Genetics for Epidemiologists National Human Genome Research Institute


























- Slides: 58

Genetics for Epidemiologists National Human Genome Research Institute National Institutes of Health U. S. Department of Health and Human Services Lecture 7: Replication and Functional Studies U. S. Department of Health and Human Services National Institutes of Health National Human Genome Research Institute Teri A. Manolio, M. D. , Ph. D. Director, Office of Population Genomics and Senior Advisor to the Director, NHGRI, for Population Genomics

Topics to be Covered • Replication – Past challenges – Criteria – Reasons for inability to replicate findings • Finding the causal variant – Neighboring regions: conservation, nearby genes – Sequencing – Protein product – Expression studies – Experimental studies: Knockdown, knockout, knockin

Chanock S, Manolio T, et al, Nature 2007; 447: 655 -660.

Need for Consensus on What Constitutes Replication: circa November, 2006 • Avalanche of GWA and candidate gene studies anticipated in near future • Replication held as sine qua non • Likelihood of single study establishing an association is low until sample sizes increase sufficiently and analytical methods improve substantially • Common problem of how to interpret confusing and spurious findings

Case in Point: DTNBP 1 and Schizophrenia • First identified as putative schizophreniasusceptibility gene in Irish pedigrees • Reported confirmation in several replication studies in independent European samples but reported risk alleles and haplotypes appeared to differ between studies • Comparison among studies difficult because different marker sets used by each group • Hap. Map data and all identified polymorphisms typed in CEPH samples to produce high density reference map Mutsuddi et al, Am J Hum Genet 2006; 79: 903 -909.

Phylogenetic Tree of Five Common Haplotypes of DTNBP 1 Mutsuddi et al, Am J Hum Genet 2006; 79: 903 -909.

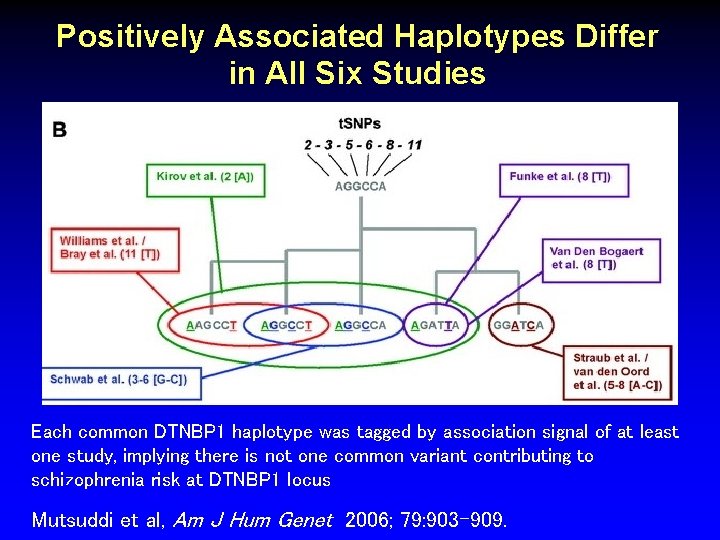
Positively Associated Haplotypes Differ in All Six Studies Each common DTNBP 1 haplotype was tagged by association signal of at least one study, implying there is not one common variant contributing to schizophrenia risk at DTNBP 1 locus Mutsuddi et al, Am J Hum Genet 2006; 79: 903 -909.

How NOT To Do A Replication Study • Use a different phenotype • Use different markers • Mix fine-mapping and replication • Use different analytic methods (haplotype vs. single marker) • Use different populations

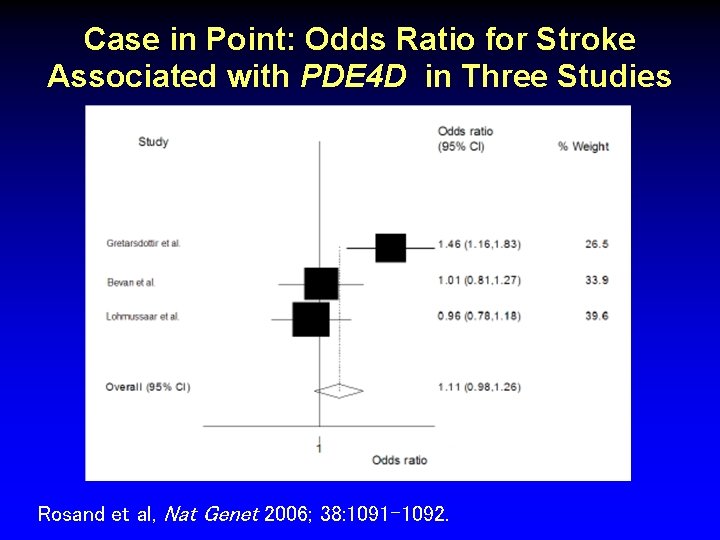
Case in Point: Odds Ratio for Stroke Associated with PDE 4 D in Three Studies Rosand et al, Nat Genet 2006; 38: 1091 -1092.

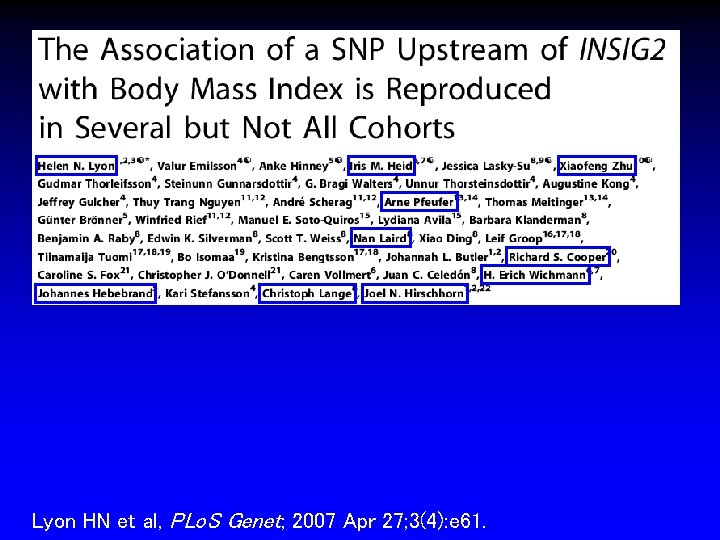
• Association between minor allele of rs 7566605 near INSIG 2 and increased BMI and homozygosity in 923 related Framingham Heart Study (FHS) participants • Association reproduced in four additional cohorts • Not seen in fifth cohort Science, 14 Apr 2006 Science, 12 Jan 2007

Lyon HN et al, PLo. S Genet; 2007 Apr 27; 3(4): e 61.

• Nine large cohorts from eight populations across multiple ethnicities • Family-based, population-based, case-control designs • Association at p < 0. 05 in five cohorts but none in three cohorts • Variability in strength of association over time • Replication both in unrelated (p = 0. 046) and familybased (p = 0. 004) samples • Suggests initial finding unlikely to be spurious but effect likely to be heterogeneous Lyon HN et al, PLo. S Genet; 2007 Apr 27; 3(4): e 61.

• Second variant (rs 1455832 -C) intronic to ROBO 1 with age-varying association to BMI over time • Minor allele homozygote associated with increased BMI diminishing after age 45 • Replicated age-varying association in same direction in five of eight other cohorts totaling 13, 584 subjects • One childhood cohort showed very strong interaction (p < 10 -9), four others 0. 003 < p < 0. 05; overall 10 -9 • In all replication cohorts but one, association would not have been detected if testing only for main genetic effect and not for age-by-’ 5832 interaction Lasky-Su J et al, Am J Hum Genet; 2008; 82: 849 -58.

Definition of Robust Initial Finding • Sufficient statistical power to observe reported effect, which will vary by magnitude of observed effect • Highly significant analysis using stable method • Consistent findings using simple, straightforward analytic approach • Consistent findings in epidemiologically sound study • Consistent findings overall and within key subgroups of initial study • Consistent findings in same or highly similar phenotypes

Value of Single/First Study • Initial study rarely definitive by itself but often represents important discovery tool – If consortium of multiple studies, stronger • What to do with studies not having option for replication? – Don’t change standards for definitiveness • Don’t just rely on GWA-- have multiple tools for identifying and understanding associations • May need different standards for findings of major clinical significance, particularly pharmacogenomic demonstration of adverse effect that would be unethical to try to replicate

Importance of Significance Level • Should we promulgate a specific number– NO, but in general, smaller is better • General agreement: range is very broad, higher threshold for difficult to measure phenotype • Beware of the very smallest • If significance depends on analytic method or multiple comparison correction, BEWARE • If significance or association depends on phenotype definition, BEWARE • Randomize the phenotypes and report number significant at that level • Biologic information may be useful A PRIORI but a posteriori can come up with almost anything

Importance of Genotyping Quality • Report results of known study sample duplicates, Hap. Map or other standard duplicates • Replicate small number of “significant” SNPs with second technology at some late stage • May not be needed if nearby SNPs in strong LD show same results • Strong caveats are needed regarding fallibility of genotyping - Results can change based on genotype calling algorithm - QC filters and consistency of results after applying them must be described

NCI-NHGRI Working Group Criteria for Positive Replication • Sufficient sample size to distinguish proposed effect from no effect convincingly • Same or very similar trait (extension to related trait may increase confidence in finding, such as consistent finding for both dichotomized obesity and continuous BMI) • Same or very similar population (extension to other populations may also increase confidence in finding, such as consistent association in populations of European, Asian, or even recent African ancestry) Chanock S, Manolio T, et al, Nature 2007; 447: 655 -660.

NCI-NHGRI Working Group Criteria for Positive Replication (continued) • Same inheritance model (dominant, codominant, recessive), though not necessarily same analytic method • Same gene, same SNP (or SNP in complete LD with prior SNP, r 2 = 1), same direction as original finding • Highly significant association • N. B. : Initial study must adequately describe these parameters Chanock S, Manolio T, et al, Nature 2007; 447: 655 -660.

Criteria for True Non-Replication or “Meaningful Negativity” • Same as for positive replication (same trait, same gene, same SNP, same direction, same genetic model) • Must be identical trait and population to claim non-replication • Powered to appropriate effect size (account for “winner’s curse”) Chanock S, Manolio T, et al, Nature 2007; 447: 655 -660.

Replication in Samples of Different Ancestral Origin • Shorter LD blocks may explain failure of associations identified in European ancestry populations to replicate in recent African ancestry samples • Shorter LD may permit better localization of risk variants • Allele frequency differences may also explain lack of replication

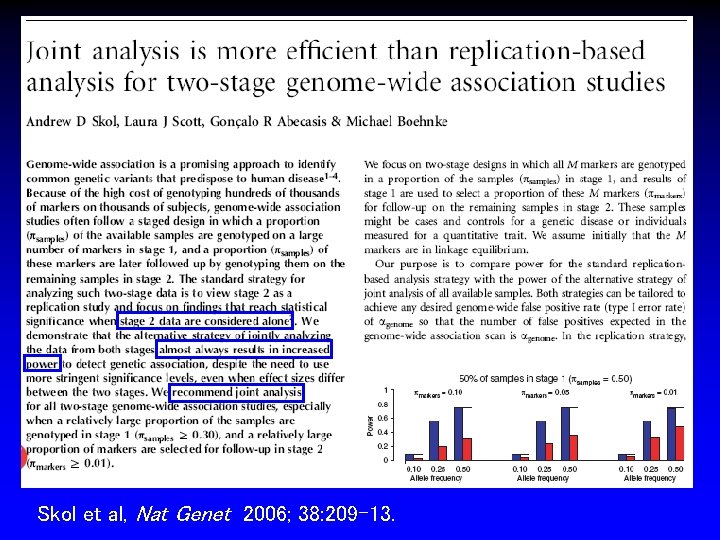
Skol et al, Nat Genet 2006; 38: 209 -13.

Narrowing the Association Region… Larson, G. The Complete Far Side. 2003.

Flow of Investigation: From Genome-Wide Association to Clinical Translation Initial Genome-Wide Association (GWA) Studies Replication/Fine Mapping Sequencing/Genotyping Functional Studies Translational Studies

Flow of Investigation: From Genome-Wide Association to Clinical Translation Initial Genome-Wide Association (GWA) Studies Replication/Fine Mapping Sequencing/Genotyping Functional Studies Translational Studies

Flow of Investigation: From Genome-Wide Association to Clinical Translation Initial Genome-Wide Association (GWA) Studies Replication/Fine Mapping Sequencing/Genotyping Functional Studies Translational Studies

Flow of Investigation: From Genome-Wide Association to Clinical Translation Initial Genome-Wide Association (GWA) Studies Replication/Fine Mapping Sequencing/Genotyping Functional Studies Translational Studies

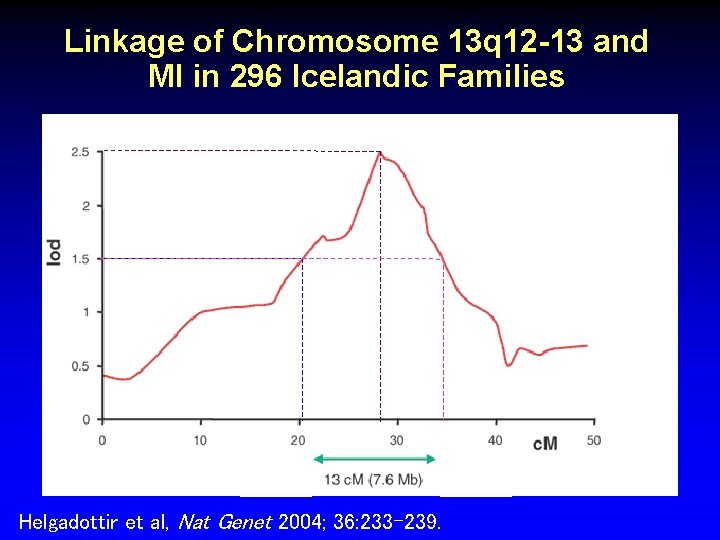
Linkage of Chromosome 13 q 12 -13 and MI in 296 Icelandic Families Helgadottir et al, Nat Genet 2004; 36: 233 -239.

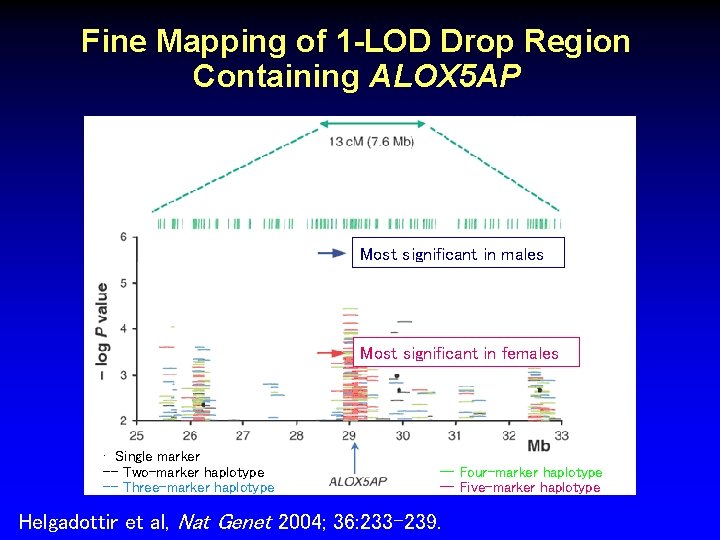
Fine Mapping of 1 -LOD Drop Region Containing ALOX 5 AP Most significant in males Most significant in females • Single marker -- Two-marker haplotype -- Three-marker haplotype -- Four-marker haplotype -- Five-marker haplotype Helgadottir et al, Nat Genet 2004; 36: 233 -239.

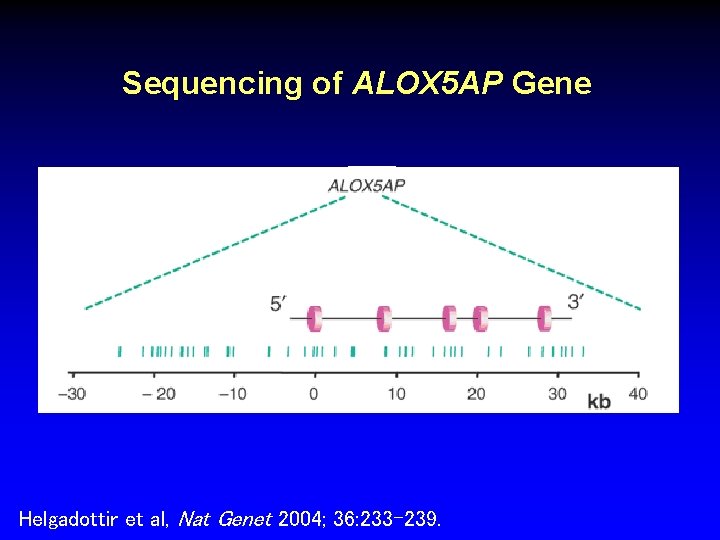
Sequencing of ALOX 5 AP Gene Helgadottir et al, Nat Genet 2004; 36: 233 -239.

Sequencing for GWA: 1, 000 Genomes Project http: //www. 1000 genomes. org/

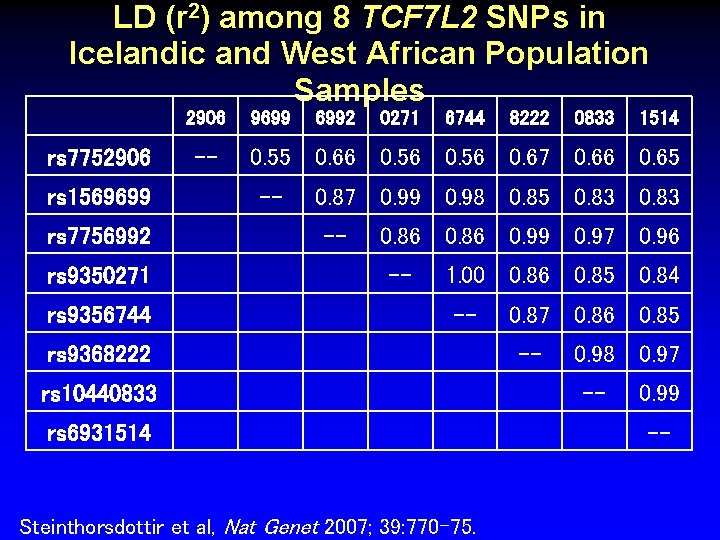
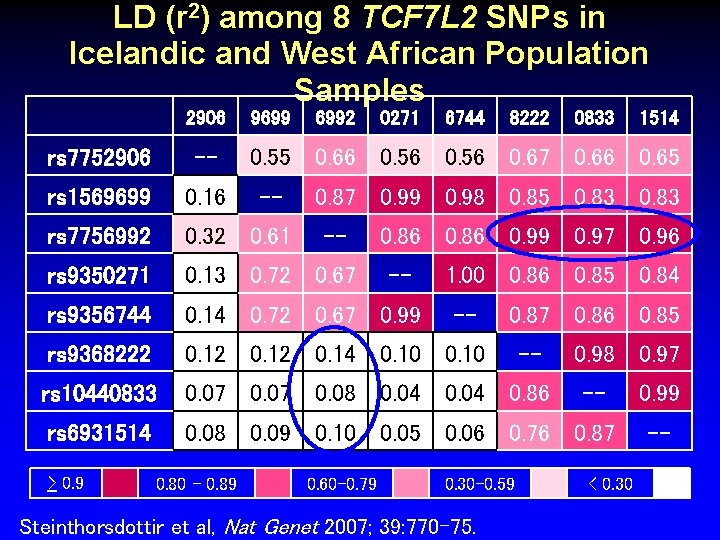
LD (r 2) among 8 TCF 7 L 2 SNPs in Icelandic and West African Population Samples 2906 rs 7752906 rs 1569699 rs 7756992 rs 9350271 rs 9356744 -- 9699 6992 0271 6744 8222 0833 1514 0. 55 0. 66 0. 56 0. 67 0. 66 0. 65 -- 0. 87 0. 99 0. 98 0. 85 0. 83 -- 0. 86 0. 99 0. 97 0. 96 -- 1. 00 0. 86 0. 85 0. 84 -- rs 9368222 rs 10440833 rs 6931514 Steinthorsdottir et al, Nat Genet 2007; 39: 770 -75. 0. 87 0. 86 0. 85 -- 0. 98 0. 97 -- 0. 99 --

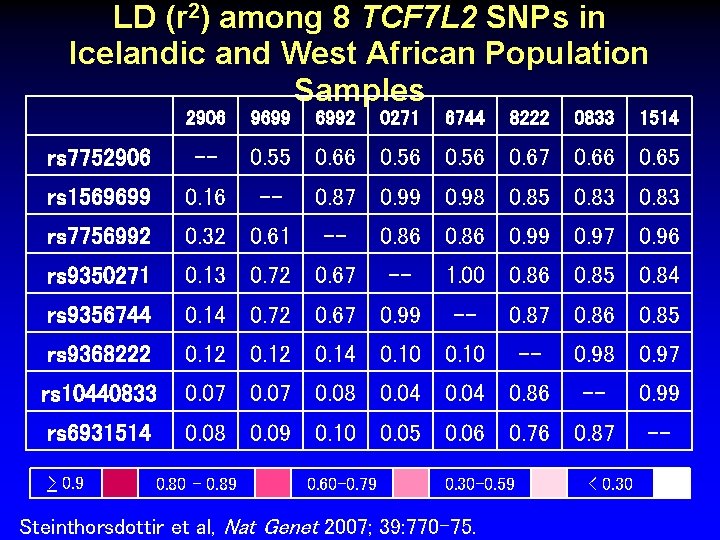
LD (r 2) among 8 TCF 7 L 2 SNPs in Icelandic and West African Population Samples 2906 9699 6992 0271 6744 8222 0833 rs 7752906 -- rs 1569699 0. 16 rs 7756992 0. 32 0. 61 rs 9350271 0. 13 0. 72 0. 67 rs 9356744 0. 14 0. 72 0. 67 0. 99 rs 9368222 0. 12 0. 14 0. 10 rs 10440833 0. 07 0. 08 0. 04 0. 86 rs 6931514 0. 08 0. 09 0. 10 0. 05 0. 06 0. 76 0. 87 > 0. 9 0. 80 – 0. 89 1514 0. 55 0. 66 0. 56 0. 67 0. 66 0. 65 -- 0. 87 0. 99 0. 98 0. 85 0. 83 -- 0. 60 -0. 79 0. 86 0. 99 0. 97 0. 96 -- 1. 00 0. 86 0. 85 0. 84 -- 0. 87 0. 86 0. 85 0. 30 -0. 59 Steinthorsdottir et al, Nat Genet 2007; 39: 770 -75. -- 0. 98 0. 97 -- < 0. 30 0. 99 --

LD (r 2) among 8 TCF 7 L 2 SNPs in Icelandic and West African Population Samples 2906 9699 6992 0271 6744 8222 0833 rs 7752906 -- rs 1569699 0. 16 rs 7756992 0. 32 0. 61 rs 9350271 0. 13 0. 72 0. 67 rs 9356744 0. 14 0. 72 0. 67 0. 99 rs 9368222 0. 12 0. 14 0. 10 rs 10440833 0. 07 0. 08 0. 04 0. 86 rs 6931514 0. 08 0. 09 0. 10 0. 05 0. 06 0. 76 0. 87 > 0. 9 0. 80 – 0. 89 1514 0. 55 0. 66 0. 56 0. 67 0. 66 0. 65 -- 0. 87 0. 99 0. 98 0. 85 0. 83 -- 0. 60 -0. 79 0. 86 0. 99 0. 97 0. 96 -- 1. 00 0. 86 0. 85 0. 84 -- 0. 87 0. 86 0. 85 0. 30 -0. 59 Steinthorsdottir et al, Nat Genet 2007; 39: 770 -75. -- 0. 98 0. 97 -- < 0. 30 0. 99 --

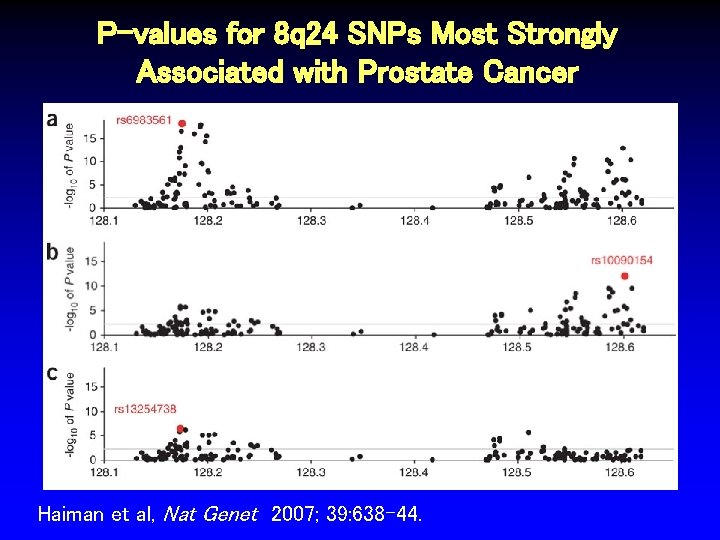
P-values for 8 q 24 SNPs Most Strongly Associated with Prostate Cancer Haiman et al, Nat Genet 2007; 39: 638 -44.

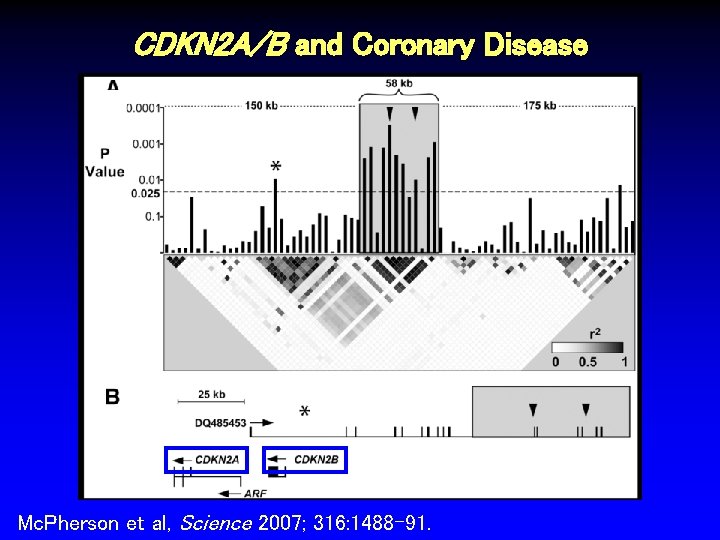
CDKN 2 A/B and Coronary Disease Mc. Pherson et al, Science 2007; 316: 1488 -91.

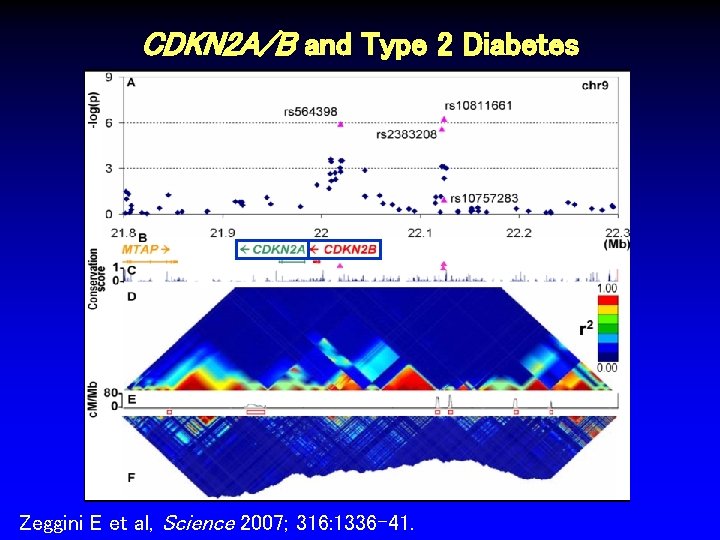
CDKN 2 A/B and Type 2 Diabetes Zeggini E et al, Science 2007; 316: 1336 -41.

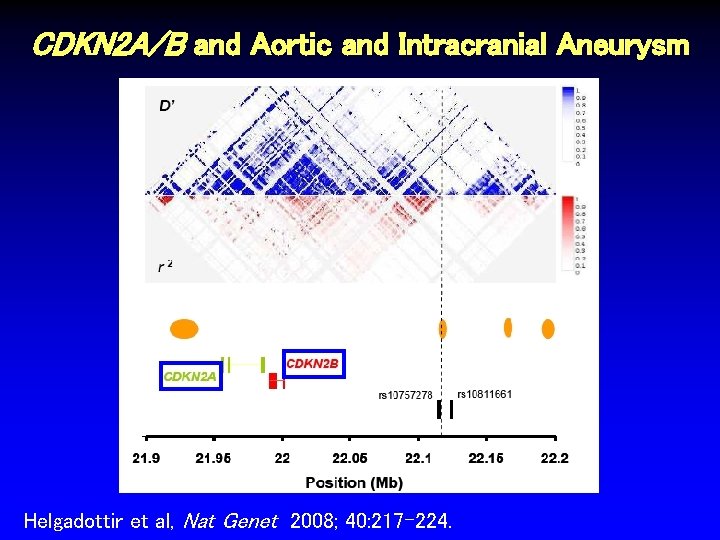
CDKN 2 A/B and Aortic and Intracranial Aneurysm Helgadottir et al, Nat Genet 2008; 40: 217 -224.

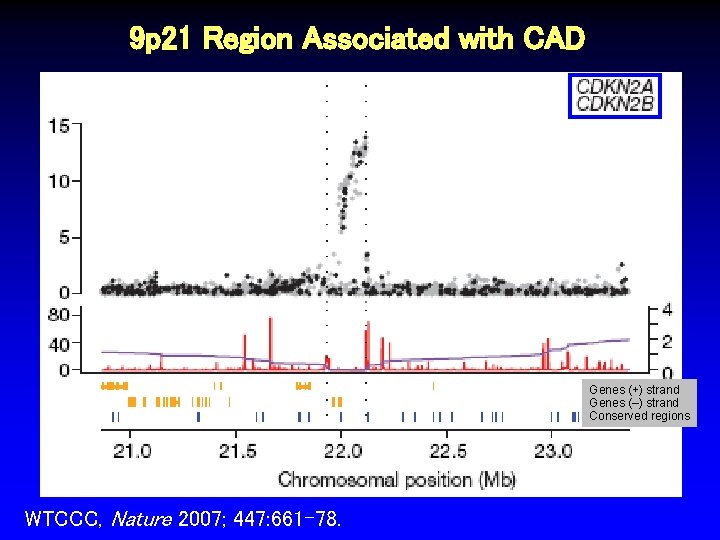
9 p 21 Region Associated with CAD Genes (+) strand Genes (–) strand Conserved regions WTCCC, Nature 2007; 447: 661 -78.

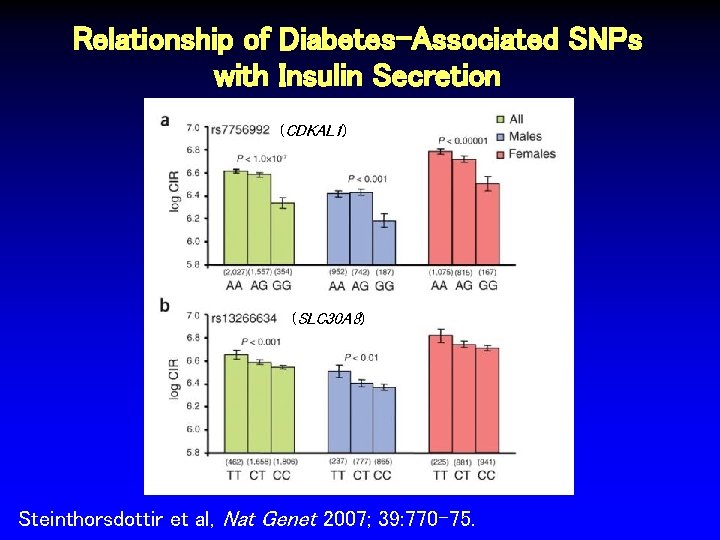
Functional Studies: Correlation of SNPs with Logical Intermediate Phenotypes • rs 7756992 on 6 p 22. 3 strongly associated with type 2 diabetes (OR 1. 20, p < 8 x 10 -8), resides in intron 5 of CDK 5 regulatory subunit associated protein 1 -like 1 (CDKAL 1) • rs 13244434 on 8 q 24 also associated with T 2 DM: OR 1. 15, p < 4 x 10 -6 • Nonsynonymous arginine to tryptophan change in last exon of solute carrier family 30 (zinc transporter), member 8 (SLC 30 A 8) • Specific to pancreas and expressed in beta cells Steinthorsdottir et al, Nat Genet 2007; 39: 770 -75.

Relationship of Diabetes-Associated SNPs with Insulin Secretion (CDKAL 1) (SLC 30 A 8) Steinthorsdottir et al, Nat Genet 2007; 39: 770 -75.

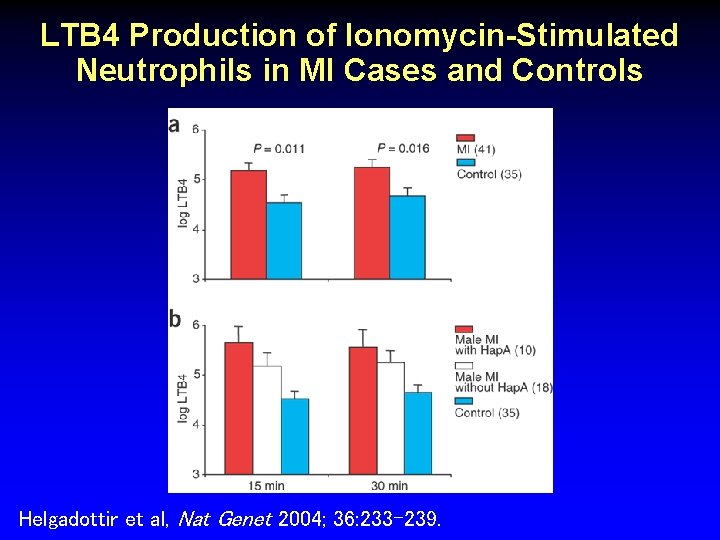
LTB 4 Production of Ionomycin-Stimulated Neutrophils in MI Cases and Controls Helgadottir et al, Nat Genet 2004; 36: 233 -239.

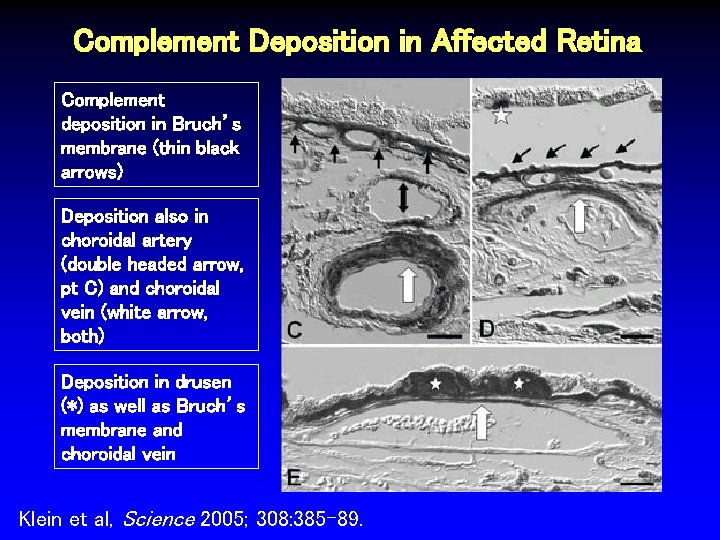
Co-Localization of Gene Product with Histopathologic Changes • CFH in retina and drusen (macular degeneration) • GAB 2 in dystrophic neurons (Alzheimers disease)

Complement Deposition in Affected Retina Complement deposition in Bruch’s membrane (thin black arrows) Deposition also in choroidal artery (double headed arrow, pt C) and choroidal vein (white arrow, both) Deposition in drusen (*) as well as Bruch’s membrane and choroidal vein Klein et al, Science 2005; 308: 385 -89.

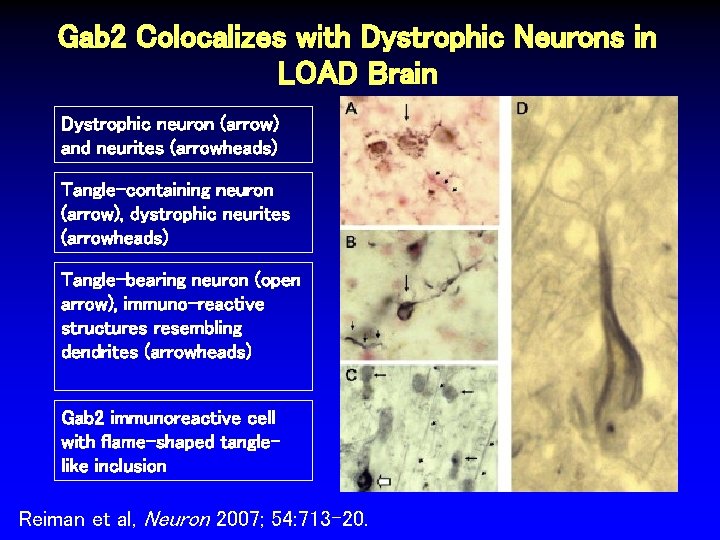
Gab 2 Colocalizes with Dystrophic Neurons in LOAD Brain Dystrophic neuron (arrow) and neurites (arrowheads) Tangle-containing neuron (arrow), dystrophic neurites (arrowheads) Tangle-bearing neuron (open arrow), immuno-reactive structures resembling dendrites (arrowheads) Gab 2 immunoreactive cell with flame-shaped tanglelike inclusion Reiman et al, Neuron 2007; 54: 713 -20.

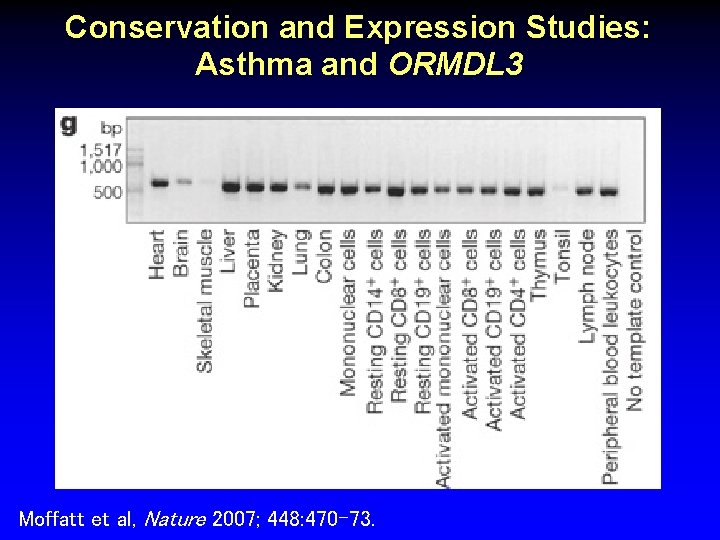
Conservation and Expression Studies: Asthma and ORMDL 3 Moffatt et al, Nature 2007; 448: 470 -73.

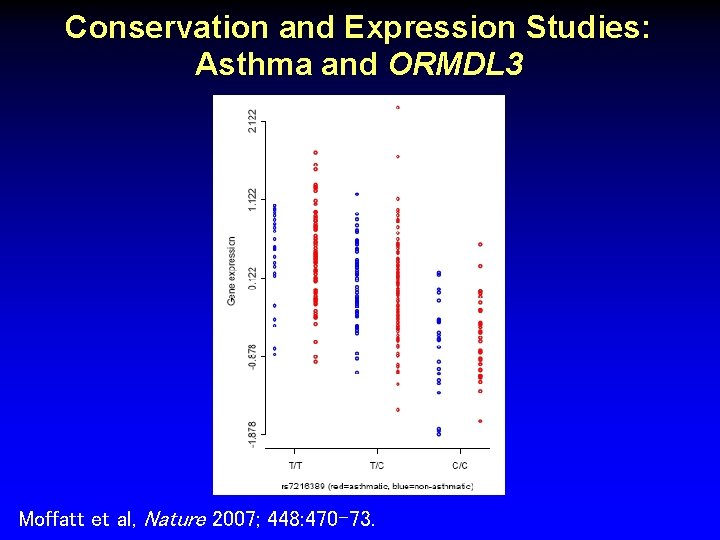
Conservation and Expression Studies: Asthma and ORMDL 3 Moffatt et al, Nature 2007; 448: 470 -73.

Knockdown and Knockout Studies • Knockdown of ATG 16 L 1 – Associated with Crohn’s disease – Reduces phagocytosis of S. typhimurium in He. La cells • Knockdown of GAB 2 – Associated with Azheimer’s disease – Increases tau phosphorylation • Knockout of MLXIPL – Associated with lower triglyceride levels – Knockout shows lower triglyceride levels – Transgenic (knockin) shows higher levels

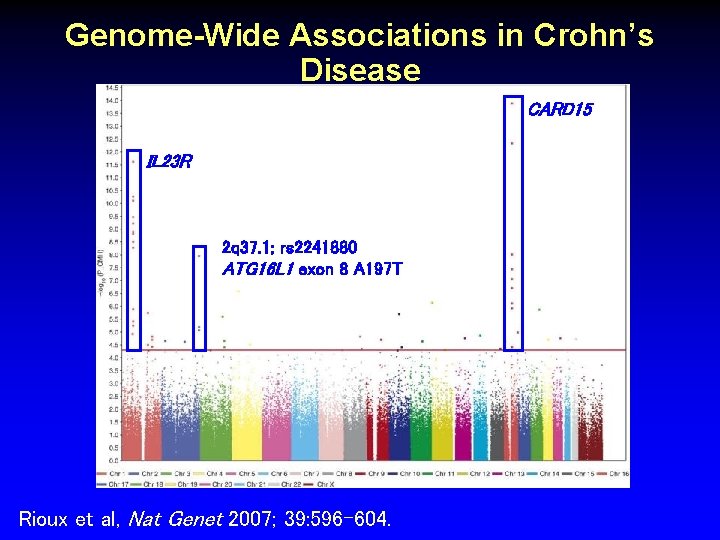
Genome-Wide Associations in Crohn’s Disease CARD 15 IL 23 R 2 q 37. 1; rs 2241880 ATG 16 L 1 exon 8 A 197 T Rioux et al, Nat Genet 2007; 39: 596 -604.

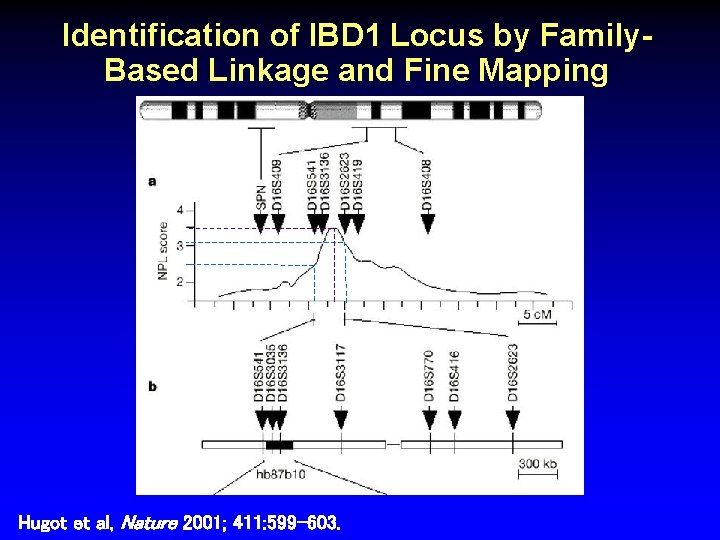
Identification of IBD 1 Locus by Family. Based Linkage and Fine Mapping Hugot et al, Nature 2001; 411: 599 -603.

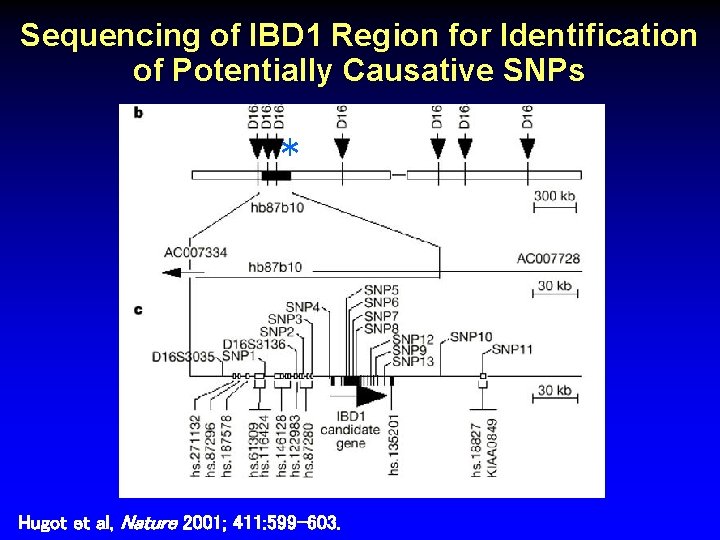
Sequencing of IBD 1 Region for Identification of Potentially Causative SNPs * Hugot et al, Nature 2001; 411: 599 -603.

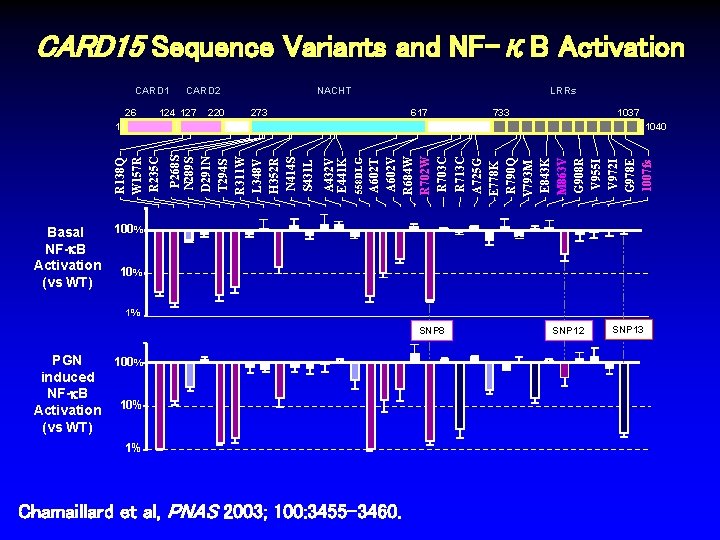
CARD 15 Sequence Variants and NF-κB Activation CARD 1 26 NACHT CARD 2 124 127 220 LRRs 273 617 1037 733 A 602 T A 602 V R 684 W R 702 W R 703 C R 713 C A 725 G E 778 K R 790 Q V 793 M E 843 K M 863 V G 908 R V 955 I V 972 I G 978 E 1007 fs 558 DLG A 432 V E 441 K P 268 S N 289 S D 291 N T 294 S R 311 W L 348 V H 352 R N 414 S S 431 L 1040 R 138 Q W 157 R R 235 C 1 100% Basal NF-k. B Activation 10% (vs WT) 1% SNP 8 PGN 100% induced NF-k. B Activation 10% (vs WT) 1% Chamaillard et al, PNAS 2003; 100: 3455 -3460. SNP 12 SNP 13

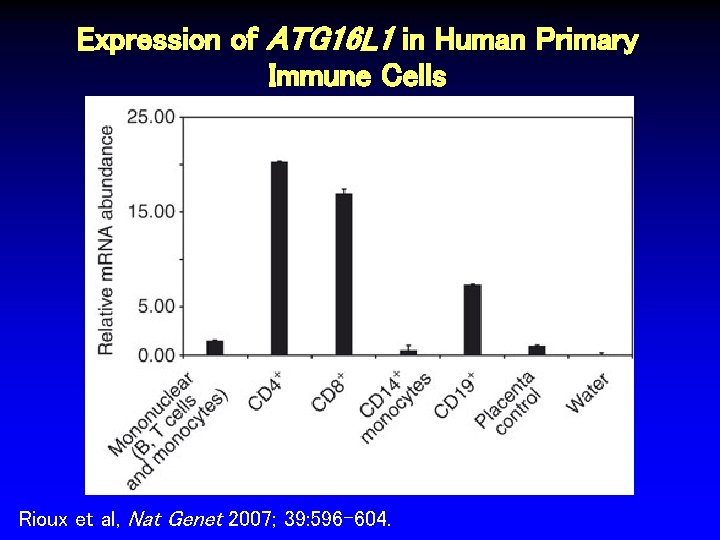
Gene Expression in Crohn’s Disease • rs 2241880 associated at p < 10 -8 • Nonsynonymous amino acid change in exon 8 of autophagy-related 16 -like 1 (ATG 16 L 1) • Autophagy is biologic process involved in protein degradation, antigen processing, absorption of cellular organelles, initiation and regulation of inflammatory response Rioux et al, Nat Genet 2007; 39: 596 -604.

Expression of ATG 16 L 1 in Human Primary Immune Cells Rioux et al, Nat Genet 2007; 39: 596 -604.

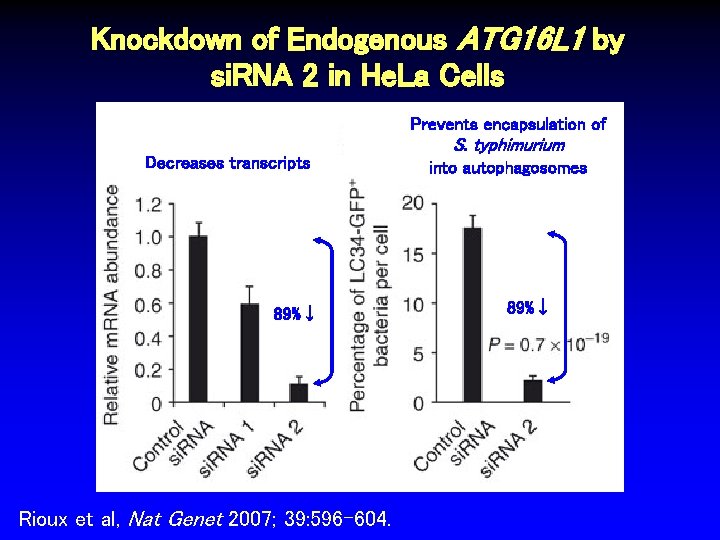
Knockdown of Endogenous ATG 16 L 1 by si. RNA 2 in He. La Cells Prevents encapsulation of Decreases transcripts 89%↓ Rioux et al, Nat Genet 2007; 39: 596 -604. S. typhimurium into autophagosomes 89%↓

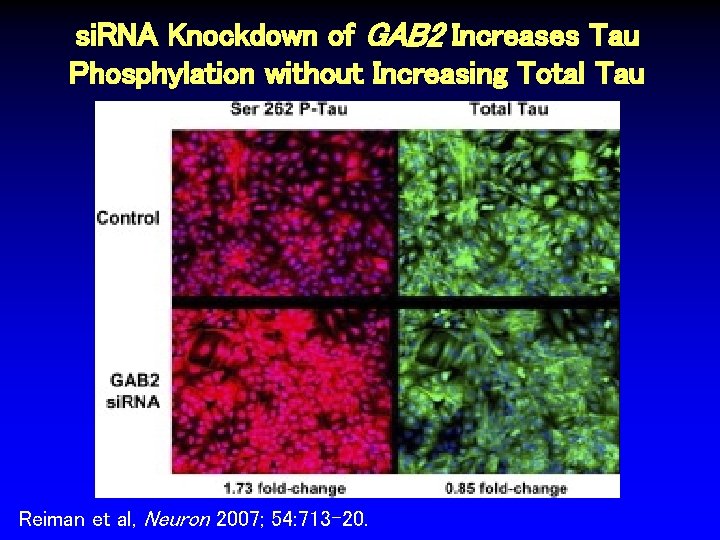
si. RNA Knockdown of GAB 2 Increases Tau Phosphylation without Increasing Total Tau Reiman et al, Neuron 2007; 54: 713 -20.

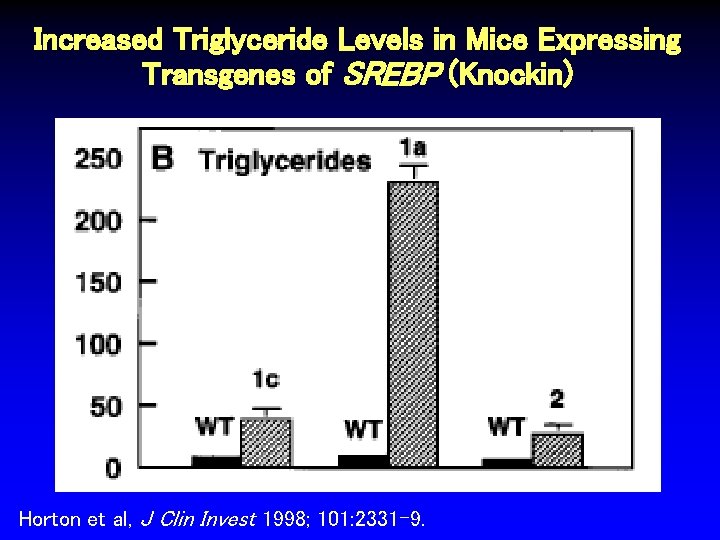
Increased Triglyceride Levels in Mice Expressing Transgenes of SREBP (Knockin) Horton et al, J Clin Invest 1998; 101: 2331 -9.

Post GWA: Finding (Putative) Causal Variants • Narrowing region with fine mapping, sequencing • Structure of association region: nearby genes, conservation • Association with levels of protein product • Co-localization with histopathologic changes • Association with expression levels • Knockdown, knockout