GENETICS genetic mapping classical approaches to study gene

























- Slides: 64

GENETICS genetic mapping, classical approaches to study gene function

Basic aims: • uncovering gene function understand mechanisms of morphogenesis, development, metabolism, physiology etc. in connection with coordinated gene expression • breeding production of plants (organisms) with improved characteristics or their combination

Terminology Gene • segment of DNA (genetic information) that specifies a trait (is a predisposition for a trait) • basic unit of heridity in living organisms • Genotype + environment + ? = phenotype • Interactions between genes/proteins (epistases – mainly metabolic and signal pathways)

Allele – form of a gene • dominant vs. recesive (codominant, …) • genesis of new alleles by mutations Locus – location of a gene on a chromosome • Genetic linkage – inheriting of certain genes (their alleles) jointly, - consequence of their presence on the same chromosome (gene distance c. M = % of recombinant gametes) HOW MANY alleles of certain gene is in the genome? of alleles = from 1 up to N = ploidy (* of loci with the gene)

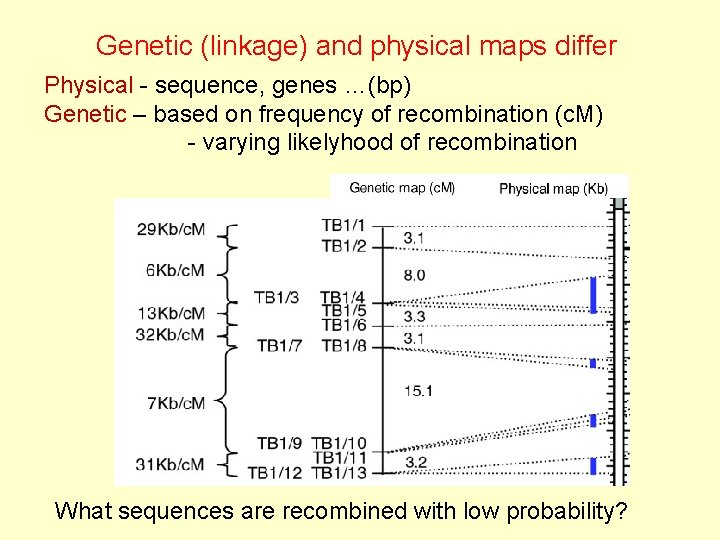
Genetic (linkage) and physical maps differ Physical - sequence, genes …(bp) Genetic – based on frequency of recombination (c. M) - varying likelyhood of recombination What sequences are recombined with low probability?

Genetics classical (direct) x reverse Direct – from a trait (phenotype) to identification of the corresponding gene Reverse – from a gene (prediction based on the sequence) to the gene’s function (phenotype) Both approaches are based on mutant analysis: Direct g. – looking for certain change in phenotype in the mutagenized Reverse g. – population (and analysing the genetic courses) targeted mutagenesis/modification of a selected gene (and analyzing the phenotypic consequences)

Reporter genes • encode proteins that can be directly visualized or enzymes, whose activity can be visualized • quantitative or qualitative assessment • promoter activity analyses, subcellular localization of proteins, optimizing transformation procedure, …. Constrains: - background (autofluorescence, natural enzyme activity within the tissue) - protein stability (mask changes in promoter activity) - …. .

Reporter genes (the most frequently used) gene product substrate detection gus. A b-glucuronidase (E. coli) MUG X-gluc luciferin xxx fluorescence histochemical luminiscence luc gfp luciferase (fire fly) green fluorescent protein fluorescence (gelly fish)

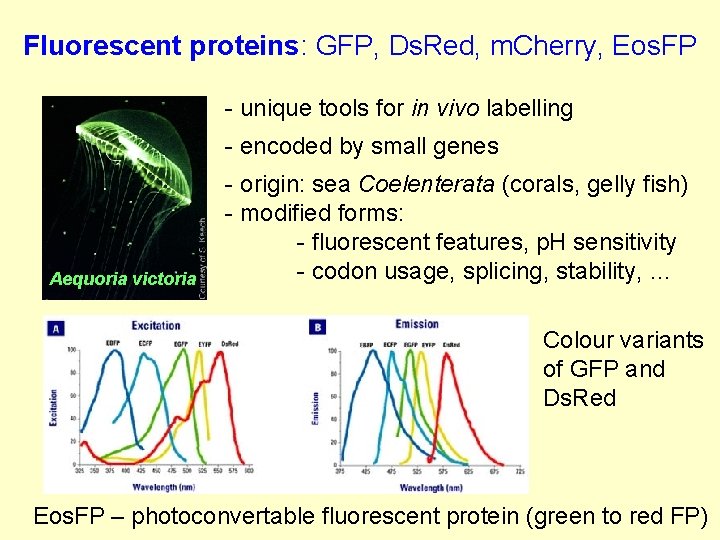
Fluorescent proteins: GFP, Ds. Red, m. Cherry, Eos. FP - unique tools for in vivo labelling - encoded by small genes Aequoria victoria - origin: sea Coelenterata (corals, gelly fish) - modified forms: - fluorescent features, p. H sensitivity - codon usage, splicing, stability, … Colour variants of GFP and Ds. Red Eos. FP – photoconvertable fluorescent protein (green to red FP)

GUS - glucuronidase Qualitative detection (X-gluc = 5 -bromo-4 -chloro-3 -indolyl glucuronide) • oxidized blue precipitate of the reaction product • low background!!! • slow diffusion • mostly in the fixed material Quantitative detection • GUS enzyme isolation, fluorimetric statement • highly sensitive, low background (MUG = 4 -methylumbelliferyl-beta-D-glucuronide)

Mutagenesis • Classical: – chemical m. – EMS (ethane metyl sulfonate; point mutations) – physical m. – RTG, gama. . . (usually short deletions) – wide spektrum of affects (regulation, interaction) – even dominant mutations, resemble natural mutations, – BUT complicated identification of mutated genes

Identification of mutated gene (responsible for the mutation) Point mutations, short deletions 1) Based on genetic map and segregation analysis + chromosom walking, sequencing (time, money and labor-consuming) 2) Using NGS (next generation seq. ) - faster, moderate expensive - even in unknown genomes!!! - mixed samples (back crosses) - comparisons of frequencies of similar oligomers Nordström et al. Nature Biotech. 2013 < > <

Mutagenesis • More advanced - insertional mutagenesis – random insertions in the genome! – by insertion of T-DNA of Agrobacterium tumefaciens (transposons) – allows simple determination of the site of insertion mutations are tagged with the inserted sequence – various strategies for gene isolation – advanced variants of insertional mutagenesis: • activation mutagenesis • promoter, enhancer trap, …

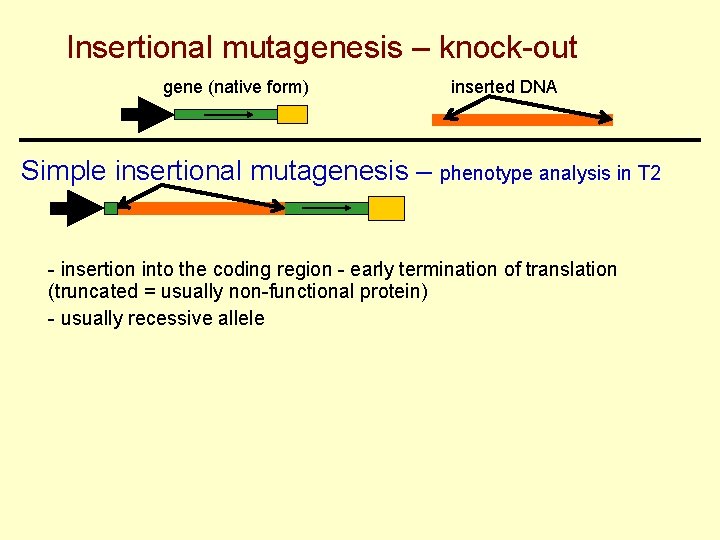
Insertional mutagenesis – knock-out gene (native form) inserted DNA Simple insertional mutagenesis – phenotype analysis in T 2 - insertion into the coding region - early termination of translation (truncated = usually non-functional protein) - usually recessive allele

Identification of mutated gene Insertional mutagenesis: • sequencing of the flanking region (too low template concentration for direct Sanger sequencing!) Technical solution (preamplification of the target locus): • • TAIL PCR (Thermal Asymmetric Inter. Laced PCR) adaptor PCR plasmid rescue i. PCR

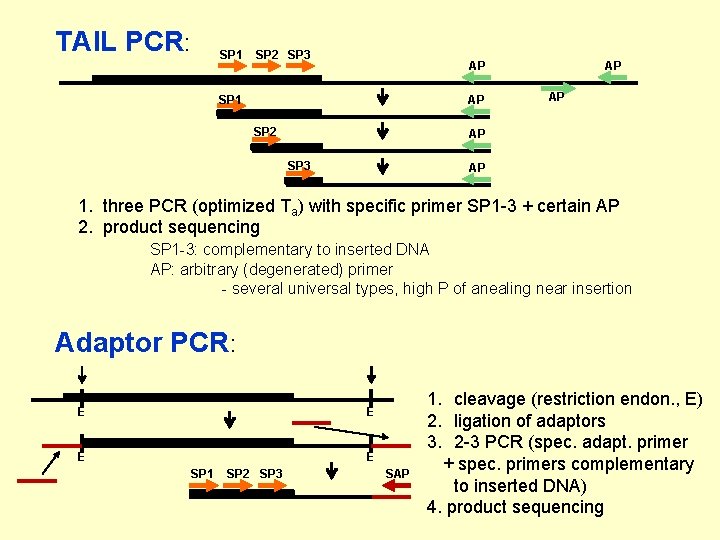
TAIL PCR: SP 1 SP 2 SP 3 AP SP 1 AP SP 2 AP AP AP SP 3 AP 1. three PCR (optimized Ta) with specific primer SP 1 -3 + certain AP 2. product sequencing SP 1 -3: complementary to inserted DNA AP: arbitrary (degenerated) primer - several universal types, high P of anealing near insertion Adaptor PCR: E E SP 1 SP 2 SP 3 SAP 1. cleavage (restriction endon. , E) 2. ligation of adaptors 3. 2 -3 PCR (spec. adapt. primer + spec. primers complementary to inserted DNA) 4. product sequencing

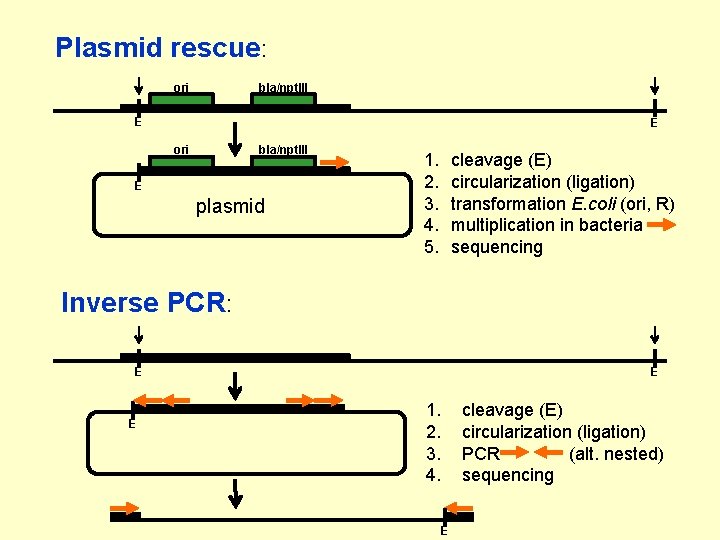
Plasmid rescue: ori bla/npt. III E E ori bla/npt. III E plasmid 1. 2. 3. 4. 5. cleavage (E) circularization (ligation) transformation E. coli (ori, R) multiplication in bacteria sequencing Inverse PCR: E E E 1. 2. 3. 4. E cleavage (E) circularization (ligation) PCR (alt. nested) sequencing

Mutagenesis Most advanced genome editing = targeted mutagenesis within the genome: ZFN all causing double stranded break TALEN in DNA CRISPR/Cas 9 Mutation: - usually inactivation (NHEJ – non homology end joining – random local mutations) - specific change of the sequence - driven by an oligonucleotide serving as a recombination template) - using modified CAS 9 (e. g. fused with C deaminase)

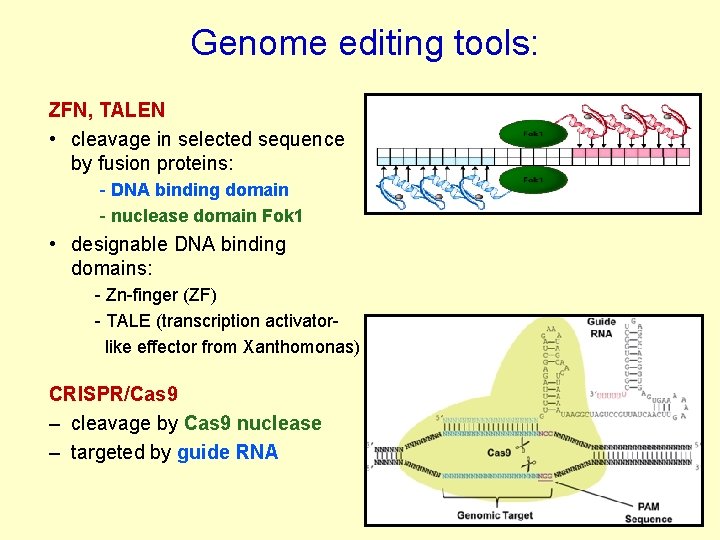
Genome editing tools: ZFN, TALEN • cleavage in selected sequence by fusion proteins: - DNA binding domain - nuclease domain Fok 1 • designable DNA binding domains: - Zn-finger (ZF) - TALE (transcription activatorlike effector from Xanthomonas) CRISPR/Cas 9 – cleavage by Cas 9 nuclease – targeted by guide RNA

Use of mutagenesis in genetics Direct: - classical mutagenesis - insertional mutagenesis (all types – see later) Reverse: - directed mutagenesis - also random insertional mutagenesis 1) generation + characterization of random insertional mutants 2) making a collection 3) ordering of mutant in selected gene from the collection

Approaches of reverse genetics - analyses of mutants with insertion in an analyzed gene - modulation of an analyzed gene expression - promoter activity analyses - analyses of subcellular protein localization (GFP fusion) - gene modification (within/outside the genome) - TILLING, …. . - analysis of the gene polymorphism in various ecotypes (1000+1 genome project)

RG 1: Analyses of insertion mutants (usually knock-out) - publicly available (Arabidopsis, rice, …) - insertions in different sites in the genome 80 % genes (A. th. ) for inactivation – optimally 5’ exons, experiment. confirmation is needed!) – mutant selection in silico, ordering seeds from the collection Gene 1 1 Gene 2 23 4 Gene 3 56 7 = sites of T-DNA insertions in individual lines (1 -8) 8 …line number

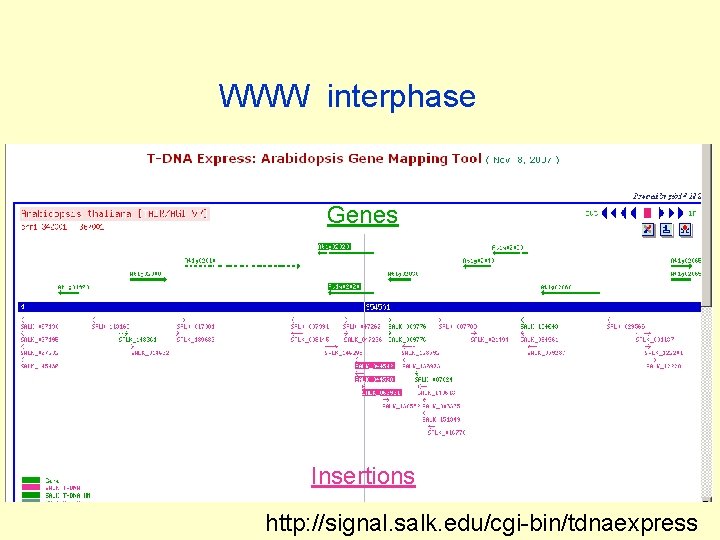
WWW interphase Genes Insertions http: //signal. salk. edu/cgi-bin/tdnaexpress

RG 2: Modulation of gene expression - increased m. RNA/protein level (overexpression) - introduction of a gene with a strong constitutive promoter (e. g. viral) - syn. gain-of-function mutations, ectopic expr. - decreased m. RNA/protein level by RNAi - silencing, knock-down - X knock–out, loss-of-function = complete inactivation by disruption of the gene

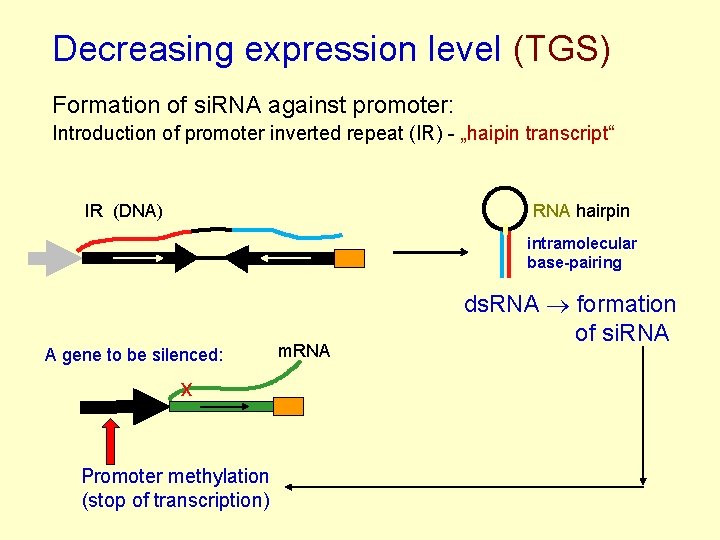
Decreasing expression level (TGS) Formation of si. RNA against promoter: Introduction of promoter inverted repeat (IR) - „haipin transcript“ IR (DNA) RNA hairpin intramolecular base-pairing A gene to be silenced: x Promoter methylation (stop of transcription) m. RNA ds. RNA formation of si. RNA

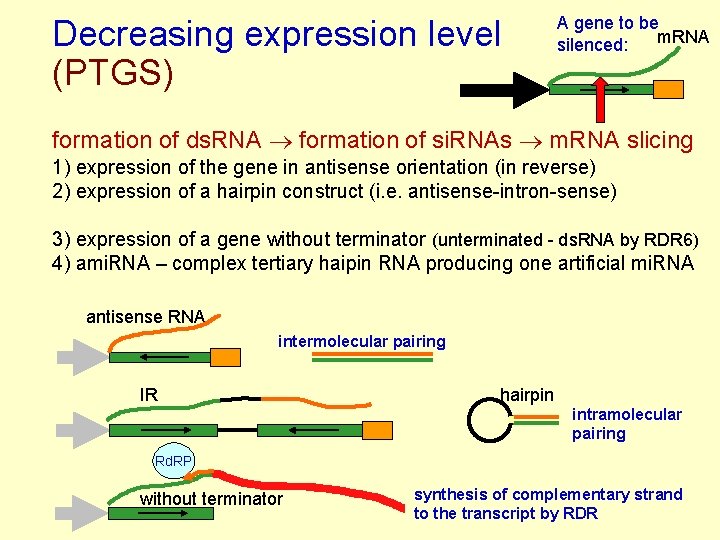
Decreasing expression level (PTGS) A gene to be m. RNA silenced: formation of ds. RNA formation of si. RNAs m. RNA slicing 1) expression of the gene in antisense orientation (in reverse) 2) expression of a hairpin construct (i. e. antisense-intron-sense) 3) expression of a gene without terminator (unterminated - ds. RNA by RDR 6) 4) ami. RNA – complex tertiary haipin RNA producing one artificial mi. RNA antisense RNA intermolecular pairing IR hairpin intramolecular pairing Rd. RP without terminator synthesis of complementary strand to the transcript by RDR

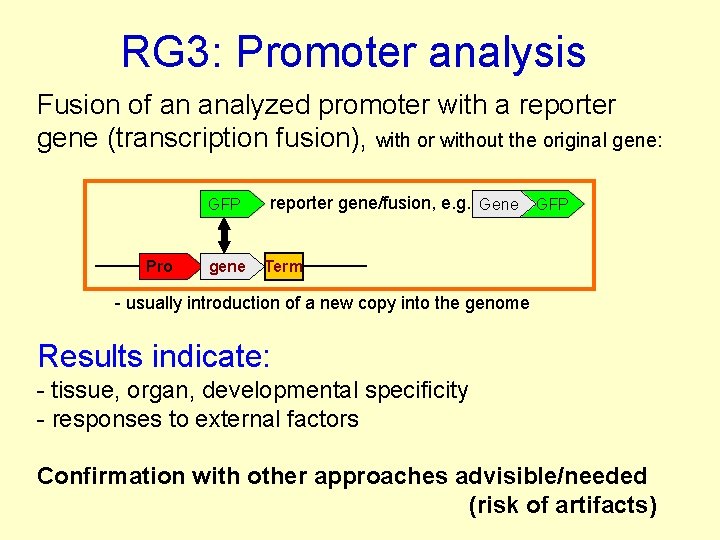
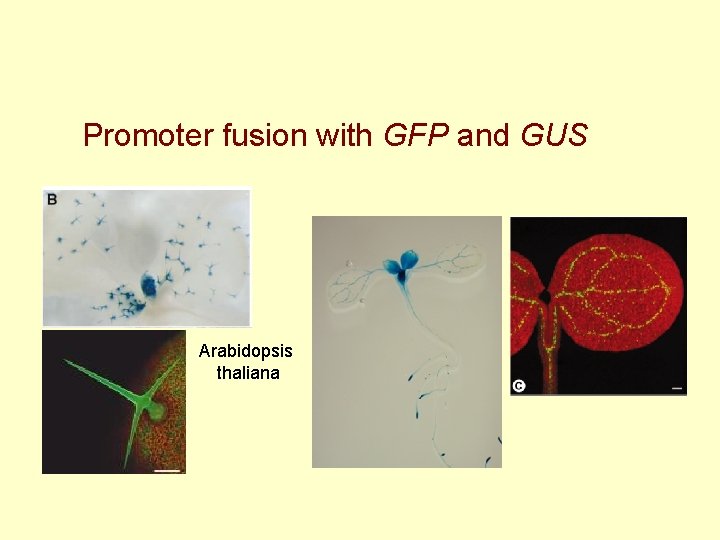
RG 3: Promoter analysis Fusion of an analyzed promoter with a reporter gene (transcription fusion), with or without the original gene: GFP Pro gene reporter gene/fusion, e. g. Gene GFP Term - usually introduction of a new copy into the genome Results indicate: - tissue, organ, developmental specificity - responses to external factors Confirmation with other approaches advisible/needed (risk of artifacts)

Promoter fusion with GFP and GUS Arabidopsis thaliana

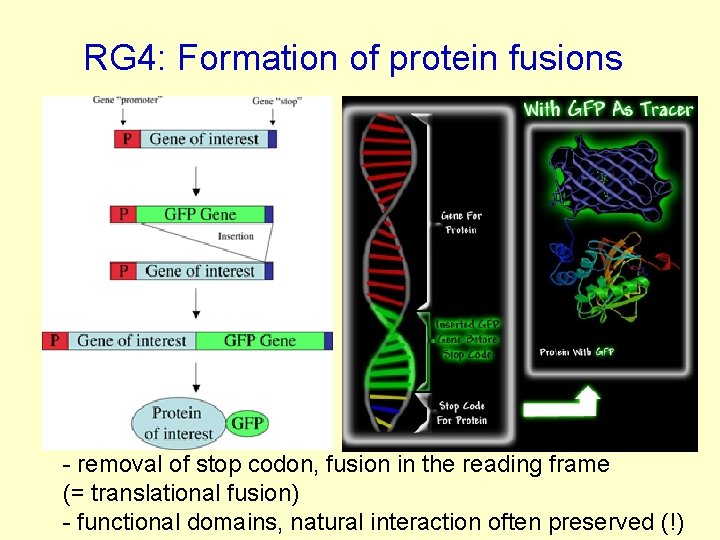
RG 4: Formation of protein fusions - removal of stop codon, fusion in the reading frame (= translational fusion) - functional domains, natural interaction often preserved (!)

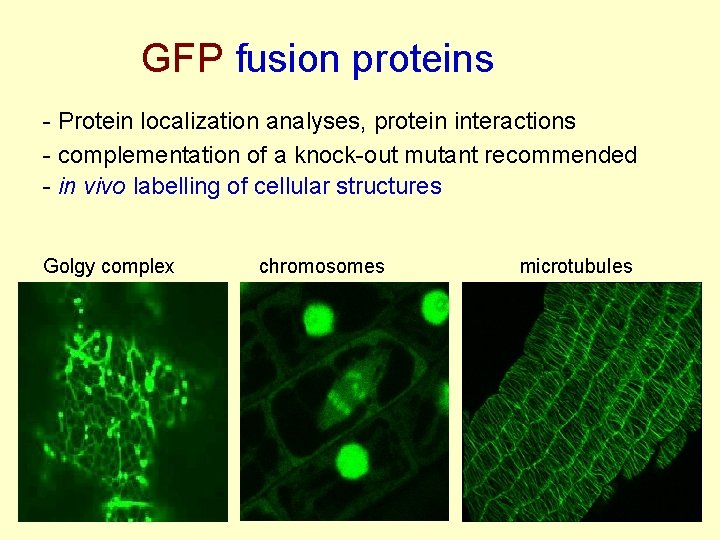
GFP fusion proteins - Protein localization analyses, protein interactions - complementation of a knock-out mutant recommended - in vivo labelling of cellular structures Golgy complex chromosomes microtubules

RG 5: Modification of the gene (outside the genome) - introduction of a modified gene copy to the knock-out mutant • gene isolation (including the native promoter) • directed mutagenesis ex vivo by PCR (primers) • selection of homozygous knock-out mutant in the gene (heterozygous mutant in the case of lethal mutations) • introduction of the modified gene (analysis of numerous lines to avoid „position-effect“ artifacts) RG 6: Gene editing in the genome (see earlier) (ZFN, TALEN, CRISPR/Cas 9 + oligonucleotides)

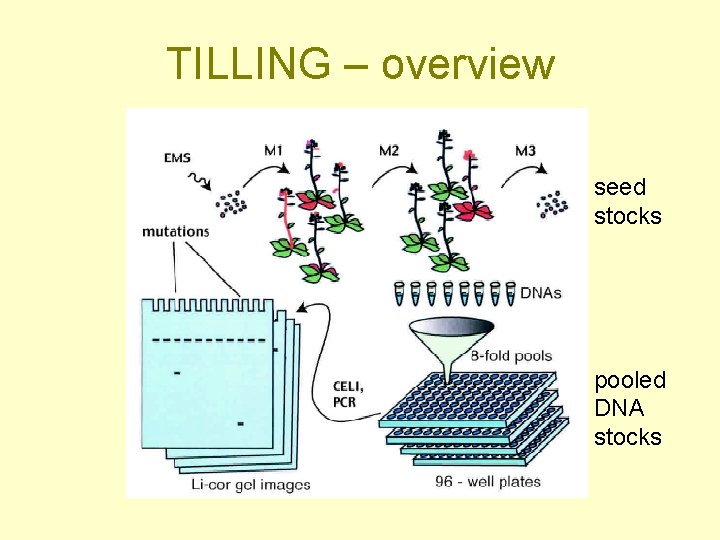
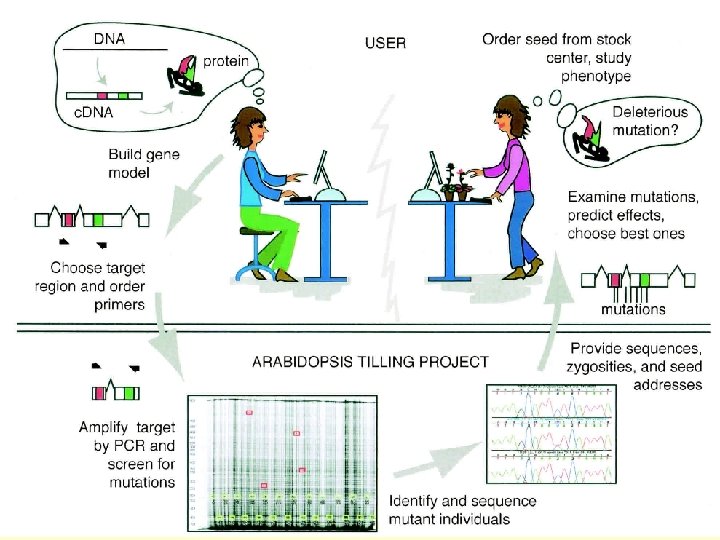
RG 7: TILLING isolation of mutants with point mutations in selected gene Targeting induced local lesions in genomes • chemical mutagenesis (EMS) – seed and DNA stocks • PCR- and heteroduplex analysis-based screen for mutants in certain gene • Point mutations! (changed regulation, interactions, …)

TILLING – overview seed stocks pooled DNA stocks

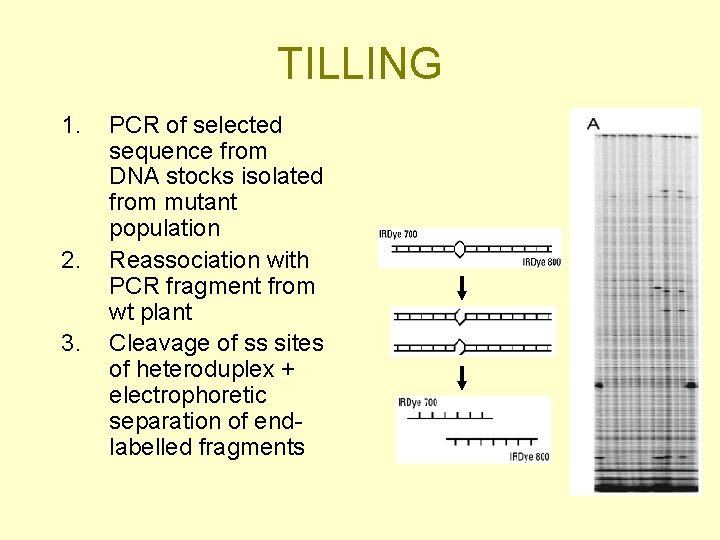
TILLING 1. 2. 3. PCR of selected sequence from DNA stocks isolated from mutant population Reassociation with PCR fragment from wt plant Cleavage of ss sites of heteroduplex + electrophoretic separation of endlabelled fragments


Direct genetics - selection of mutants by altered phenotype shootmeristemless agamous

Mutant screens – phenotype, conditions, treatments, … - looking for outstanding individuals

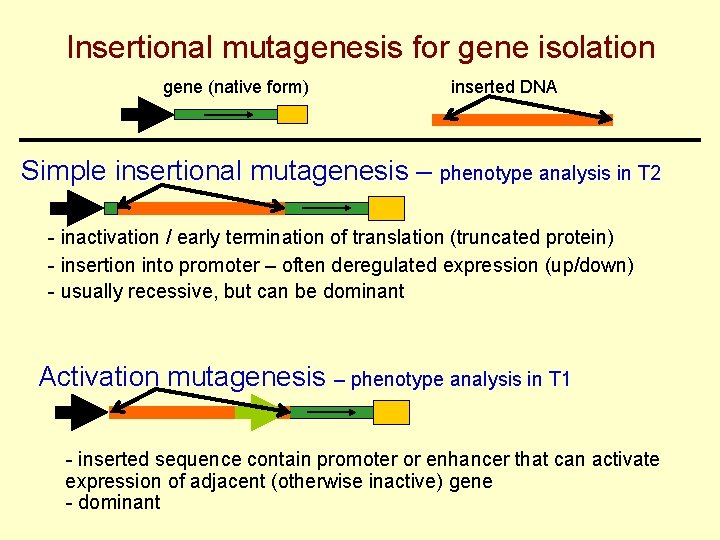
Insertional mutagenesis for gene isolation gene (native form) inserted DNA Simple insertional mutagenesis – phenotype analysis in T 2 - inactivation / early termination of translation (truncated protein) - insertion into promoter – often deregulated expression (up/down) - usually recessive, but can be dominant Activation mutagenesis – phenotype analysis in T 1 - inserted sequence contain promoter or enhancer that can activate expression of adjacent (otherwise inactive) gene - dominant

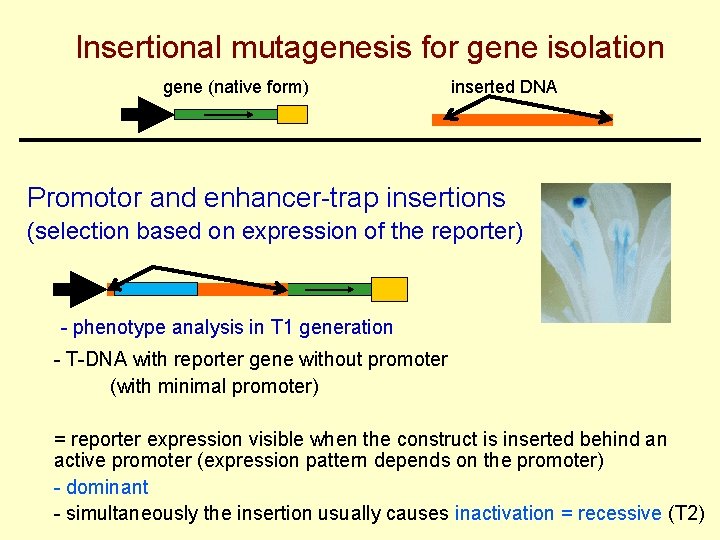
Insertional mutagenesis for gene isolation gene (native form) inserted DNA Promotor and enhancer-trap insertions (selection based on expression of the reporter) - phenotype analysis in T 1 generation - T-DNA with reporter gene without promoter (with minimal promoter) = reporter expression visible when the construct is inserted behind an active promoter (expression pattern depends on the promoter) - dominant - simultaneously the insertion usually causes inactivation = recessive (T 2)

Direct genetics – identification of mutated gene - the same phenotypic change can result from different mutations „there are numerous ways how to build up a house incorrectly“ - allelic mutations – mutation in the same gene (x different g. ) How to find out if two homozygous mutants with the same phenotype are mutated in the same gene or not (in case of recesive mutations)? Crossing of homozygous mutants F 1 - wt phenotype = different genes (complementation, heterozygous in both genes) - mutant phenotype = allelic (same genes were mutated) = no complementation

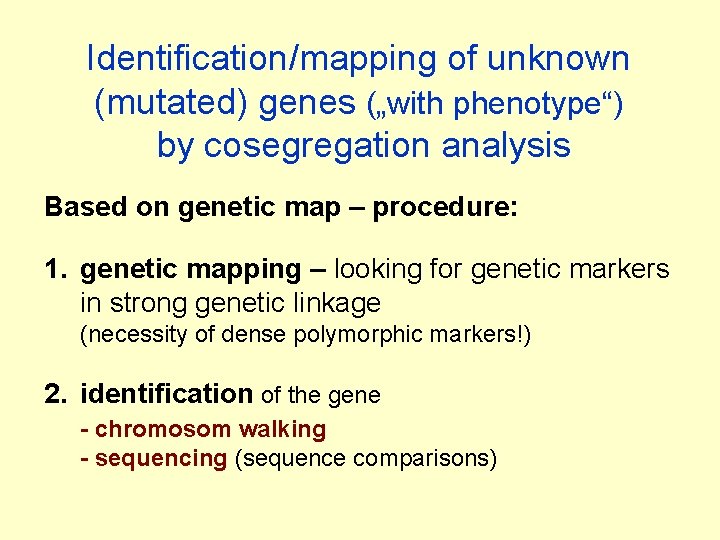
Identification/mapping of unknown (mutated) genes („with phenotype“) by cosegregation analysis Based on genetic map – procedure: 1. genetic mapping – looking for genetic markers in strong genetic linkage (necessity of dense polymorphic markers!) 2. identification of the gene - chromosom walking - sequencing (sequence comparisons)

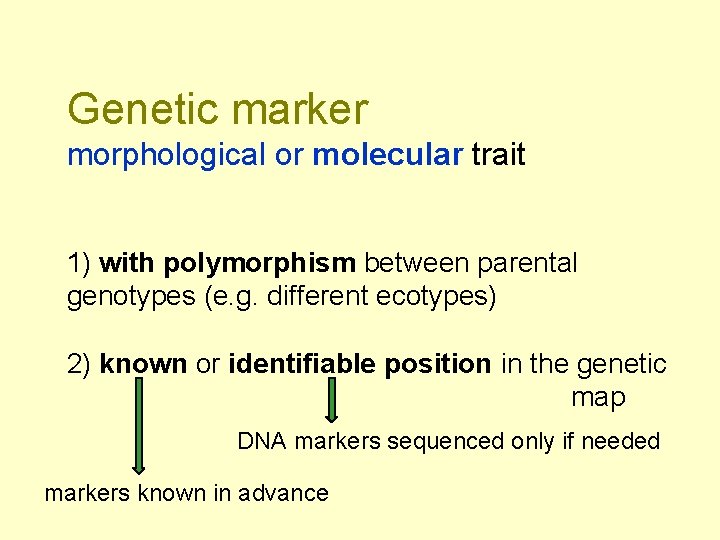
Genetic marker morphological or molecular trait 1) with polymorphism between parental genotypes (e. g. different ecotypes) 2) known or identifiable position in the genetic map DNA markers sequenced only if needed markers known in advance

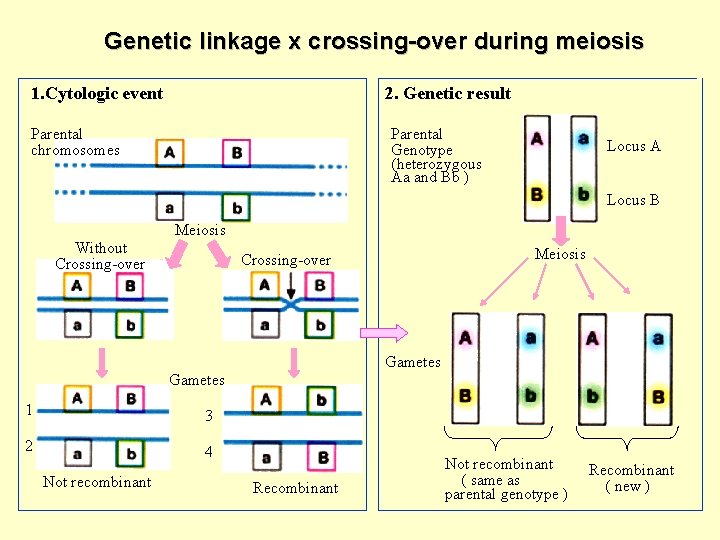
Genetic linkage x crossing-over during meiosis 1. Cytologic event 2. Genetic result Parental chromosomes Parental Genotype (heterozygous Aa and Bb ) Locus A Locus B Meiosis Without Crossing-over Meiosis Crossing-over Gametes 1 3 2 4 Not recombinant Recombinant Not recombinant ( same as parental genotype ) Recombinant ( new )

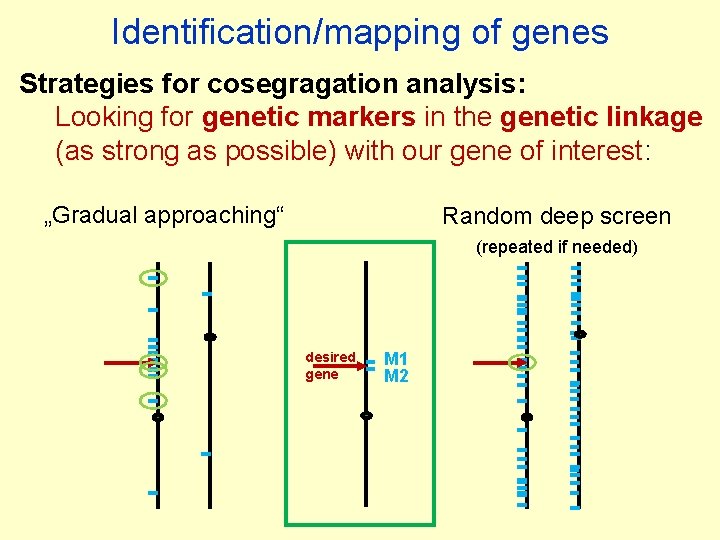
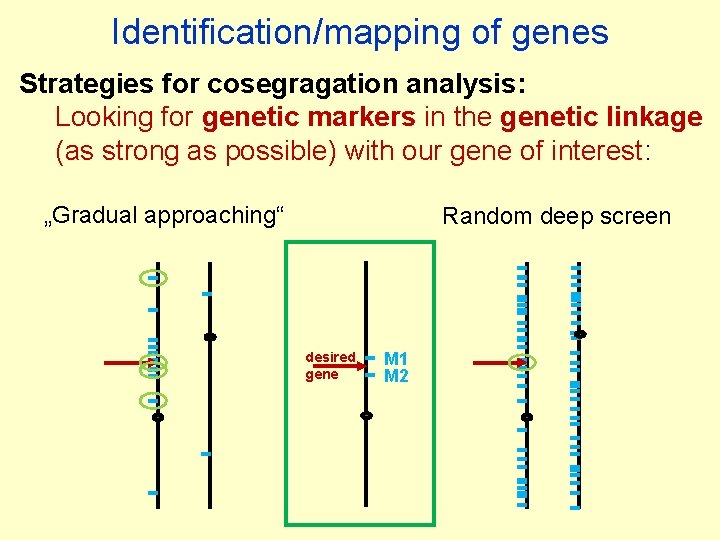
Identification/mapping of genes Strategies for cosegragation analysis: Looking for genetic markers in the genetic linkage (as strong as possible) with our gene of interest: „Gradual approaching“ Random deep screen (repeated if needed) desired gene M 1 M 2

Types of genetic markers = trait with known or identifiable position in genetic map with polymorphism between parental genotypes (e. g. different ecotypes) • Morphological (limited number) • Molecular

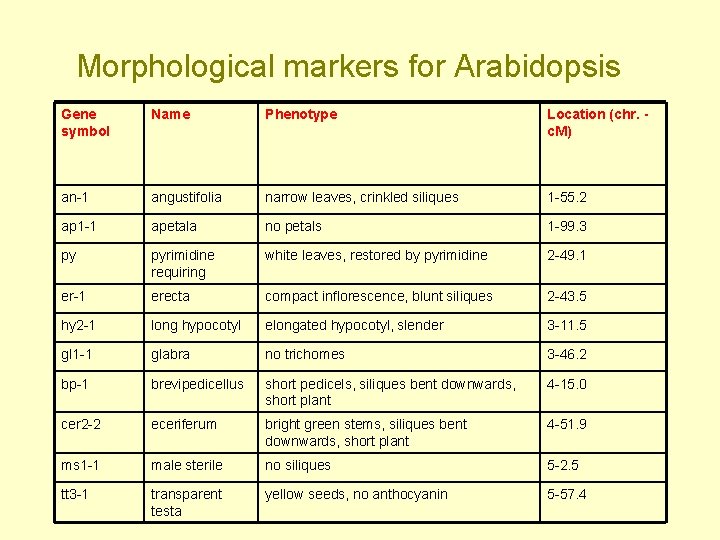
Morphological markers for Arabidopsis Gene symbol Name Phenotype Location (chr. c. M) an-1 angustifolia narrow leaves, crinkled siliques 1 -55. 2 ap 1 -1 apetala no petals 1 -99. 3 py pyrimidine requiring white leaves, restored by pyrimidine 2 -49. 1 er-1 erecta compact inflorescence, blunt siliques 2 -43. 5 hy 2 -1 long hypocotyl elongated hypocotyl, slender 3 -11. 5 gl 1 -1 glabra no trichomes 3 -46. 2 bp-1 brevipedicellus short pedicels, siliques bent downwards, short plant 4 -15. 0 cer 2 -2 eceriferum bright green stems, siliques bent downwards, short plant 4 -51. 9 ms 1 -1 male sterile no siliques 5 -2. 5 tt 3 -1 transparent testa yellow seeds, no anthocyanin 5 -57. 4

Types of genetic markers • Morphological (limited number) • Molecular – Isozymes (the same activity, but different mobility in gel electrophoresis) – DNA markers – differences in DNA sequence known OR detectable position in the genome!

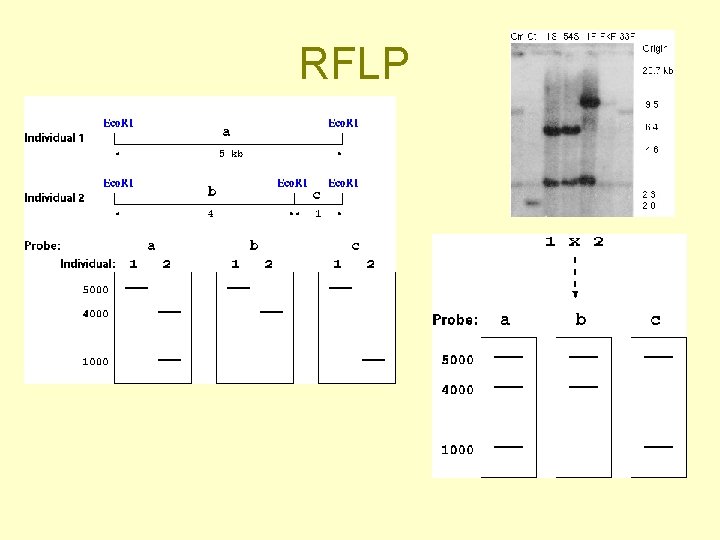
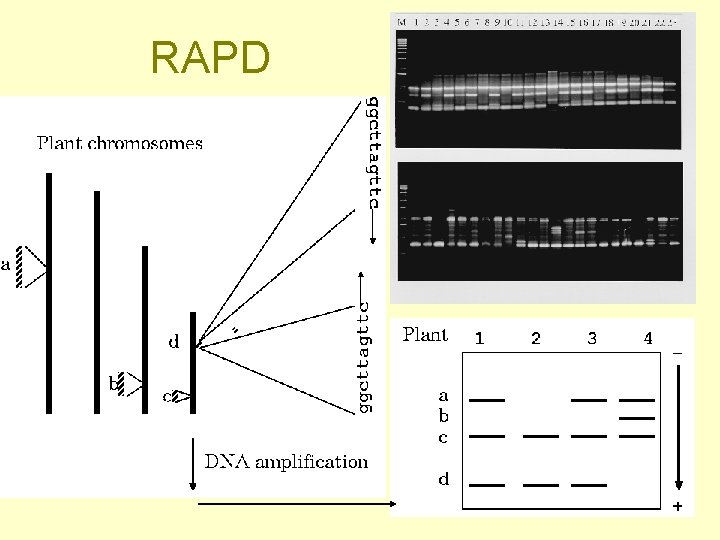
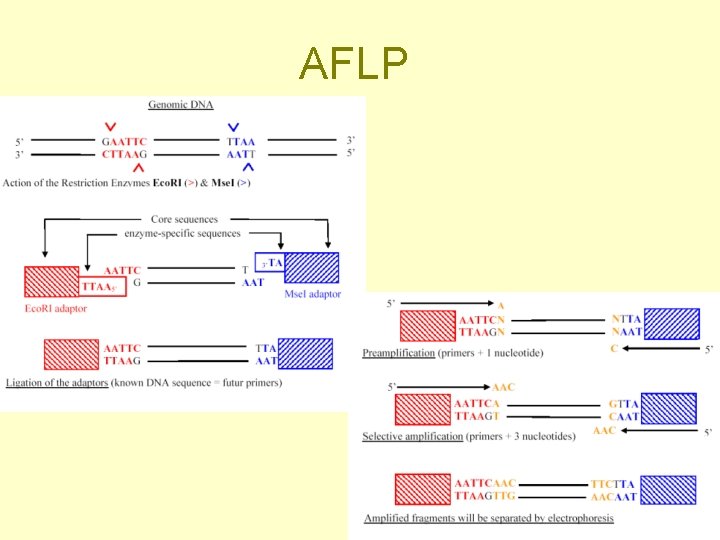
DNA molecular markers (= usually an electrophoretic band) • RFLP (Restriction fragment length polymorfism) + Southern • RAPD (Random amplified polymorphism detection) • AFLP (Amplified fragment length polymorphism)

Examples of DNA molecular markers Known position in the genome • RFLP (Restriction fragment length polymorfism) + Southern hybridization • SSR (Simple sequence repeats) Unknown sequence and position - randomly amplified DNA segments - sequence and position determined subsequently (only for those of interest - in genetic linkage with a trait) • RAPD (Random amplified polymorphism detection) • AFLP (Amplified fragment length polymorphism)

RFLP

RAPD

AFLP

Identification/mapping of genes Strategies for cosegragation analysis: Looking for genetic markers in the genetic linkage (as strong as possible) with our gene of interest: „Gradual approaching“ Random deep screen desired gene M 1 M 2

„Gradual approaching“ • Crossing of different genotypes with high polymorphism (multiple differences in markers)!!! Basic set of genetic markers in Arabidopsis thaliana 2 -3 in every chromosomal arm

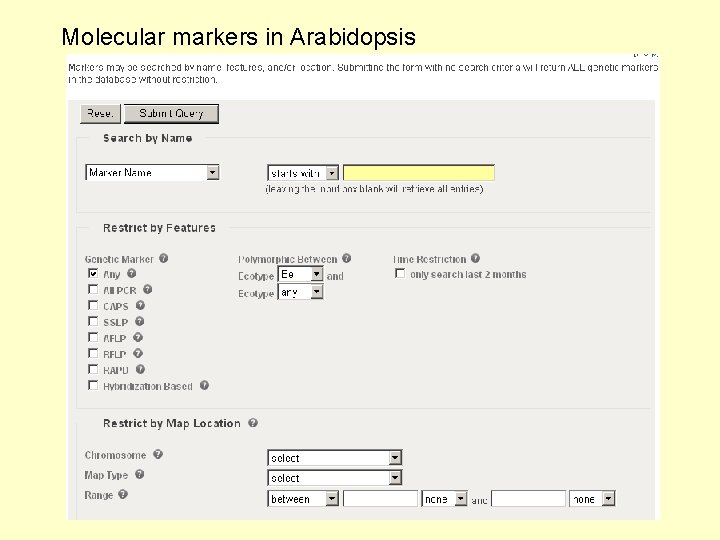
Molecular markers in Arabidopsis

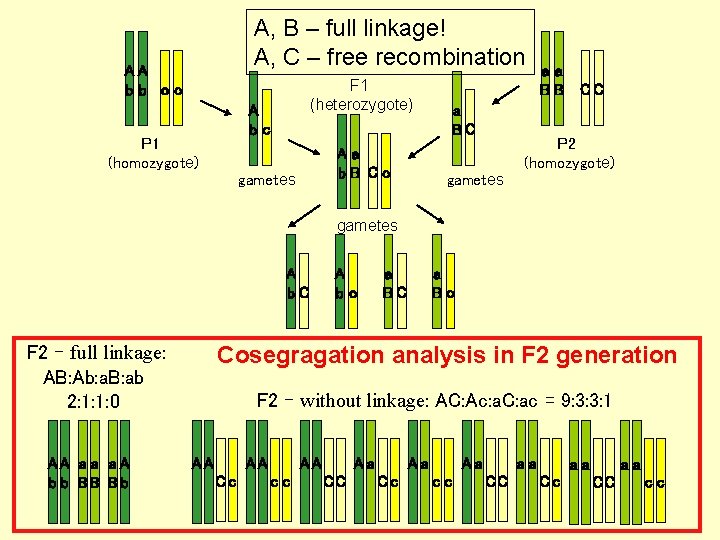
A, B – full linkage! A, C – free recombination AA bb cc F 1 (heterozygote) A bc P 1 (homozygote) Aa b. B Cc gametes a BC aa BB CC P 2 (homozygote) gametes A b. C F 2 – full linkage: AB: Ab: a. B: ab 2: 1: 1: 0 AA a a a A b b BB B b A bc a BC a Bc Cosegragation analysis in F 2 generation F 2 – without linkage: AC: Ac: a. C: ac = 9: 3: 3: 1 AA AA Cc AA cc Aa CC Aa Cc Aa cc aa CC aa Cc aa CC cc

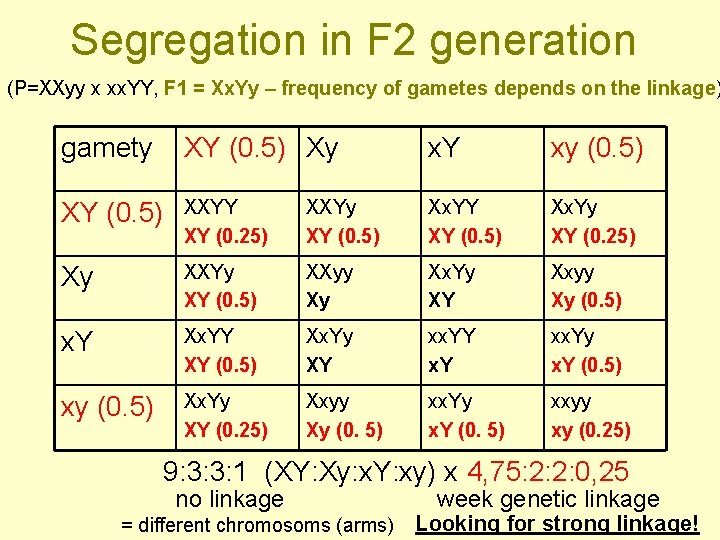
Segregation in F 2 generation (P=XXyy x xx. YY, F 1 = Xx. Yy – frequency of gametes depends on the linkage) gamety XY (0. 5) Xy x. Y xy (0. 5) XY (0. 5) XXYY XY (0. 25) XXYy XY (0. 5) Xx. YY XY (0. 5) Xx. Yy XY (0. 25) Xy XXYy XY (0. 5) XXyy Xy Xx. Yy XY Xxyy Xy (0. 5) x. Y Xx. YY XY (0. 5) Xx. Yy XY xx. YY x. Y xx. Yy x. Y (0. 5) xy (0. 5) Xx. Yy XY (0. 25) Xxyy Xy (0. 5) xx. Yy x. Y (0. 5) xxyy xy (0. 25) 9: 3: 3: 1 (XY: Xy: x. Y: xy) x 4, 75: 2: 2: 0, 25 no linkage = different chromosoms (arms) week genetic linkage Looking for strong linkage!

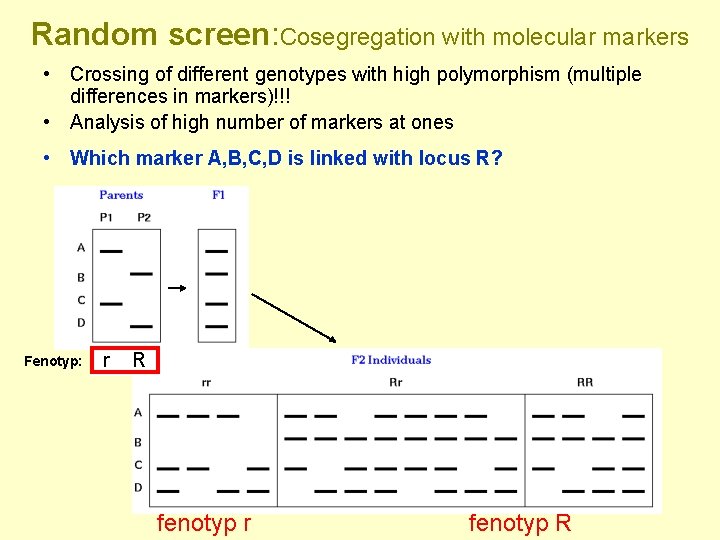
Random screen: Cosegregation with molecular markers • Crossing of different genotypes with high polymorphism (multiple differences in markers)!!! • Analysis of high number of markers at ones • Which marker A, B, C, D is linked with locus R? Fenotyp: r R fenotyp r fenotyp R

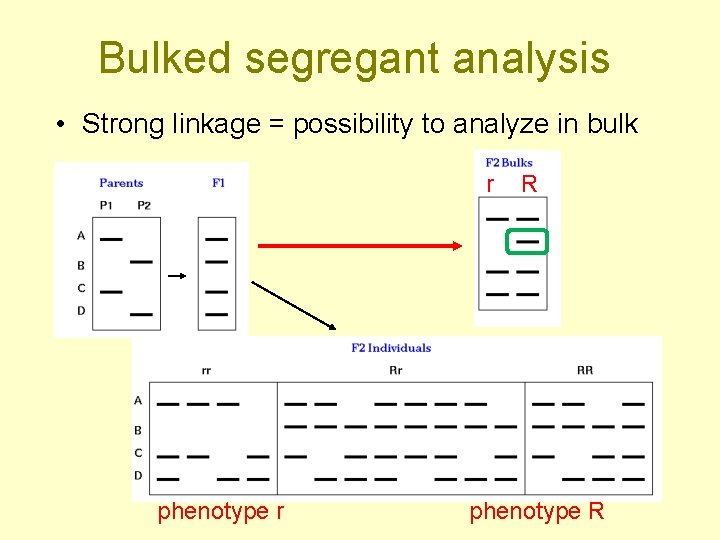
Bulked segregant analysis • Strong linkage = possibility to analyze in bulk r phenotype r R phenotype R

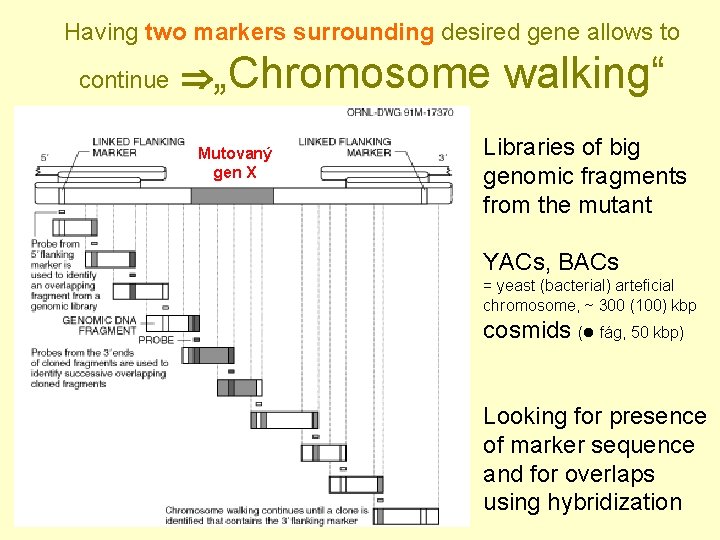
Having two markers surrounding desired gene allows to continue „Chromosome Mutovaný gen X walking“ Libraries of big genomic fragments from the mutant YACs, BACs = yeast (bacterial) arteficial chromosome, ~ 300 (100) kbp cosmids ( fág, 50 kbp) Looking for presence of marker sequence and for overlaps using hybridization

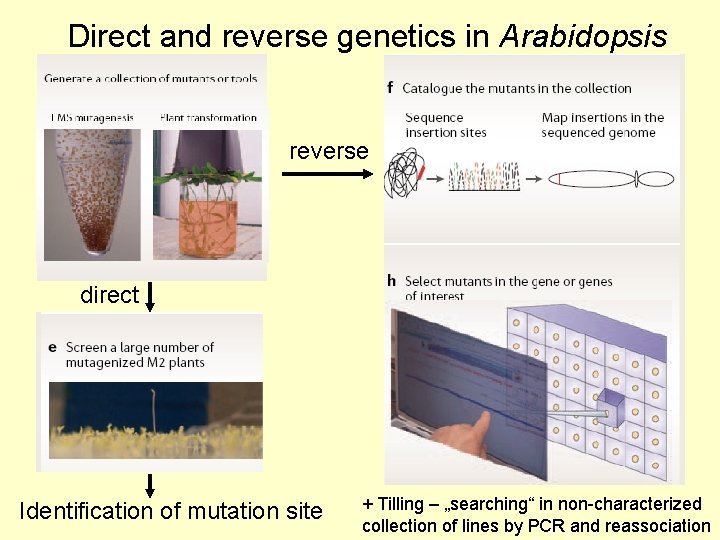
Direct and reverse genetics in Arabidopsis reverse direct Identification of mutation site + Tilling – „searching“ in non-characterized collection of lines by PCR and reassociation

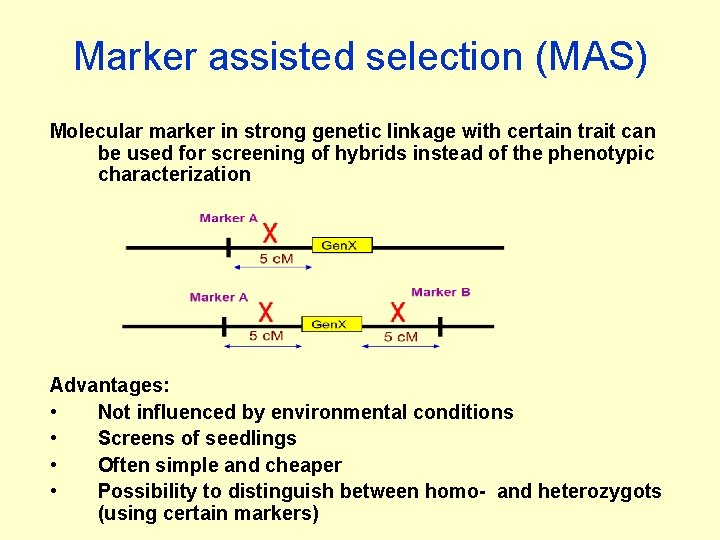
Marker assisted selection (MAS) Molecular marker in strong genetic linkage with certain trait can be used for screening of hybrids instead of the phenotypic characterization Advantages: • Not influenced by environmental conditions • Screens of seedlings • Often simple and cheaper • Possibility to distinguish between homo- and heterozygots (using certain markers)

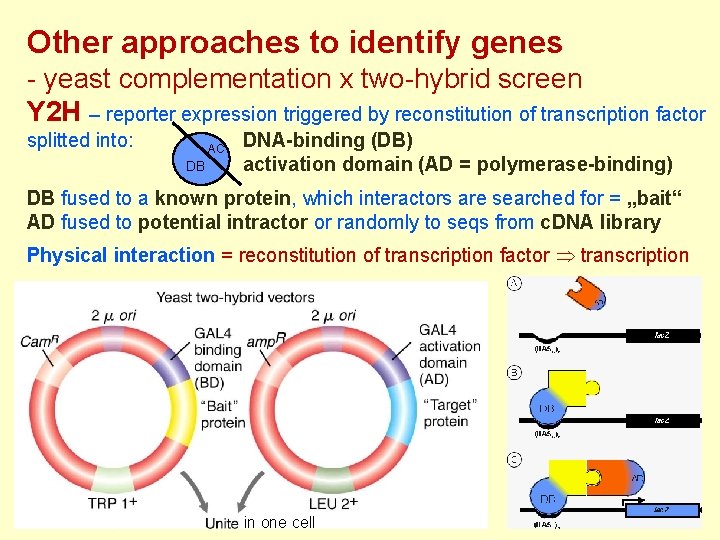
Other approaches to identify genes - yeast complementation x two-hybrid screen Y 2 H – reporter expression triggered by reconstitution of transcription factor splitted into: AC DB DNA-binding (DB) activation domain (AD = polymerase-binding) DB fused to a known protein, which interactors are searched for = „bait“ AD fused to potential intractor or randomly to seqs from c. DNA library Physical interaction = reconstitution of transcription factor transcription in one cell

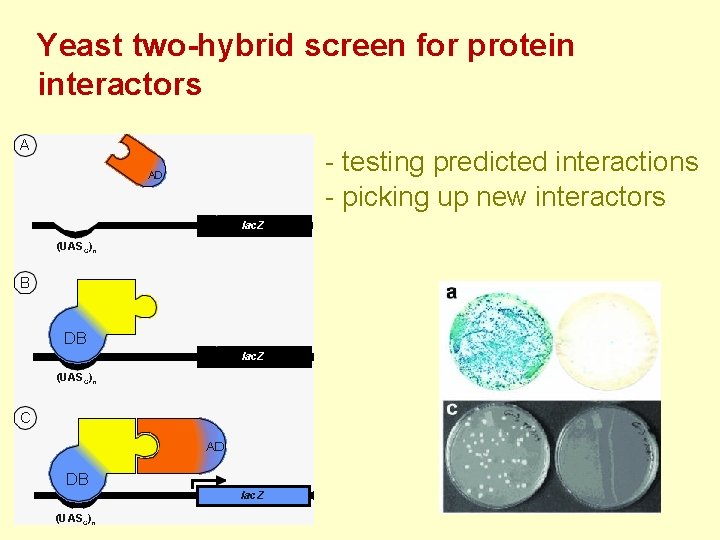
Yeast two-hybrid screen for protein interactors - testing predicted interactions - picking up new interactors