Genetics Lec 3 Dr Mohammed Hussein M B

Genetics Lec. 3 Dr. Mohammed Hussein M. B. Ch. B, MSC, Ph. D, DCH (UK), MRCPCH

Important principles that can characterize single-gene diseases 1. 2. 3. 4. 5. 6. 7. 8. 9. Variable expression Incomplete penetrance Pleiotropy Locus heterogeneity New mutations Delayed age of onset Anticipation Imprinting Uniparental disomy

Variable Expression • Most genetic diseases vary in the degree of phenotypic expression: Some individuals may be severely affected, whereas others are more mildly affected. • This can be the result of several factors: 1. 2. 3. 4. Environmental Influences Allelic Heterogeneity Heteroplasmy Modifier Loci
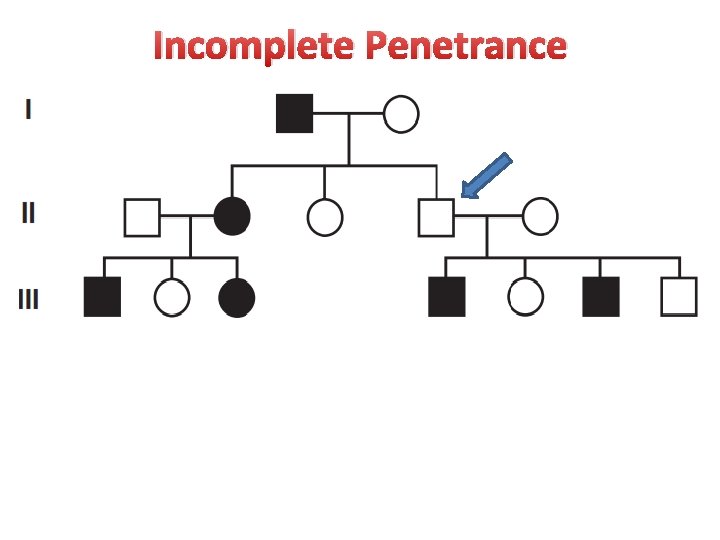
Incomplete Penetrance

Pleiotropy • Pleiotropy exists when a single disease-causing mutation affects multiple organ systems. • Pleiotropy is a common feature of genetic diseases. • Marfan syndrome provides a good example of the principle of pleiotropy
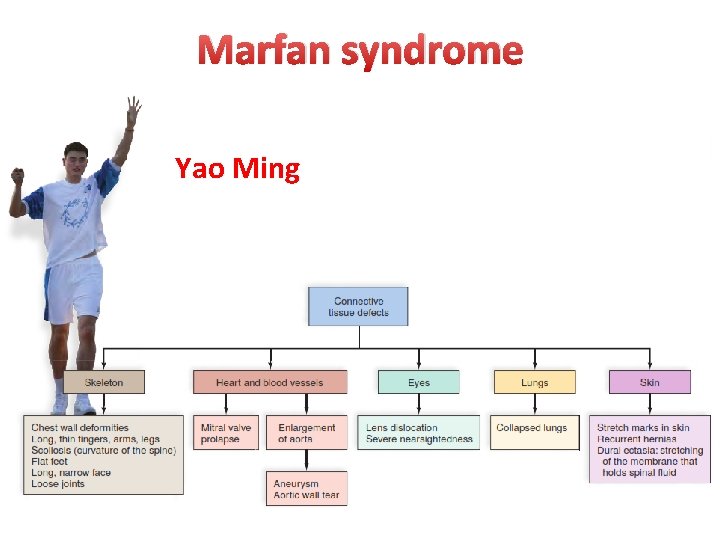
Marfan syndrome Yao Ming

Locus Heterogeneity • Locus heterogeneity exists when the same disease phenotype can be caused by mutations in different loci. • For example, retinitis pigmentosa has autosomal dominant, autosomal recessive, and X-linked origins.
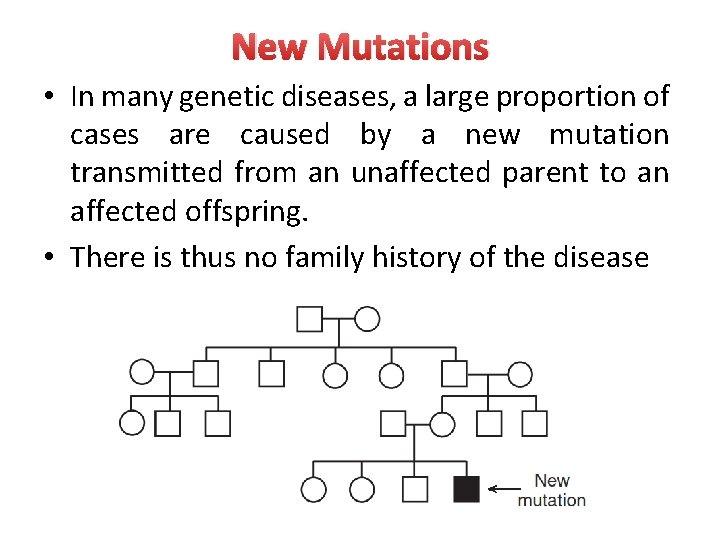
New Mutations • In many genetic diseases, a large proportion of cases are caused by a new mutation transmitted from an unaffected parent to an affected offspring. • There is thus no family history of the disease

Delayed Age of Onset • Many individuals who carry a disease-causing mutation do not manifest the phenotype until later in life. • This can complicate the interpretation of a pedigree because it may be difficult to distinguish genetically normal individuals from those who have inherited the mutation but have not yet displayed the phenotype. • Example: Familial breast cancer

Genetic Anticipation • Anticipation refers to a pattern of inheritance in which individuals in the most recent generations of a pedigree develop a disease at an earlier age or with greater severity than do those in earlier generations. • For a number of genetic diseases, this phenomenon can be attributed to the gradual expansion of trinucleotide repeat polymorphisms within or near a coding gene.
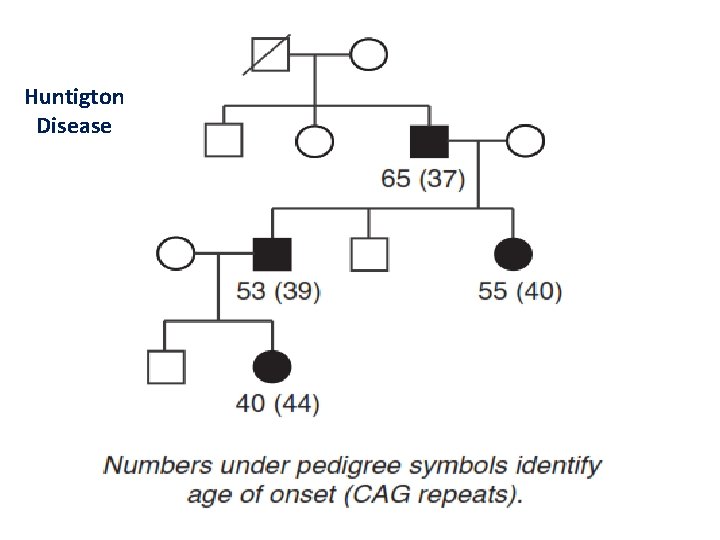
Huntigton Disease

Imprinting • Imprinting refers to the fact that a small number of genes are transcriptionally active only when transmitted by one of the two sexes. • The homologous locus in the other parent is rendered transcriptionally inactive. • Thus, for imprinted loci, it is normal to have only the maternal (for some loci) active, or only the paternal (for other loci) active.
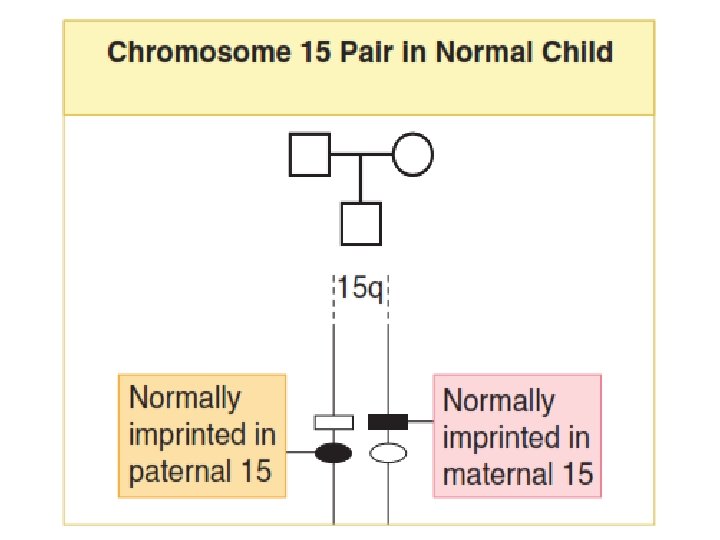

• On rare occasion, the transcriptionally active gene may be deleted from the chromosome during gametogenesis. • This leaves the offspring with no active gene at that locus. • The gene from one parent is inactivated due to normal imprinting, and the gene from the other parent deleted by a mutation. • This situation may result in a genetic disease.

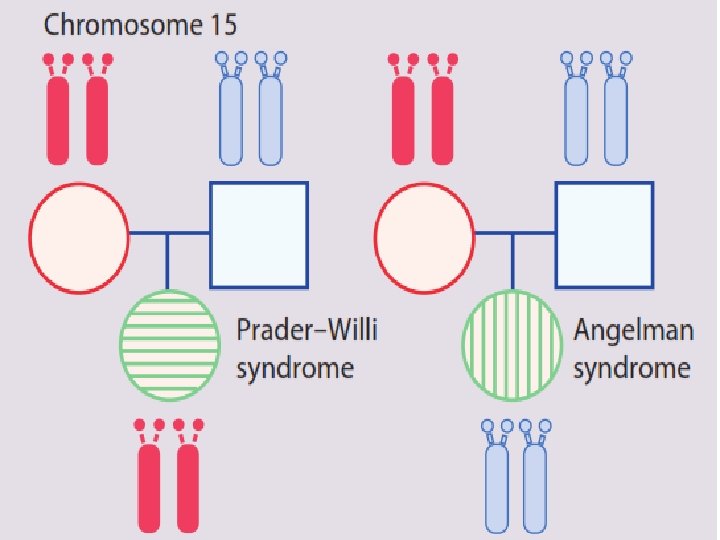
Prader-Willi Syndrome and Angelman Syndrome
Prader-Willi Syndrome

Angelman Syndrome

Uniparental Disomy • Uniparental disomy is a rare condition in which both copies of a particular chromosome are contributed by one parent.
Population Genetics

Definition • Population genetics is the study of genetic variation in populations. • Allow us to understand how and why the prevalence of various genetic diseases differs among populations.

: British Mathematician : German Physician

Hardy-Weinberg Equilibrium

• If a population is large and if individuals mate at random with respect to their genotypes at a locus, the population should be in H-W equilibrium.
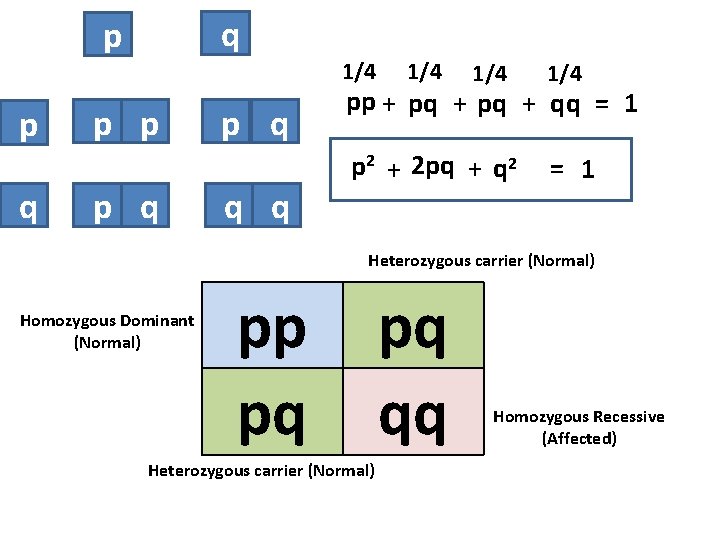
q p p q 1/4 p p p q q q 1/4 1/4 pp + pq + qq = 1 p 2 + 2 pq + q 2 = 1 Heterozygous carrier (Normal) Homozygous Dominant (Normal) pp pq Heterozygous carrier (Normal) pq qq Homozygous Recessive (Affected)

2 P + 2 pq + 2 q =1 p = frequency the normal allele (most common) q = frequency the disease-producing allele (minor) p 2 = frequency of homozygous normal q 2= frequency of homozygous affected (diseased) 2 pq = frequency of heterozygous carrier

Simplification • Generally p, the normal allele frequency in the population, is very close to 1 (e. g. , most of the alleles of this gene are normal). • In this case, we may assume that p ~ 1, and the equation simplifies to: 2 P + 2 pq + 1 + 2 q + 2 q =1 =1

• In summary, there are three major terms one usually works with in the Hardy Weinberg equation applied to AR conditions: 1. q, the frequency of the disease-causing allele 2. 2 q, the carrier frequency 3. q 2, the disease prevalence

A Practical Application of the H-W Principle • A 20 -year-old female has an AR genetic disease called Phenylketonuria (PKU). She asks you 2 questions: 1. What is the chance that she had to marry a man with the disease-producing allele. 2. What is the probability that she will have a child with PKU? • Note: the prevalence of PKU in the population is 1/10, 000.

Answer q = disease-causing allele 2 q = carrier frequency q 2 = disease prevalence • Disease prevalence = 1/10, 000 (given) • So q 2 = 1/10, 000 • So q = square root of 1/10, 000, which is 1/100. This is the frequency of the disease producing allele • Carrier frequency = 2 q = 2 x 1/100 = 1/50 • The chance that she had to marry a man with the disease-producing allele is 1/50

2. What is the probability that she will have a child with PKU? • The answer to the second question is based on the joint occurrence of 4 events: 1. The probability that the female carry the abnormal allele 2. The probability that the she will pass her abnormal allele to the child 3. The probability that the male carry the abnormal allele 4. The probability that the he will pass his abnormal allele to the child 1 * 2 * 3 * 4 = the risk of having a child with the disease

• The answer to the second question is based on the joint occurrence of 4 events: 1. The probability that the female carry the abnormal allele (1) as she is known affected 2. The probability that the she will pass her abnormal allele to the child (1) as the disease is AR 3. The probability that the male carry the abnormal allele (1/50) as calculated before 4. The probability that the he will pass his abnormal allele to the child (1/2) 1 * 1/50*1/2 = 1/100

• A couple come for genetic counseling about the risk that their child will have cystic fibrosis. • The husband's sister has the disease, but he is not affected and there is no history in the wife's family of the disease. • Note: the prevalence of CF is 4/10000.

Answer q = disease-causing allele 2 q = carrier frequency q 2 = disease prevalence • Prevalence = 4/10000 = q 2 • So q = 2/100 • So 2 q = 4/100 = 1/25 (carrier risk of CF in population) which is the risk for the wife. • Regarding the husband, as his sister is affected, so both parents must be carriers. Since he is not affected therefore there is a 2/3 chance that he is a carrier. 1/25 * 1/2 * 2/3 * 1/2 * = 1/150

• In AD diseases the homozygous and the heterozygous are the affected • The heterozygous affected are the most common, so the prevalence is equal to the heterozygous (2 q) • While the homozygous affected (q 2 ) are less common but have more sever symptoms pp pq pq qq

q = allele frequency 2 q = heterozygous affected = prevalence of the disease q 2 = homozygous affected (sever disease)

Example q = disease-causing allele 2 q = heterozygous affected (disease prevalence) q 2 = homozygous affected (sever disease) • The prevalence of hyperlipidemia (AD disease) in USA is 1/500, calculate the homozygous affected. • As the disease is AD, so the • Prevalence = 2 q, so q= 1/1000 • q 2 = 1/100000 = 1/106 = homozygous affected (severely affected).

• Most cases occur in hemizygous males (x. Y). • So, q = disease-producing allele frequency also equals the prevalence of affected males. • q 2= prevalence of disease in females. (very rare) • 2 q = prevalence of female carriers.

q = allele frequency = prevalence of affected males 2 q = heterozygous female carriers q 2 = homozygous affected females

Example q = disease-causing allele (disease prevalence in male) 2 q = heterozygous female carrier q 2 = homozygous affected female (sever disease) • The prevalence of hemophilia in males is 1/10, 000, calculate the prevalence of female carriers and the females affected. • The prevalence of the disease = q = 1/10, 000 • 2 q is the female carriers = 2/10, 000. • q 2 is the affected females = 1/10000 = 1/108

q AR diseases: Ø q = allele frequency Ø 2 q = heterozygous carriers Ø q 2 = homozygous affected = prevalence of the disease q AD diseases: Ø q = allele frequency Ø 2 q = heterozygous affected = prevalence of the disease Ø q 2 = homozygous affected (sever disease) q X-L diseases: Ø q = allele frequency = prevalence of affected males Ø 2 q = heterozygous female carriers Ø q 2 = homozygous affected females

• Although human populations are typically in H -W equilibrium for most loci, deviations from equilibrium can be produced by: 1. Mutation 2. Natural Selection 3. Genetic Drift 4. Gene Flow 5. Consanguinity

• Mutation is ultimately the source of all new genetic variation in populations. • Founder effect

Natural Selection

• Natural selection acts upon genetic variation, increasing the frequencies of alleles that promote survival or fertility (referred to as fitness) and decreasing the frequencies of alleles that reduce fitness. q Cystic fibrosis (heterozygote resistance to typhoid fever) q Hemochromatosis (heterozygote advantage in ironpoor environments) q Glucose-6 -phosphate dehydrogenase deficiency (heterozygote resistance to malaria)

• Mutation rates do not vary significantly from population to population, although they can result in significant differences in allele frequencies when they occur in small populations

• Gene flow refers to the exchange of genes among populations. • Because of gene flow, populations located close to one another often tend to have similar gene frequencies

• Consanguinity refers to the mating of individuals who are related to one another (typically, a union is considered to be consanguineous if it occurs between individuals related at the second -cousin level or closer). • Consequently, there is an increased risk of genetic disease in the offspring of consanguineous matings


Determining type of inheritance in a pedigree • Ask yourself the following questions 1. Dose an affected individual have an affected parent? 2. 3. 4. o Yes: Dominant disease o No: Recessive disease If it is Recessive, then ask: Dose all affected are males? o Yes: X-linked recessive disease o No: Autosomal recessive disease If it is Dominant, then ask: Is there male-male transmission? o Yes: autosomal dominant disease o No: may be an X-linked dominant Are all daughters of an affected male also affected? o Yes: X-linked dominant o No: autosomal dominant disease
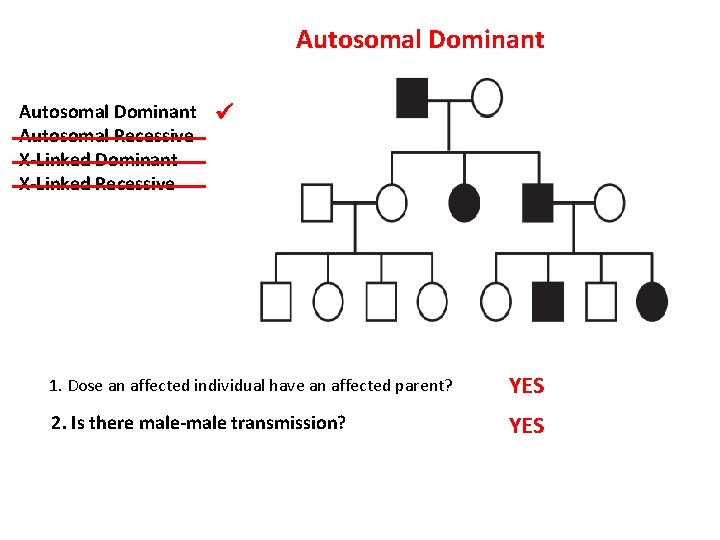
Autosomal Dominant Autosomal Recessive X-Linked Dominant X-Linked Recessive 1. Dose an affected individual have an affected parent? YES 2. Is there male-male transmission? YES
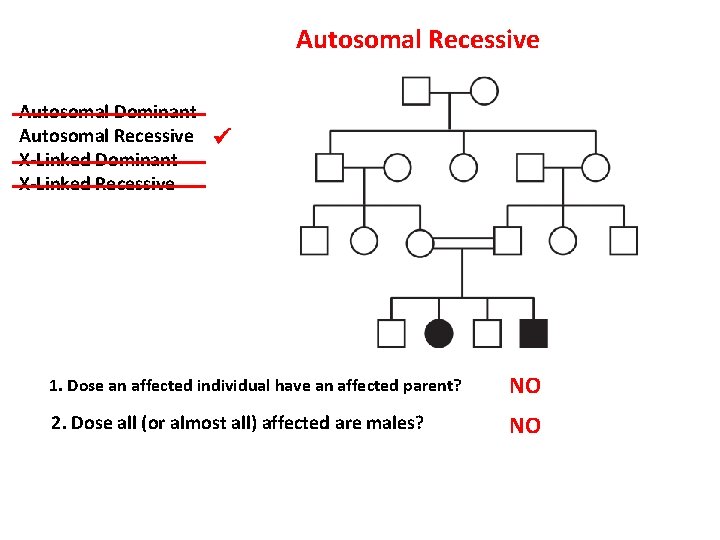
Autosomal Recessive Autosomal Dominant Autosomal Recessive X-Linked Dominant X-Linked Recessive 1. Dose an affected individual have an affected parent? NO 2. Dose all (or almost all) affected are males? NO
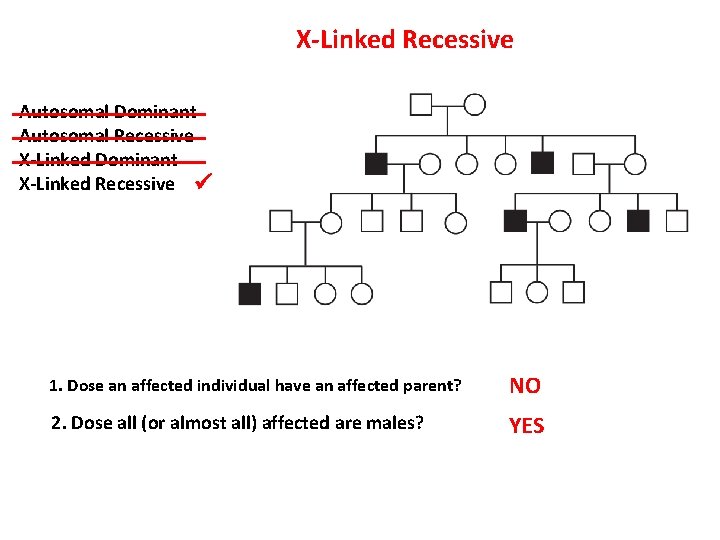
X-Linked Recessive Autosomal Dominant Autosomal Recessive X-Linked Dominant X-Linked Recessive 1. Dose an affected individual have an affected parent? NO 2. Dose all (or almost all) affected are males? YES
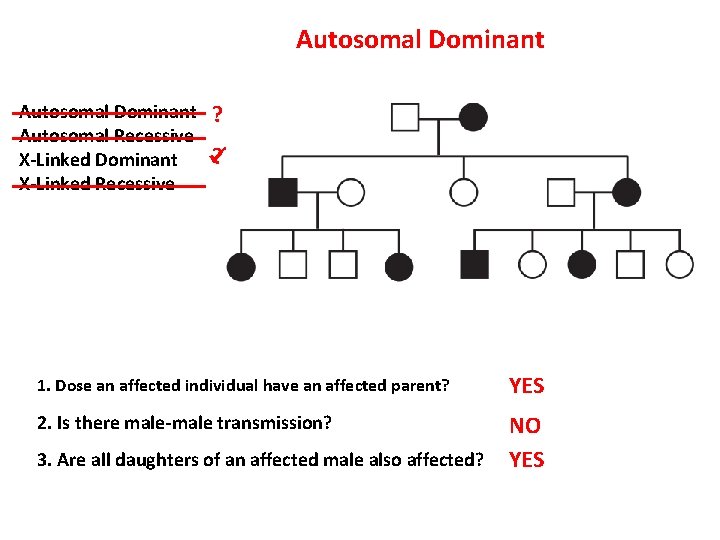
Autosomal Dominant Autosomal Recessive X-Linked Dominant X-Linked Recessive ? ? 1. Dose an affected individual have an affected parent? YES 2. Is there male-male transmission? NO YES 3. Are all daughters of an affected male also affected?
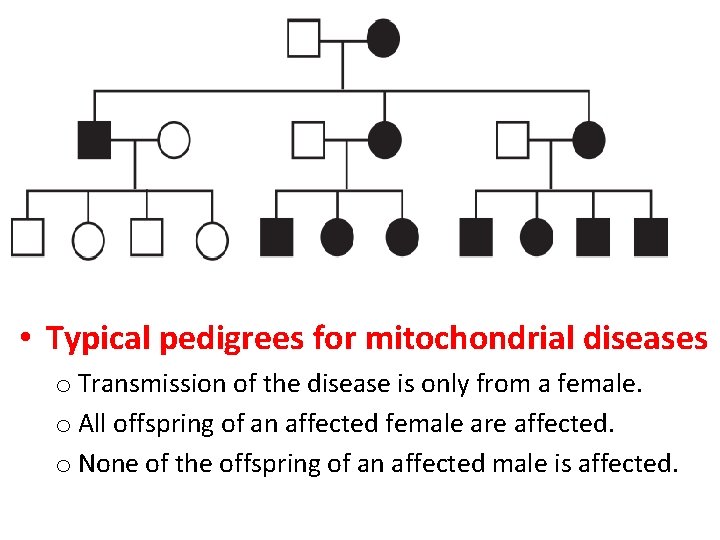
• Typical pedigrees for mitochondrial diseases o Transmission of the disease is only from a female. o All offspring of an affected female are affected. o None of the offspring of an affected male is affected.

Dr. Mohammed Hussein M. B. Ch. B, MSC, Ph. D, DCH (UK), MRCPCH
- Slides: 56