at high risk for T 2 D • Guide treatment regimens for T 2 D – Individualize therapy

Genetics and Prevention of T 2 D n T 2 D is preventable – Maintaining age-appropriate body weight – Physical activity n New genes will provide insight to etiology n Public health messages may have a greater influence on genetically susceptible n Will genetic testing prevent T 2 D? – Unclear whether knowledge of one’s genetic risk will lead to behavior modifications

Genetics and Prevention of T 2 D n Challenges include: – Predictive values of most test is low – How to communicate risk information? – Health care professionals may not be able to interpret genetic tests – Genetic testing may lead to distress, etc. – Insurance and employment discrimination – Confidentiality and stigmatization – Direct to consumer marketing for genetic testing
Maturity Onset Diabetes of the Young (MODY)

MODY n Account for ~ 5% of type 2 diabetes n Single gene defects n Early age at onset (< age 25 years) n Characterized by the absence of obesity, no ketosis and no evidence of beta cell autoimmunity n Hyperglycemia often corrected by diet – Autosomal dominant inheritance – Multiple generations affected
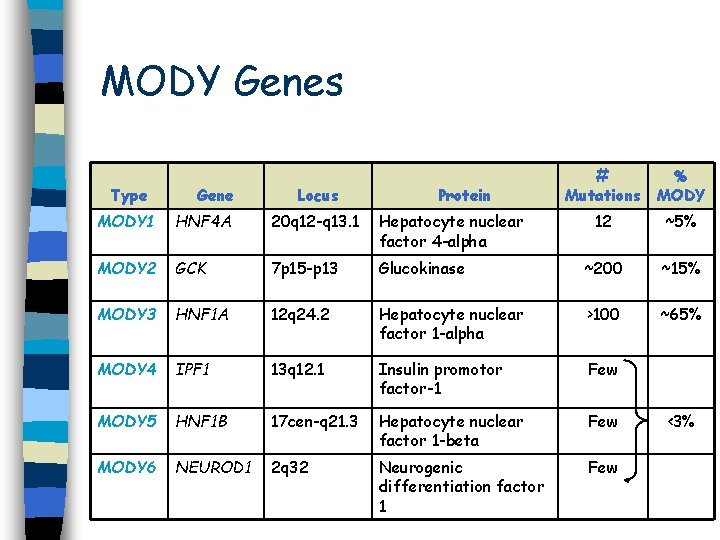
MODY Genes Type Gene Locus Protein # Mutations % MODY 12 ~5% MODY 1 HNF 4 A 20 q 12 -q 13. 1 Hepatocyte nuclear factor 4 -alpha MODY 2 GCK 7 p 15 -p 13 Glucokinase ~200 ~15% MODY 3 HNF 1 A 12 q 24. 2 Hepatocyte nuclear factor 1 -alpha >100 ~65% MODY 4 IPF 1 13 q 12. 1 Insulin promotor factor-1 Few MODY 5 HNF 1 B 17 cen-q 21. 3 Hepatocyte nuclear factor 1 -beta Few MODY 6 NEUROD 1 2 q 32 Neurogenic differentiation factor 1 Few <3%

MODY 1 is HNF 4 A (hepatocyte nuclear factor 4 -alpha) on 20 q 12 -q 13. 1 n Transcription factor – Expressed in the liver, kidney, intestine and pancreatic islet cells – Has been associated with T 2 D n Controls genes involved in glucose, cholesterol and fatty acid metabolism n Controls transcription of HNF 1 A (MODY 3) n Several mutations/splicing defects identified – Account for ~5% of all MODY cases

MODY 2 is GCK (glucokinase) on 7 p 15 -p 13 n Only MODY gene that is not a transcription factor n Required for glucose metabolism and insulin secretion; acts as a glucose ‘sensor’ n MODY 2 is generally a mild form of diabetes n ~ 200 mutations have been identified – VNTR, nonsense and missense mutations – Account for ~15% of all MODY cases

MODY 3 is HNF 1 A (hepatocyte nuclear factor 1 -alpha) on 12 q 24. 2 n Regulates expression of insulin and other genes involved in glucose transport / metabolism – Influences expression of HNF 4 A (MODY 1) n Results in a severe insulin secretory defect n More than 100 genetic variants have been identified n Mutations in MODY 3 are the most common cause of MODY – May contribute to abnormal islet cell development – Account for ~65% of all MODY cases – Sensitive to sulphonylureas

MODY 4 is IPF 1 (insulin promoter factor-1) on 13 q 12. 1 n Transcription factor that regulates expression of insulin, somatostatin and other genes – Involved in the development of the pancreas – In adults, expressed only in pancreatic cells n Mutations lead to decreased binding activity to the insulin promoter – Reduced activation of insulin gene in response to glucose n Genetic variants include frameshift, insertions and missense mutations – Accounts for a very small proportion of MODY cases

MODY 5 is HNF 1 B (hepatocyte nuclear factor 1 -beta) on 17 cen-q 21. 3 n Transcription factor required for liver-specific expression of a variety of genes n Is highly homologous to HNF 1 A (MODY 3) n HNF 1 A and HNF 1 B likely interact to regulate gene expression n Individuals have lower renal threshold to glucose n Is a rare cause of MODY – Recognizes same binding site as HNF 1 A

MODY 6 is NEUROD 1 (neurogenic differentiation factor 1) on 2 q 32 n Is a transcription factor involved in the differentiation of neurons n Regulates insulin gene expression by binding to a critical motif on the insulin promoter n Few genetic variants identified – Missense and nonsense mutations – Account for ~1% of all MODY cases

Summary of MODY Genetics n All MODY genes are expressed in the pancreas, and play a role in: – The metabolism of glucose – The regulation of insulin or other genes involved in glucose transport – The development of the fetal pancreas n MODY phenotype depends on the MODY genotype (on next slide) n Knowing the genotype is important to determine treatment
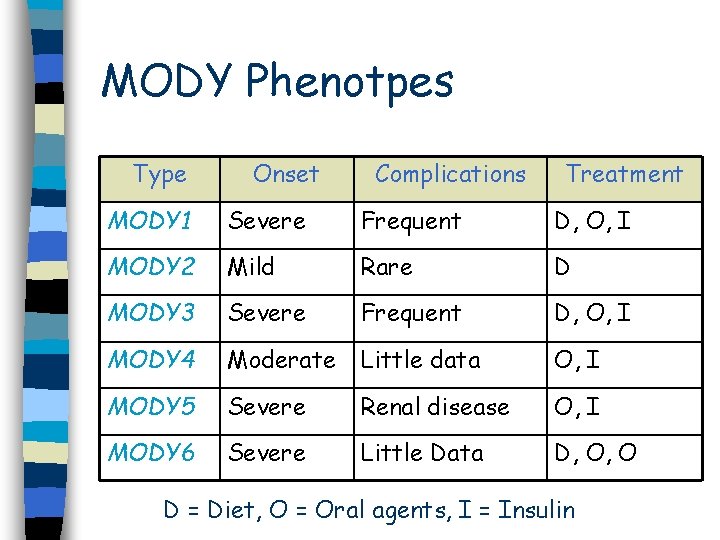
MODY Phenotpes Type Onset Complications Treatment MODY 1 Severe Frequent D, O, I MODY 2 Mild Rare D MODY 3 Severe Frequent D, O, I MODY 4 Moderate Little data O, I MODY 5 Severe Renal disease O, I MODY 6 Severe Little Data D, O, O D = Diet, O = Oral agents, I = Insulin
- Slides: 54