General Biology 1 BIO 1101 Syllabus Textbook http
General Biology 1 BIO 1101 Syllabus & Textbook: http: //goo. gl/rvgdr. H Lecturer: Michael Gotesman, Ph. D Email: mgotesman@citytech. cuny. edu OER Lecture: https: //openlab. citytech. cuny. edu/bio-oer/page/2/ Lab: https: //openlab. citytech. cuny. edu/bio-oer/ Grade Breakdown: Exams (4): 20% Each Quizzes: 20% Average

Recap: Lecture 26 1. Search for Genetic Molecule of Information: 1911 – Thomas Hunt Morgan (Fruit flies red/white eye) 1856 -1863 – Gregor Johann Mendel (Pea Plants) 2. DNA – Heritable Molecule for Genetic Information: 1929 – Frederick Griffith (R - nonlethal vs S - lethal Pneumonia strain) 1944 -- Oswald T. Avery, Colin M. Mac. Leod, and Maclyn Mc. Carty (Fractionation of S-strain macromolecules) 1952 -- Alfred Hershey and Martha Chase (radioactively labeled bacteriophage) 3. (Erwin) Chargaff’s Rules A. The amounts of A, T, G, and C in DNA: 1. Identical in identical twins 2. Varies between individuals of a species 3. Varies more from species to species B. In each species, there are equal amounts of: A & T, G & C 4. DNA – Double helix 1. Rosalind Franklin – X-ray patterns 2. Maurice Wilkins provided the diffraction pattern to Watson and Crick 3. Double helix with the phosphate-sugar backbone on the outside
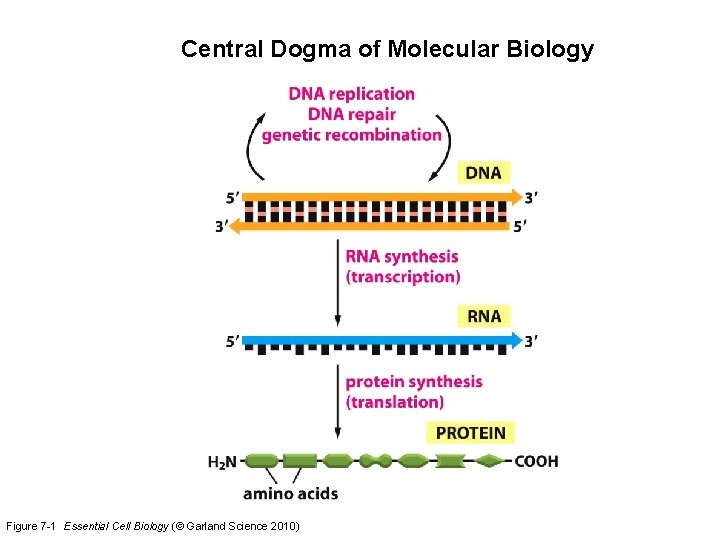
Central Dogma of Molecular Biology Figure 7 -1 Essential Cell Biology (© Garland Science 2010)

Structure of DNA l DNA contains: l Two Nucleotides with purine bases Adenine (A) l Guanine (G) l l Two Nucleotides with pyrimidine bases Thymine (T) l Cytosine (C) l 4
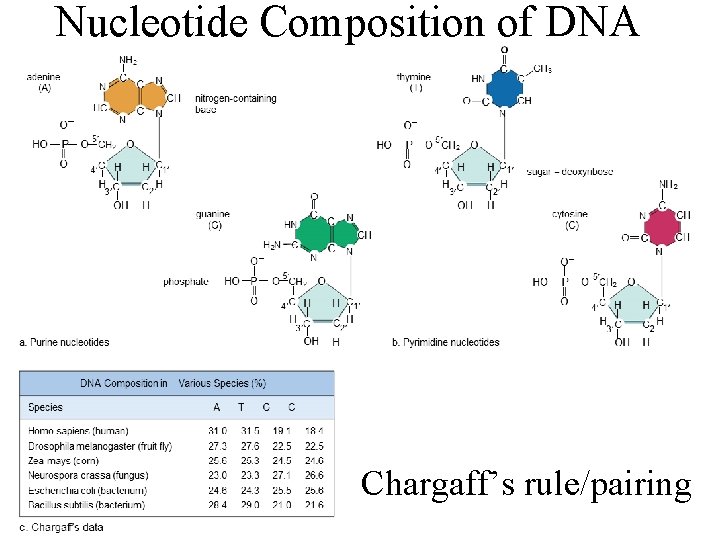
Nucleotide Composition of DNA Chargaff’s rule/pairing
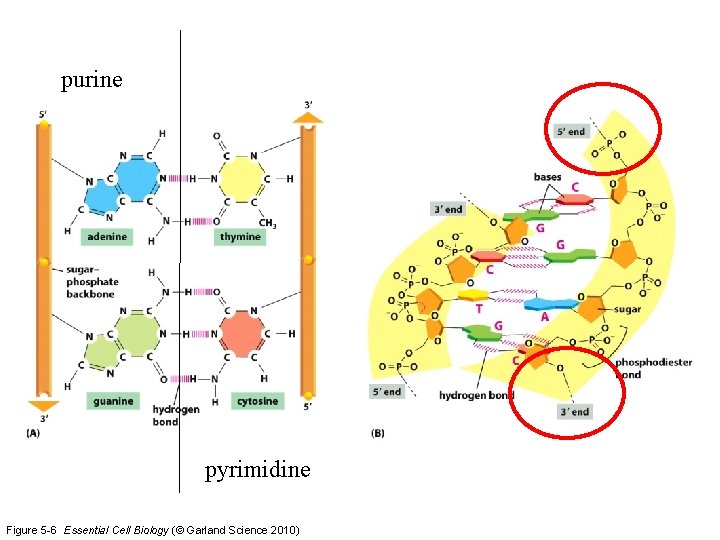
purine pyrimidine Figure 5 -6 Essential Cell Biology (© Garland Science 2010)
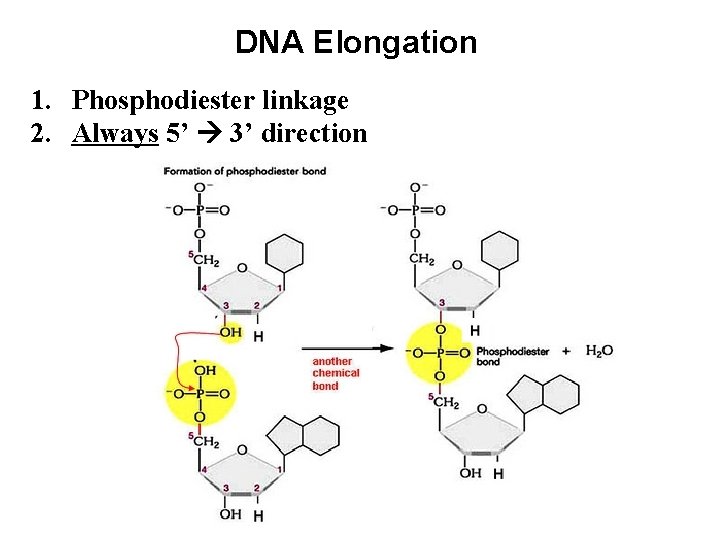
DNA Elongation 1. Phosphodiester linkage 2. Always 5’ 3’ direction
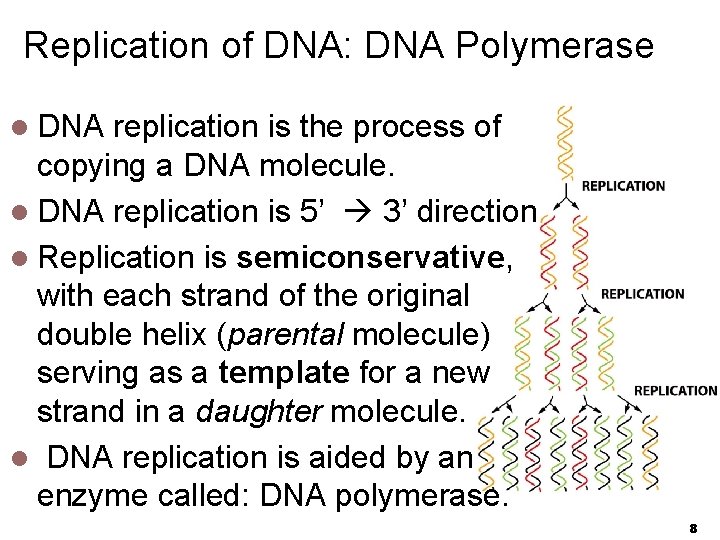
Replication of DNA: DNA Polymerase l DNA replication is the process of copying a DNA molecule. l DNA replication is 5’ 3’ direction l Replication is semiconservative, with each strand of the original double helix (parental molecule) serving as a template for a new strand in a daughter molecule. l DNA replication is aided by an enzyme called: DNA polymerase. 8

Replication Errors l Genetic variations are the raw material for evolutionary change l Mutation: l l Why should you care? A permanent (but unplanned) change in basepair sequence l Some due to errors in DNA replication l Others are due to DNA damage DNA repair enzymes are usually available to reverse most errors 9

Function of Genes l Genes Specify Enzymes l Beadle and Tatum: Experiments on fungus Neurospora crassa l Proposed that each gene specifies the synthesis of one enzyme l One-gene-one-enzyme hypothesis l l Genes Specify a Polypeptide A gene is a segment of DNA that specifies the sequence of amino acids in a polypeptide l Suggests that genetic mutations cause changes in the primary structure of a protein l What is primary structure of a protein? 10
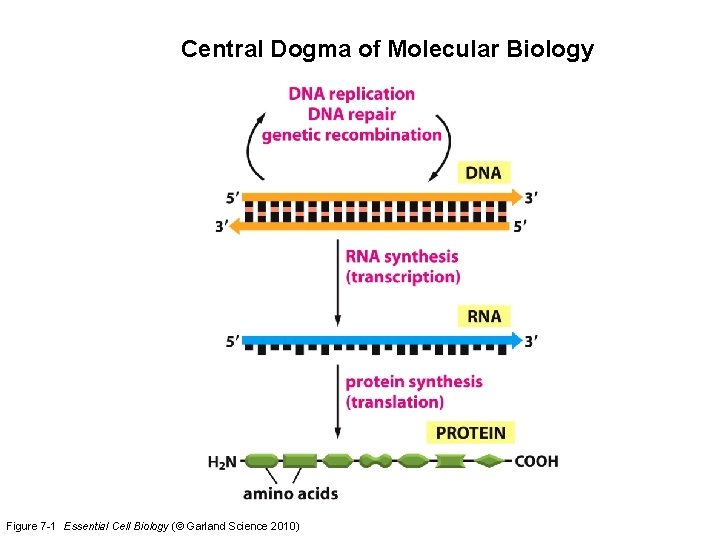
Central Dogma of Molecular Biology Figure 7 -1 Essential Cell Biology (© Garland Science 2010)

Types of RNA l RNA is a polymer of RNA nucleotides l RNA Nucleotides are of four types: Uracil, Adenine, Cytosine, and Guanine l Uracil (U) replaces thymine (T) of DNA l Types of RNA Messenger (m. RNA) - Takes genetic message from DNA in nucleus to ribosomes in cytoplasm l Ribosomal (r. RNA) - Makes up ribosomes which read the message in m. RNA l Transfer (t. RNA) - Transfers appropriate amino acid to ribosome when “instructed” l 12
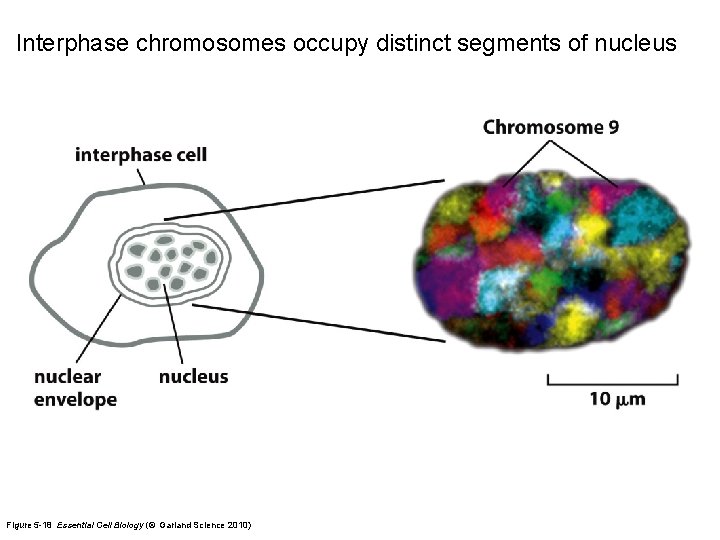
Interphase chromosomes occupy distinct segments of nucleus Figure 5 -18 Essential Cell Biology (© Garland Science 2010)
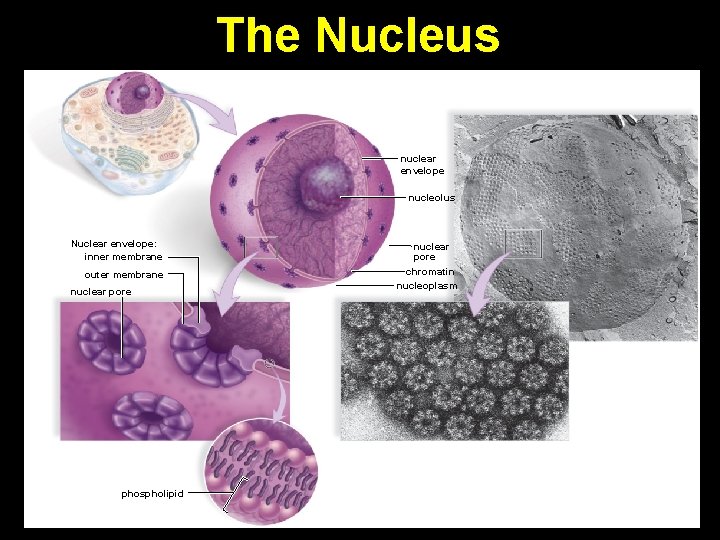
The Nucleus nuclear envelope nucleolus Nuclear envelope: inner membrane outer membrane nuclear pore phospholipid nuclear pore chromatin nucleoplasm

Protein Synthesis: From DNA to RNA to Protein l The mechanism of gene expression DNA in genes specify information, but information is not structure and function of DNA l Genetic info is expressed into structure & function through protein synthesis l l The expression of genetic info into structure & function: DNA in gene controls the sequence of nucleotides in an RNA molecule l RNA controls the primary structure of a protein l 15

Steps in Gene Expression: Transcription l Transcription Gene unzips and exposes unpaired bases l Serves as template for m. RNA formation l Loose RNA nucleotides bind to exposed DNA bases using the C=G & A=U rule l When entire gene is transcribed into m. RNA, result is a pre-m. RNA transcript of the gene l The base sequence in the pre-m. RNA is complementary to the base sequence in DNA l 16

Animation: Prokaryote Please note that due to differing operating systems, some animations will not appear until the presentation is viewed in Presentation Mode (Slide Show view). You may see blank slides in the “Normal” or “Slide Sorter” views. All animations will appear after viewing in Presentation Mode and playing each animation. Most animations will require the latest version of the Flash Player, which is available at http: //get. adobe. com/flashplayer. 17
Animation: Eukaryote Please note that due to differing operating systems, some animations will not appear until the presentation is viewed in Presentation Mode (Slide Show view). You may see blank slides in the “Normal” or “Slide Sorter” views. All animations will appear after viewing in Presentation Mode and playing each animation. Most animations will require the latest version of the Flash Player, which is available at http: //get. adobe. com/flashplayer.

Processing Messenger RNA l Pre-m. RNA, is modified before leaving the eukaryotic nucleus. l Modifications to ends of primary transcript: l Cap of modified guanine on 5′ end l l The cap is a modified guanine (G) nucleotide Helps a ribosome where to attach when translation begins Poly-A tail of 150+ adenines on 3′ end l Facilitates the transport of m. RNA out of the nucleus l Inhibits degradation of m. RNA by hydrolytic enzymes. l 19

Processing Messenger RNA l Pre-m. RNA, is composed of exons and introns. l l The exons will be expressed, The introns, occur in between the exons. l l Allows a cell to pick and choose which exons will go into a particular m. RNA splicing: l Primary transcript consists of: l l l Performed by spliceosome complexes in nucleoplasm l l l Some segments that will not be expressed (introns) Segments that will be expressed (exons) Introns are excised Remaining exons are spliced back together Result is mature m. RNA transcript 20
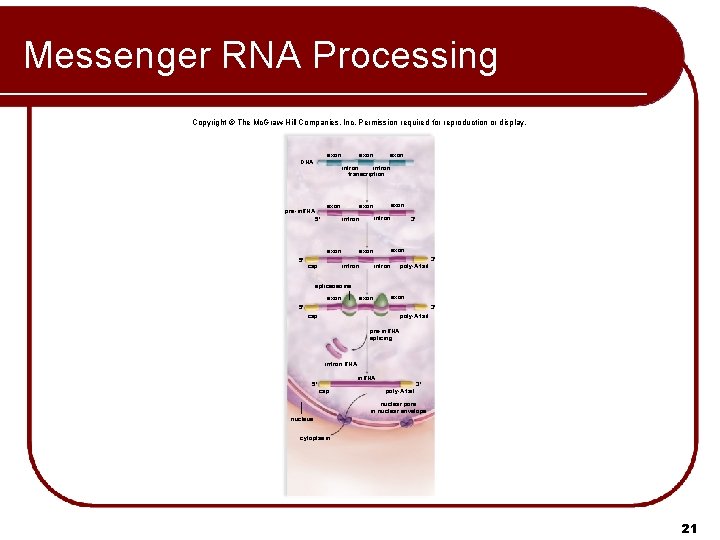
Messenger RNA Processing Copyright © The Mc. Graw-Hill Companies, Inc. Permission required for reproduction or display. exon DNA exon pre-m. RNA exon cap exon intron 5' 5' exon intron transcription 3' exon poly-A tail intron 3' spliceosome exon 3' 5' cap poly-A tail pre-m. RNA splicing intron RNA m. RNA 5' cap 3' poly-A tail nuclear pore in nuclear envelope nucleus cytoplasm 21
Animation Please note that due to differing operating systems, some animations will not appear until the presentation is viewed in Presentation Mode (Slide Show view). You may see blank slides in the “Normal” or “Slide Sorter” views. All animations will appear after viewing in Presentation Mode and playing each animation. Most animations will require the latest version of the Flash Player, which is available at http: //get. adobe. com/flashplayer.

Functions of Introns l As organismal complexity increases; l l l Number of protein-coding genes does not keep pace But the proportion of the genome that is introns increases Humans: l l l Possible functions of introns: l More bang for buck l l Genome has only about 25, 000 coding genes Up to 95% of this DNA genes is introns Exons might combine in various combinations Would allow different m. RNAs to result from one segment of DNA Introns might regulate gene expression Exciting new picture of the genome is emerging 23
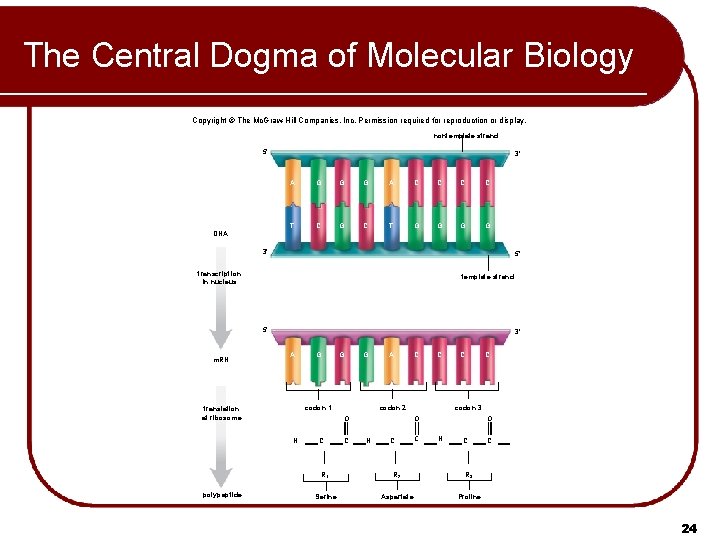
The Central Dogma of Molecular Biology Copyright © The Mc. Graw-Hill Companies, Inc. Permission required for reproduction or display. nontemplate strand 5' 3' A G G G A C C T C G C T G G DNA 3' 5' transcription in nucleus template strand 5' m. RN 3' A G G codon 1 translation at ribosome A C C C codon 2 O N polypeptide G C C codon 3 O N C C C O N C R 1 R 2 R 3 Serine Aspartate Proline C 24
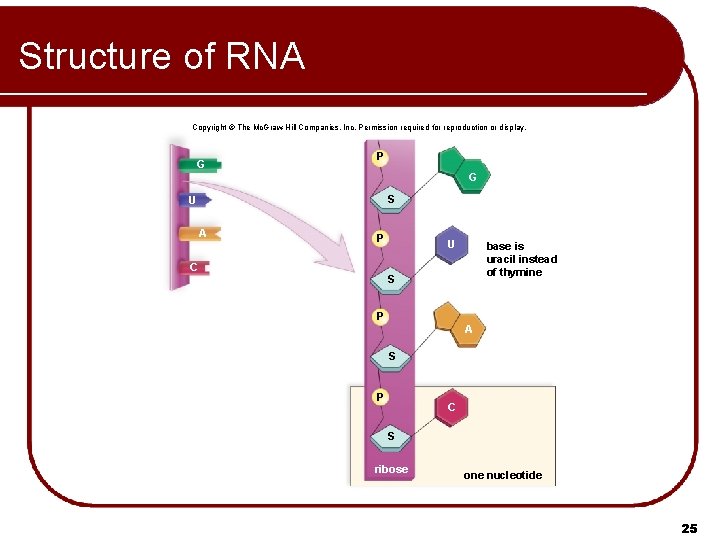
Structure of RNA Copyright © The Mc. Graw-Hill Companies, Inc. Permission required for reproduction or display. G P G S U A P C U base is uracil instead of thymine S P A S P C S ribose one nucleotide 25
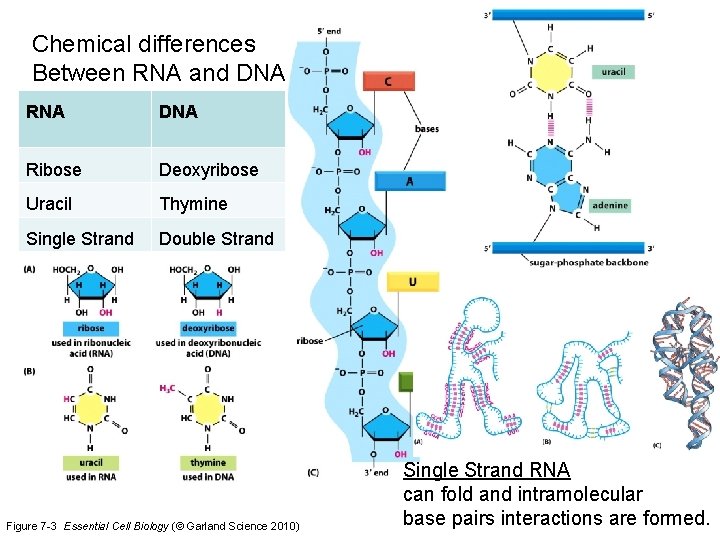
Chemical differences Between RNA and DNA RNA DNA Ribose Deoxyribose Uracil Thymine Single Strand Double Strand Figure 7 -3 Essential Cell Biology (© Garland Science 2010) Single Strand RNA can fold and intramolecular base pairs interactions are formed.
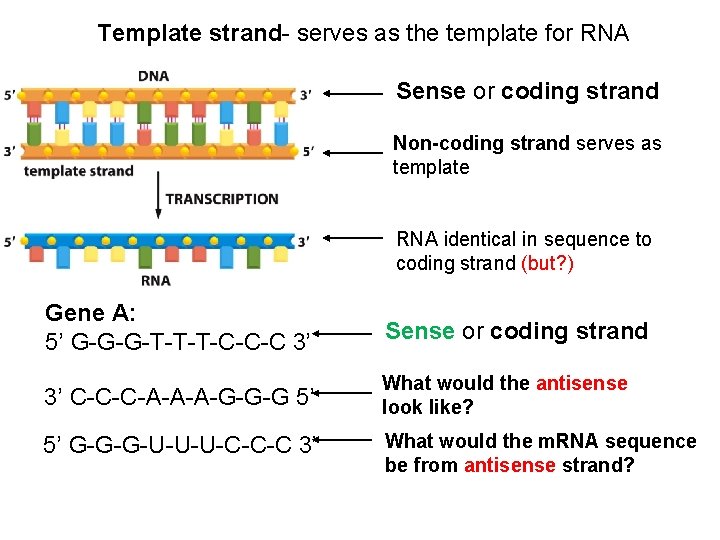
Template strand- serves as the template for RNA Sense or coding strand Non-coding strand serves as template RNA identical in sequence to coding strand (but? ) Gene A: 5’ G-G-G-T-T-T-C-C-C 3’ Sense or coding strand 3’ C-C-C-A-A-A-G-G-G 5’ What would the antisense look like? 5’ G-G-G-U-U-U-C-C-C 3’ What would the m. RNA sequence be from antisense strand?

RNA vs. DNA structure 28
General Biology 1 BIO 1101 Syllabus & Textbook: http: //goo. gl/rvgdr. H Lecturer: Michael Gotesman, Ph. D Email: mgotesman@citytech. cuny. edu OER Lecture: https: //openlab. citytech. cuny. edu/bio-oer/page/2/ Lab: https: //openlab. citytech. cuny. edu/bio-oer/ Grade Breakdown: Exams (4): 20% Each Quizzes: 20% Average
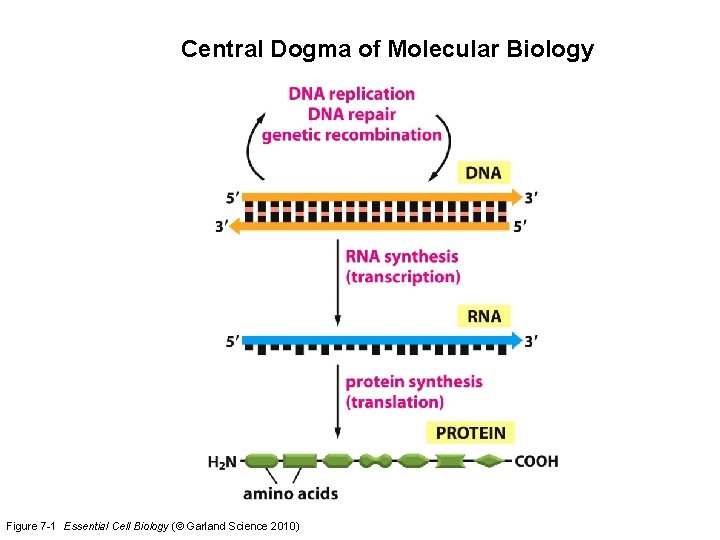
Central Dogma of Molecular Biology Figure 7 -1 Essential Cell Biology (© Garland Science 2010)

Types of RNA l RNA is a polymer of RNA nucleotides l RNA Nucleotides are of four types: Uracil, Adenine, Cytosine, and Guanine l Uracil (U) replaces thymine (T) of DNA l Types of RNA Messenger (m. RNA) - Takes genetic message from DNA in nucleus to ribosomes in cytoplasm l Transfer (t. RNA) - Transfers appropriate amino acid to ribosome when “instructed” l Ribosomal (r. RNA) - Makes up ribosomes which read the message in m. RNA l 31
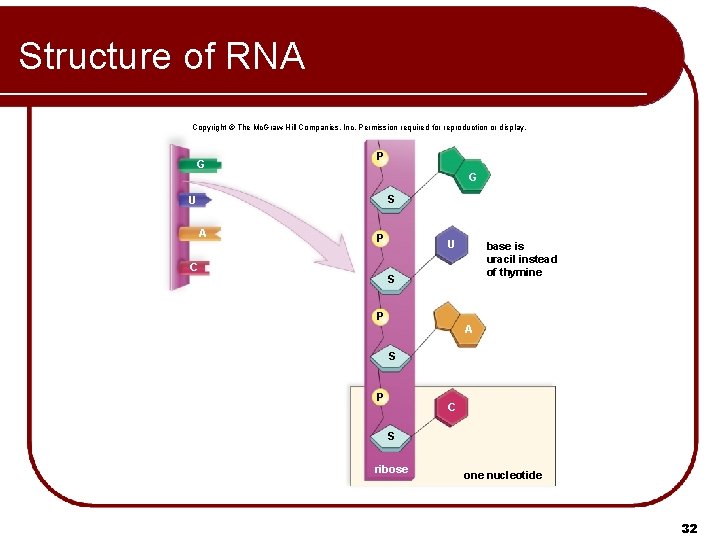
Structure of RNA Copyright © The Mc. Graw-Hill Companies, Inc. Permission required for reproduction or display. G P G S U A P C U base is uracil instead of thymine S P A S P C S ribose one nucleotide 32
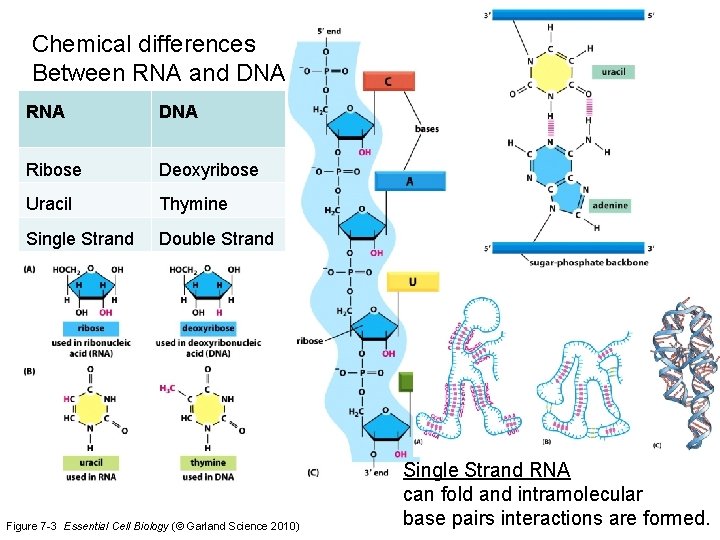
Chemical differences Between RNA and DNA RNA DNA Ribose Deoxyribose Uracil Thymine Single Strand Double Strand Figure 7 -3 Essential Cell Biology (© Garland Science 2010) Single Strand RNA can fold and intramolecular base pairs interactions are formed.
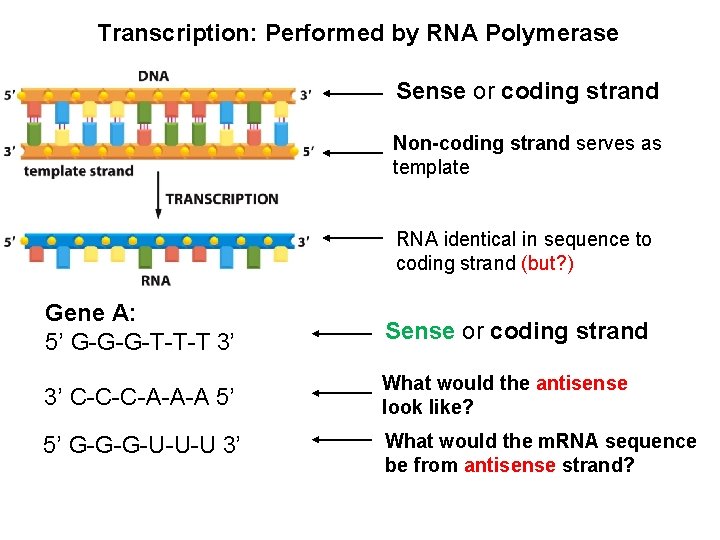
Transcription: Performed by RNA Polymerase Sense or coding strand Non-coding strand serves as template RNA identical in sequence to coding strand (but? ) Gene A: 5’ G-G-G-T-T-T 3’ Sense or coding strand 3’ C-C-C-A-A-A 5’ What would the antisense look like? 5’ G-G-G-U-U-U 3’ What would the m. RNA sequence be from antisense strand?

The minimal unit of information corresponding to an amino acid is 3 bases • Codon is the minimal triplet bases required to encode an amino acid • 4 bases possible in triplets = 43 or 64 combinations • Since there are only 20 amino acids, then 64 permits for degenerate coding Triplet = codon 5’ end always on the left Figure 7 -24 Essential Cell Biology (© Garland Science 2010)
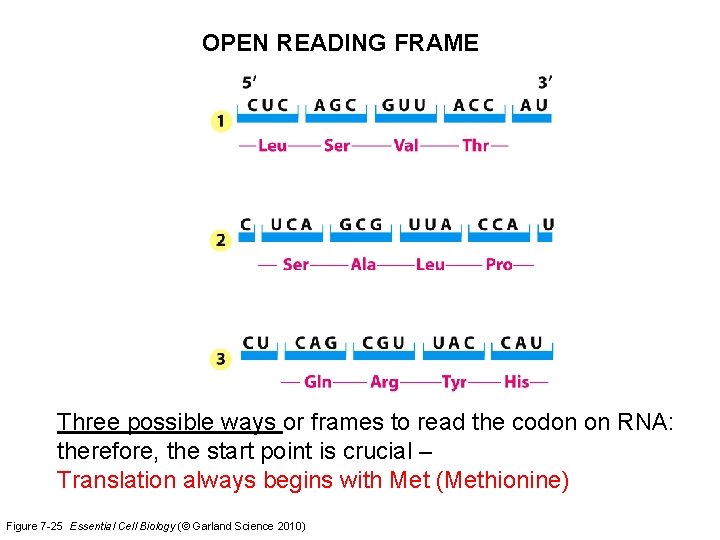
OPEN READING FRAME Three possible ways or frames to read the codon on RNA: therefore, the start point is crucial – Translation always begins with Met (Methionine) Figure 7 -25 Essential Cell Biology (© Garland Science 2010)

t. RNA molecules come in 64 different kinds l All very similar except that l l l One end bears a specific triplet (of the 64 possible) called the anticodon Other end binds with a specific amino acid type t. RNA synthetases attach correct amino acid to the correct t. RNA molecule All t. RNA molecules with a specific anticodon will always bind with the same amino acid 37
Animation Please note that due to differing operating systems, some animations will not appear until the presentation is viewed in Presentation Mode (Slide Show view). You may see blank slides in the “Normal” or “Slide Sorter” views. All animations will appear after viewing in Presentation Mode and playing each animation. Most animations will require the latest version of the Flash Player, which is available at http: //get. adobe. com/flashplayer.
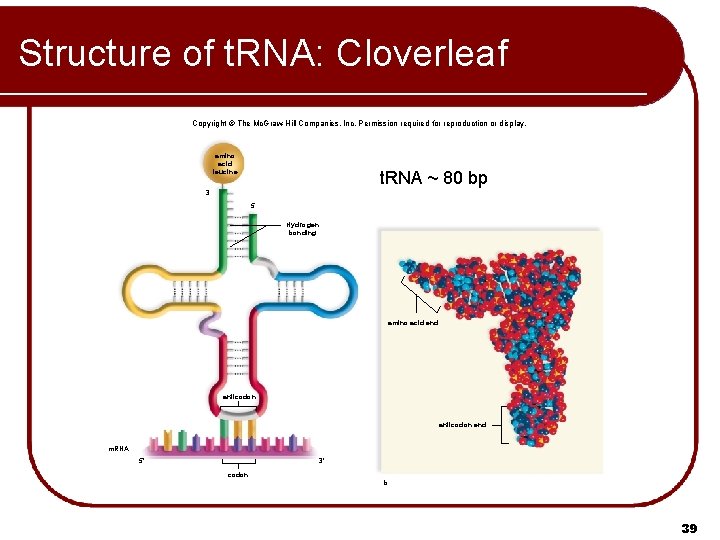
Structure of t. RNA: Cloverleaf Copyright © The Mc. Graw-Hill Companies, Inc. Permission required for reproduction or display. amino acid leucine t. RNA ~ 80 bp 3 5 Hydrogen bonding amino acid end anticodon end m. RNA 5' 3' codon b. 39

Ribosomes l l Ribosomal RNA (r. RNA): l Produced from a DNA template in the nucleolus l Combined with proteins into large and small ribosomal subunits A completed ribosome has three binding sites to facilitate pairing between t. RNA and m. RNA l The E (for exit) site l The P (for peptide) site, and l The A (for amino acid) site 40

Animation Please note that due to differing operating systems, some animations will not appear until the presentation is viewed in Presentation Mode (Slide Show view). You may see blank slides in the “Normal” or “Slide Sorter” views. All animations will appear after viewing in Presentation Mode and playing each animation. Most animations will require the latest version of the Flash Player, which is available at http: //get. adobe. com/flashplayer.
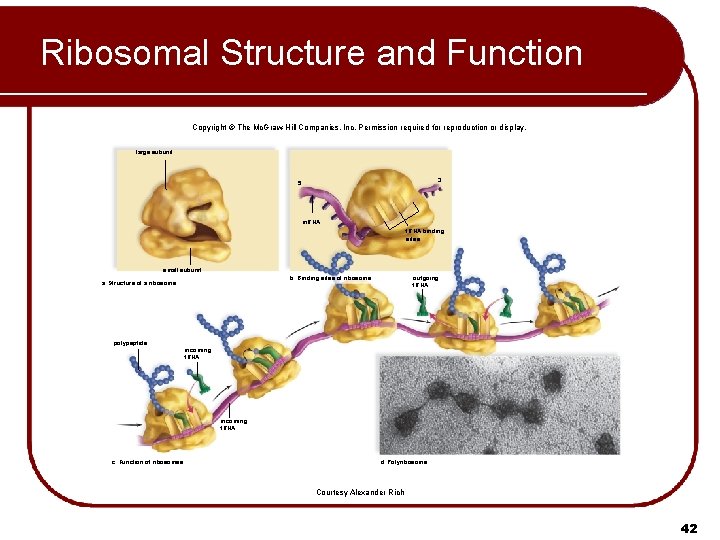
Ribosomal Structure and Function Copyright © The Mc. Graw-Hill Companies, Inc. Permission required for reproduction or display. large subunit 3 5 m. RNA t. RNA binding sites small subunit polypeptide outgoing t. RNA b. Binding sites of ribosome a. Structure of a ribosome incoming t. RNA c. Function of ribosomes d. Polyribosome Courtesy Alexander Rich 42
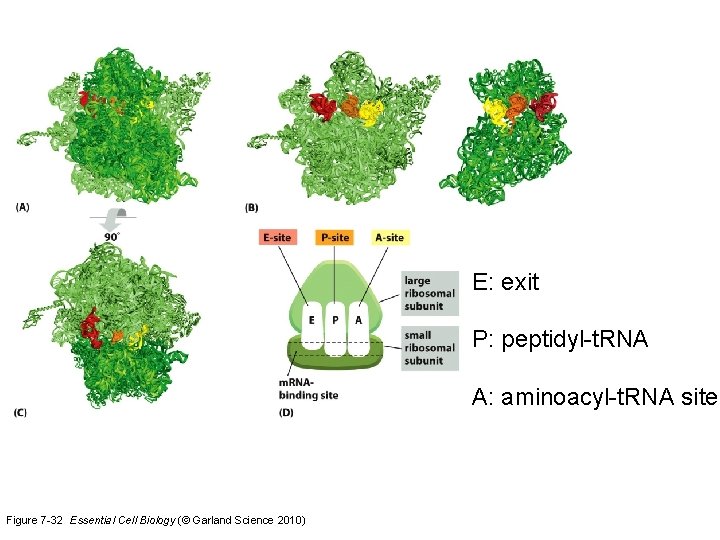
E: exit P: peptidyl-t. RNA A: aminoacyl-t. RNA site Figure 7 -32 Essential Cell Biology (© Garland Science 2010)

Animation: Translation Please note that due to differing operating systems, some animations will not appear until the presentation is viewed in Presentation Mode (Slide Show view). You may see blank slides in the “Normal” or “Slide Sorter” views. All animations will appear after viewing in Presentation Mode and playing each animation. Most animations will require the latest version of the Flash Player, which is available at http: //get. adobe. com/flashplayer.

Steps in Translation: Initiation l Small ribosomal subunit attaches to m. RNA transcript l Beginning of transcript always has the START codon (AUG) – Codes for Methionine (Met) l Initiator t. RNA (UAC) attaches to P site l Large ribosomal subunit joins the small subunit 45
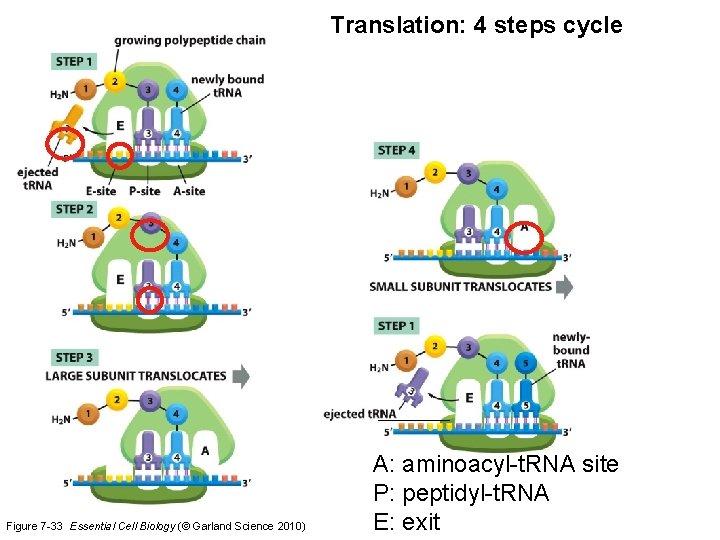
Translation: 4 steps cycle Figure 7 -33 Essential Cell Biology (© Garland Science 2010) A: aminoacyl-t. RNA site P: peptidyl-t. RNA E: exit
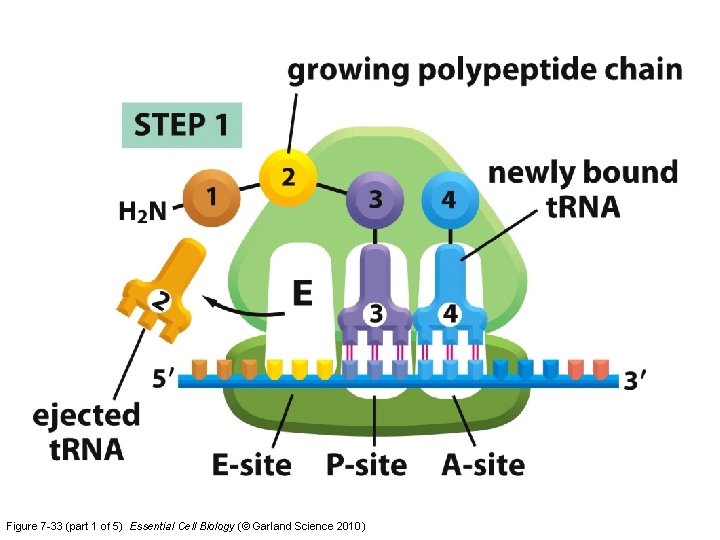
Figure 7 -33 (part 1 of 5) Essential Cell Biology (© Garland Science 2010)
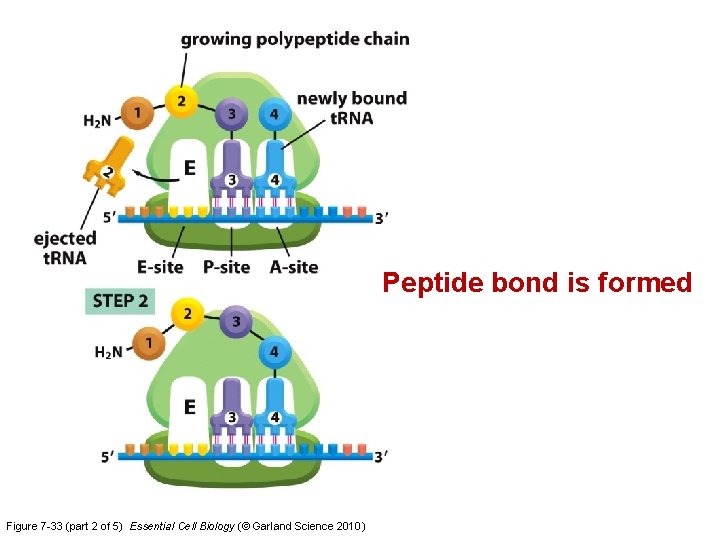
Peptide bond is formed Figure 7 -33 (part 2 of 5) Essential Cell Biology (© Garland Science 2010)
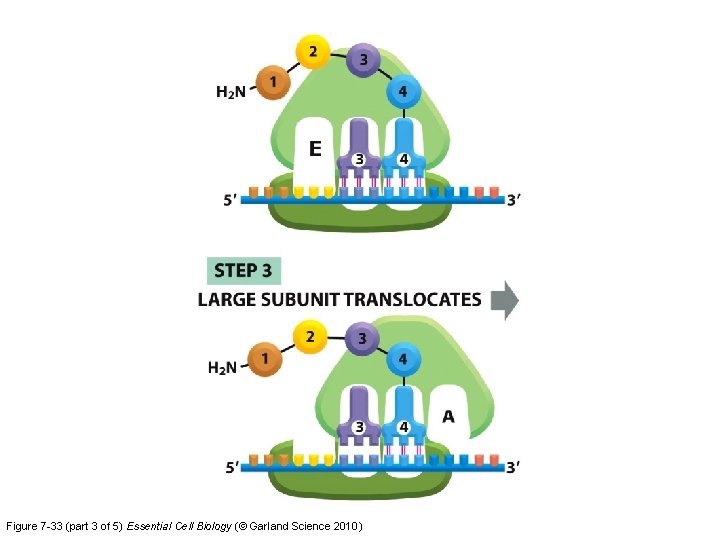
Figure 7 -33 (part 3 of 5) Essential Cell Biology (© Garland Science 2010)
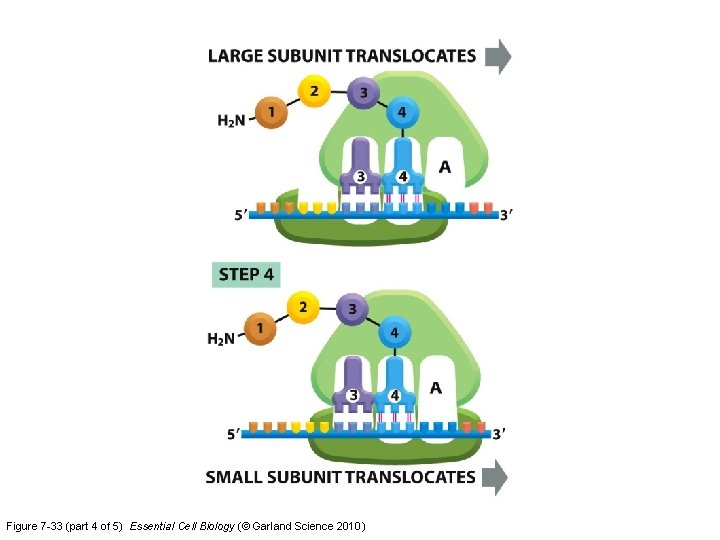
Figure 7 -33 (part 4 of 5) Essential Cell Biology (© Garland Science 2010)
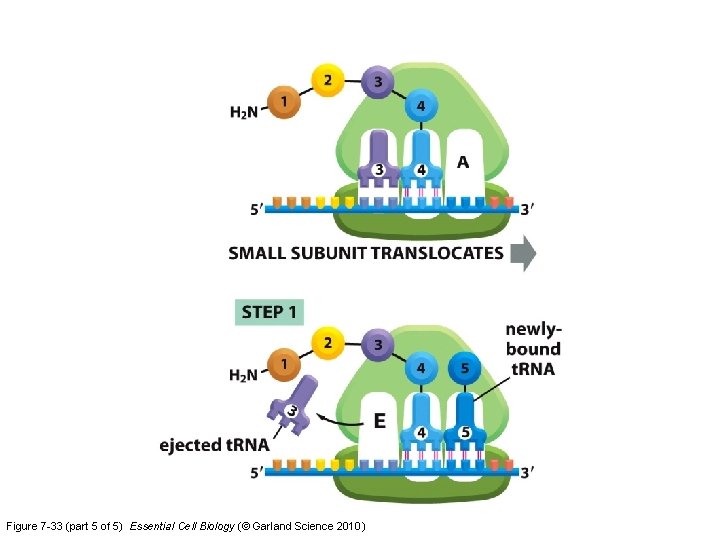
Figure 7 -33 (part 5 of 5) Essential Cell Biology (© Garland Science 2010)
Animation Please note that due to differing operating systems, some animations will not appear until the presentation is viewed in Presentation Mode (Slide Show view). You may see blank slides in the “Normal” or “Slide Sorter” views. All animations will appear after viewing in Presentation Mode and playing each animation. Most animations will require the latest version of the Flash Player, which is available at http: //get. adobe. com/flashplayer.
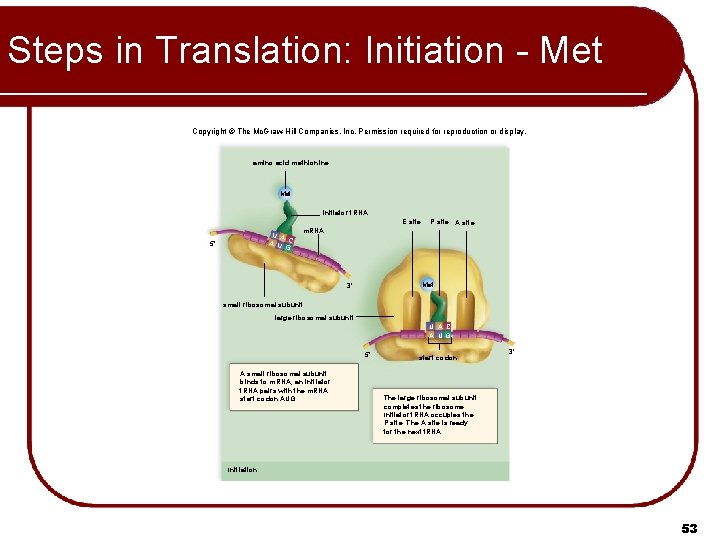
Steps in Translation: Initiation - Met Copyright © The Mc. Graw-Hill Companies, Inc. Permission required for reproduction or display. amino acid methionine Met initiator t. RNA E site U A A U C G 5' P site A site m. RNA Met 3' small ribosomal subunit large ribosomal subunit U A C A UG 5' A small ribosomal subunit binds to m. RNA; an initiator t. RNA pairs with the m. RNA start codon AUG. start codon 3' The large ribosomal subunit completes the ribosome. Initiator t. RNA occupies the P site. The A site is ready for the next t. RNA. Initiation 53

Steps in Translation: Elongation l “Elongation” refers to the growth in length of the polypeptide l RNA molecules bring their amino acid fares to the ribosome l Ribosome reads a codon in the m. RNA Allows only one type of t. RNA to bring its amino acid l Must have the anticodon complementary to the m. RNA codon being read l Joins the ribosome at it’s A site l l Methionine of initiator is connected to amino acid of 2 nd t. RNA by peptide bond 54

Steps in Translation: Elongation Second t. RNA moves to P site (translocation) l Spent initiator moves to E site and exits l Ribosome reads the next codon in the m. RNA l l Allows only one type of t. RNA to bring its amino acid l l l Must have the anticodon complementary to the m. RNA codon being read Joins the ribosome at it’s A site Dipeptide on 2 nd amino acid is connected to amino acid of 3 nd t. RNA by peptide bond 55
Animation Please note that due to differing operating systems, some animations will not appear until the presentation is viewed in Presentation Mode (Slide Show view). You may see blank slides in the “Normal” or “Slide Sorter” views. All animations will appear after viewing in Presentation Mode and playing each animation. Most animations will require the latest version of the Flash Player, which is available at http: //get. adobe. com/flashplayer.
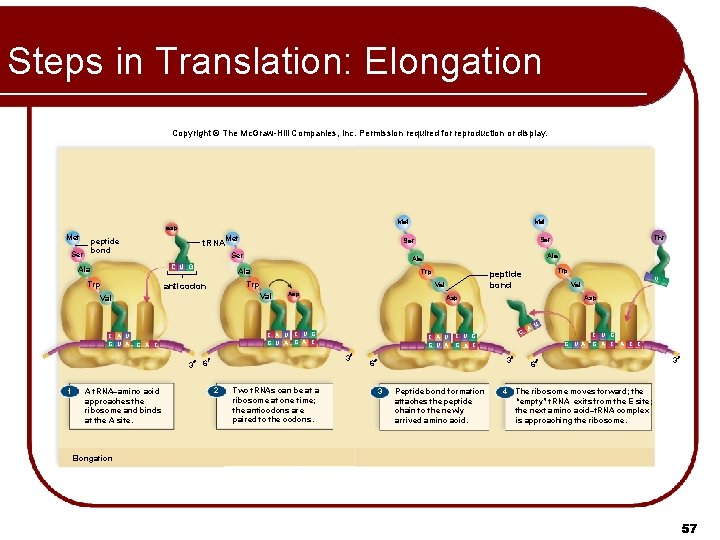
Steps in Translation: Elongation Copyright © The Mc. Graw-Hill Companies, Inc. Permission required for reproduction or display. Met asp Met peptide Ser bond Ser C U G Trp G A C A t. RNA–amino acid approaches the ribosome and binds at the A site. C A U G U A 3 6 2 Two t. RNAs can be at a ribosome at one time; the anticodons are paired to the codons. C Peptide bond formation attaches the peptide chain to the newly arrived amino acid. A U C U G G U A 3 3 G G Asp C U G G A C 6 U Val Asp C A U C U G G U A G A C 3 1 Asp Trp peptide bond Val Val C A U G U A Trp anticodon Ala Ala Thr Ser t. RNA Ala Met 4 G A C C 6 3 The ribosome moves forward; the “empty” t. RNA exits from the E site; the next amino acid–t. RNA complex is approaching the ribosome. Elongation 57

Steps in Translation: Termination Previous t. RNA moves to P site l Spent t. RNA moves to E site and exits l Ribosome reads the STOP codon at the end of the m. RNA l l l UAA, UAG, or UGA Does not code for an amino acid Polypeptide is released from last t. RNA by release factor l Ribosome releases m. RNA and dissociates into subunits l m. RNA read by another ribosome l 58
Animation Please note that due to differing operating systems, some animations will not appear until the presentation is viewed in Presentation Mode (Slide Show view). You may see blank slides in the “Normal” or “Slide Sorter” views. All animations will appear after viewing in Presentation Mode and playing each animation. Most animations will require the latest version of the Flash Player, which is available at http: //get. adobe. com/flashplayer.
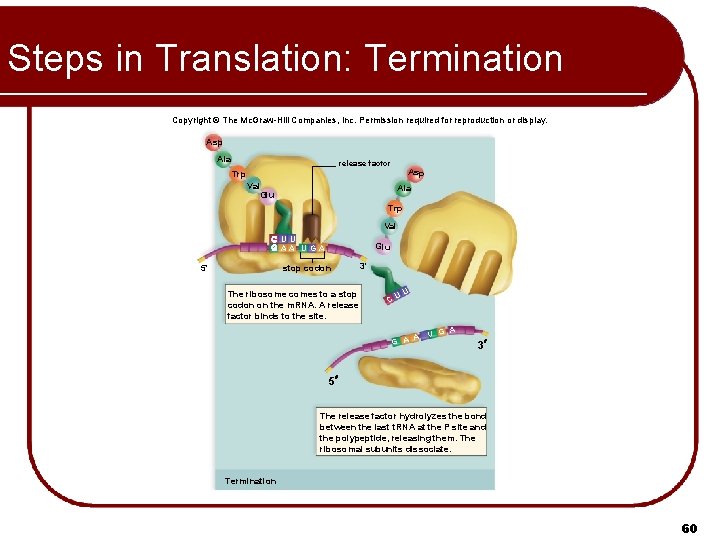
Steps in Translation: Termination Copyright © The Mc. Graw-Hill Companies, Inc. Permission required for reproduction or display. Asp Ala release factor Asp Trp Val Glu Ala Trp Val U U AA UGA Glu stop codon 5' The ribosome comes to a stop codon on the m. RNA. A release factor binds to the site. 3' U CU A G A A U G 3 5 The release factor hydrolyzes the bond between the last t. RNA at the P site and the polypeptide, releasing them. The ribosomal subunits dissociate. Termination 60
Animation Please note that due to differing operating systems, some animations will not appear until the presentation is viewed in Presentation Mode (Slide Show view). You may see blank slides in the “Normal” or “Slide Sorter” views. All animations will appear after viewing in Presentation Mode and playing each animation. Most animations will require the latest version of the Flash Player, which is available at http: //get. adobe. com/flashplayer. 61
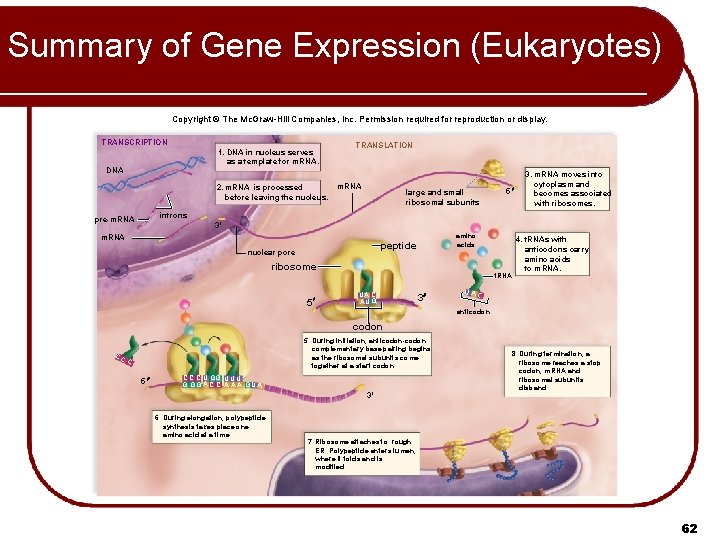
Summary of Gene Expression (Eukaryotes) Copyright © The Mc. Graw-Hill Companies, Inc. Permission required for reproduction or display. TRANSCRIPTION 1. DNA in nucleus serves as a template for m. RNA. TRANSLATION DNA 2. m. RNA is processed before leaving the nucleus. m. RNA large and small ribosomal subunits 3. m. RNA moves into cytoplasm and becomes associated with ribosomes. 5 introns pre-m. RNA 3' m. RNA amino acids peptide nuclear pore ribosome 5 t. RNA UA C A UG 3 UA 4. t. RNAs with anticodons carry amino acids to m. RNA. C anticodon CC 5. During initiation, anticodon-codon complementary base pairing begins as the ribosomal subunits come together at a start codon. C 5 C C C U GG U U U G GG A C C A A A G U A 3' 6. During elongation, polypeptide synthesis takes place one amino acid at a time. 8. During termination, a ribosome reaches a stop codon; m. RNA and ribosomal subunits disband. 7. Ribosome attaches to rough ER. Polypeptide enters lumen, where it folds and is modified. 62
Animation Please note that due to differing operating systems, some animations will not appear until the presentation is viewed in Presentation Mode (Slide Show view). You may see blank slides in the “Normal” or “Slide Sorter” views. All animations will appear after viewing in Presentation Mode and playing each animation. Most animations will require the latest version of the Flash Player, which is available at http: //get. adobe. com/flashplayer.

Review l Genetic Material l l DNA Structure l l Transformation Watson and Crick DNA Replication l Prokaryotic versus Eukaryotic l Replication Errors l Transcription l Translation l Structure of Eukaryotic Chromosome 64
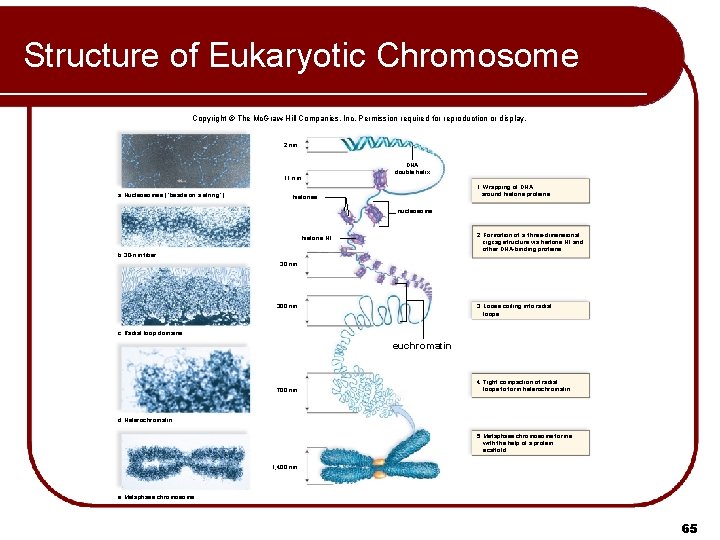
Structure of Eukaryotic Chromosome Copyright © The Mc. Graw-Hill Companies, Inc. Permission required for reproduction or display. 2 nm DNA double helix 11 nm a. Nucleosomes (“beads on a string”) 1. Wrapping of DNA around histone proteins. histones nucleosome 2. Formation of a three-dimensional zigzag structure via histone H 1 and other DNA-binding proteins. histone H 1 b. 30 -nm fiber 30 nm 300 nm 3. Loose coiling into radial loops. c. Radial loop domains euchromatin 700 nm 4. Tight compaction of radial loops to form heterochromatin. d. Heterochromatin 5. Metaphase chromosome forms with the help of a protein scaffold. 1, 400 nm e. Metaphase chromosome 65
- Slides: 65