Generalized Linear Models II Distributions link functions diagnostics

Generalized Linear Models II Distributions, link functions, diagnostics (linearity, homoscedasticity, leverage)

Dichotomous key: picking a distribution for your data
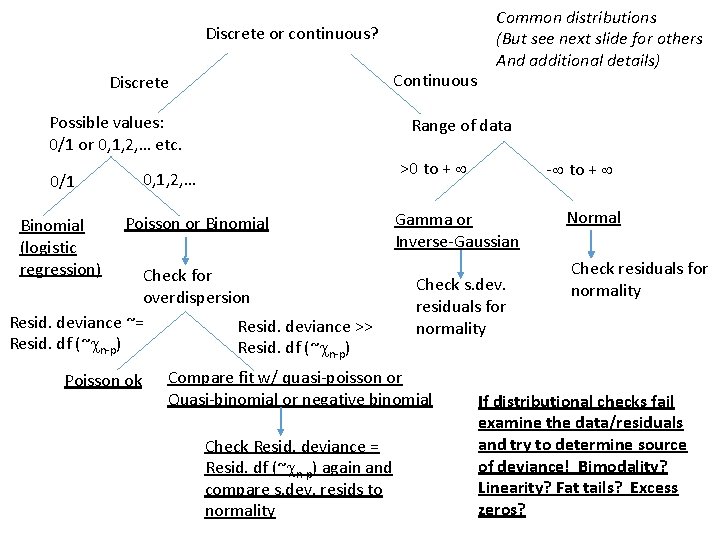
Discrete or continuous? Continuous Discrete Possible values: 0/1 or 0, 1, 2, … etc. Binomial (logistic regression) Range of data >0 to + 0, 1, 2, … 0/1 Poisson or Binomial Check for overdispersion Resid. deviance ~= Resid. deviance >> Resid. df (~ n-p) Poisson ok Common distributions (But see next slide for others And additional details) Gamma or Inverse-Gaussian Check s. dev. residuals for normality Compare fit w/ quasi-poisson or Quasi-binomial or negative binomial Check Resid. deviance = Resid. df (~ n-p) again and compare s. dev. resids to normality - to + Normal Check residuals for normality If distributional checks fail examine the data/residuals and try to determine source of deviance! Bimodality? Linearity? Fat tails? Excess zeros?

Discrete Possible values: 0/1 Bernoulli (successs/failure, logistic regresion? ) 0, 1, 2, … N (known) Binomial Multinomial (# successes in fixed # trials) (more than 2 categories, fixed # trials) 0, 1, 2, … infinity Geometric (# trials to 1 st success) Poisson (#successes in large # trials) Negative Binomial (#trials to nth success or over-dispersed Poisson) 0 to 1 Beta (fraction of total, proportions) >0 to + Exponential Gamma Inverse-Gaussian (time to 1 st success) (time to nth success) ( 1/x is normal) - to + Normal Continuous Check out Wikipedia pages for each distribution for more info!

As sample sizes get large, many distributions converge on the normal distribution • See, e. g. • http: //en. wikipedia. org/wiki/Negative_binomial_di stribution • http: //en. wikipedia. org/wiki/Gamma_distribution

Group exercise • Get a partner • Describe a real dataset to your partner • Partner picks a potentially appropriate distribution • Switch roles • Repeat!

Link Functions • Enforce appropriate range for expected response • (e. g. 0, 1 for ‘probability of success’, >0 for counts, etc) • Linearize relationship between expected response and predictors G(E(y)) = b 0 + b 1 x 1+ b 2 x 2+ etc • Be careful to interpret coefficients properly given a link function! E(y) =G-1( b 0 + b 1 x 1+ b 2 x 2+ etc) • E. g. Link Constraint Log E(y)>0 Logit E(y) in (0, 1) Inverse See Table 15. 1 in GLM chapter for lots more!

Canonical link functions

Sample problems for count data • Binomial vs. poisson • http: //personal. maths. surrey. ac. uk/st/J. Deane/Teach/se 202/poiss_bin. html
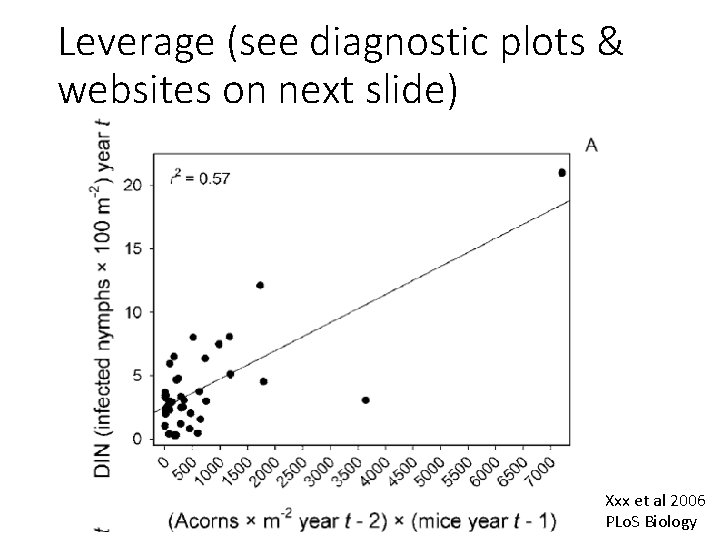
Leverage (see diagnostic plots & websites on next slide) Xxx et al 2006 PLo. S Biology

R: example GLM with data • #read in data • bd=read. csv("c: /marm/teaching/293 qe/bat_lambda. csv") • str(bd); head(bd) • #What not to do- run models blindly! • b 1=glm(Lambda~Pre. WNS_Pop, family=Gamma, data=bd); summary(b 1) • #What to do - plot data • plot(Lambda~Pre. WNS_Pop, data=bd) • #What does it suggest would be a good idea? • bd$Lpop=log(bd$Pre. WNS_Pop) • plot(Lambda~Lpop, data=bd) • b 1=glm(Lambda~Lpop, family=Gamma, data=bd); summary(b 1) • b 2=glm(Lambda~Lpop+Species, family=Gamma, data=bd); summary(b 2) • b 3=glm(Lambda~Lpop*Species, family=Gamma, data=bd); summary(b 3) • anova(b 1, b 2, b 3, test="Chisq") • AIC(b 1, b 2, b 3) • plot(b 3) • http: //stats. stackexchange. com/questions/52089/what-does-having-constant-variance-in-a-linear-regression-model-mean • http: //stats. stackexchange. com/questions/58141/interpreting-plot-lm
- Slides: 11