Gene Weaver A prototype for bioinformatics Michael Luck

Gene. Weaver: A prototype for bioinformatics Michael Luck University of Southampton, UK Kevin Bryson and David Jones, UCL Mike Joy, University of Warwick
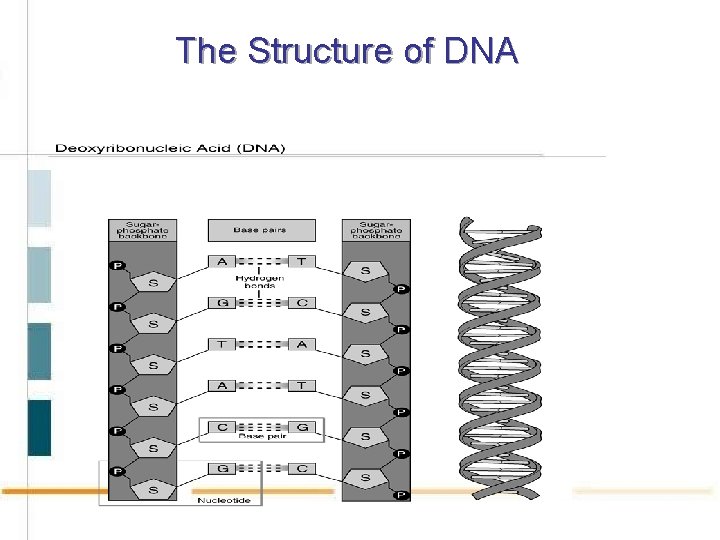
The Structure of DNA

The Result of 15 Years Hard Work > contig 1 TAAGTTATTATTTAGTTAATACTTTTAACAATATTATTAAGGTATTTAAAAAATACTATTATAGTATTTA ACATAGTTAAATACCTTAATACTGTTAAATTATATTCAATACATATATAATATTATTAAAAT ACTTGATAAGTATTATTTAGATATTAGACAAATACTAATTTTATATTGCTTTAATACTTAATACTA CTTATGTATTAAGTAAATATTACTGTAATACTAATAACAATATTATTACAATATGCTAGAATAATATTGC TAGTATCAATAATTACTAATATAGTATTAGGAAAATACCATAATAATATTTCTACATAATACTAAGTTAA TACTATGTGTAGAATAATAATCAGATTAAAAAAATTTTATCTGAAACATATTTAATCAATTG AACTGATTATTTTCAGCAGTAATAATTACATATGTACATATGTAAAATATCATTAATTTCTGT TATAATAGTATCTATTTTAGAGAGTATTATTACTATAATTAAGCATTTATGCTTAATTATAA GCTTTTTATGAACAAAATTATAGACATTTTAGTTCTTATAATAATAGATATTAAAGAAAATAAAAA AATAGAAATATCATAACCCTTGATAACCCAGAAATTAATACTTAATCAAAAATGAAAATATTAATT AATAAAAGTGAATAAAATTTTGGGAAAAAATGAATAACGTTATTATTTCCAATAACAAAATAAAA CCACATCATATTTTTTAATAGAGGCAAAAGAAATAAACTTTTATGCTAACAATGAATACT TTTCTGTCAAATGTAATTTAAAAATATTGATATTCTTGAACAAGGCTCCTTAATTGTTAAAGGAAA AATTTTTAACGATCTTATTAATGGCATAAAAGAAGAGATTATTACTATTCAAGAAAAAGATCAAACACTT TTGGTTAAAACAAAACAAGTATTAATTTAAACACAATTAATGTGAATTTCCAAGAATAAGGT TTAATGAAAAAAACGATTTAAGTGAATTTAATCAATTCAAAATTATTCACTTTTAGTAAAAGGCAT TAAAAAAATTTTTCACTCAGTTTCAAATAATCGTGAAATATCTTCTAAATTTAATGGAGTAAATTTCAAT GGATCCAATGGAAAAGAAATATTTTTAGAAGCTTCTGACACTTATAAACTATCTGTTTTTGAGATAAAGC AAGAAACAGAACCATTTGATTTCATTTTGGAGAGTAATTTACTTAGTTTCATTAATTCTTTTAATCCTGA AGAAGATAAATCTATTGTTTTTTATTACAGAAAAGATAATAAAGATAGCTTTAGTACAGAAATGTTGATT TCAATGGATAACTTTATGATTAGTTACACATCGGTTAATGAAAAATTTCCAGAGGTAAACTACTTTTTTG AATTTGAACCTGAAACTAAAATAGTTGTTCAAAAAAATGAATTAAAAGATGCACTTCAAAGAATTCAAA etc etc
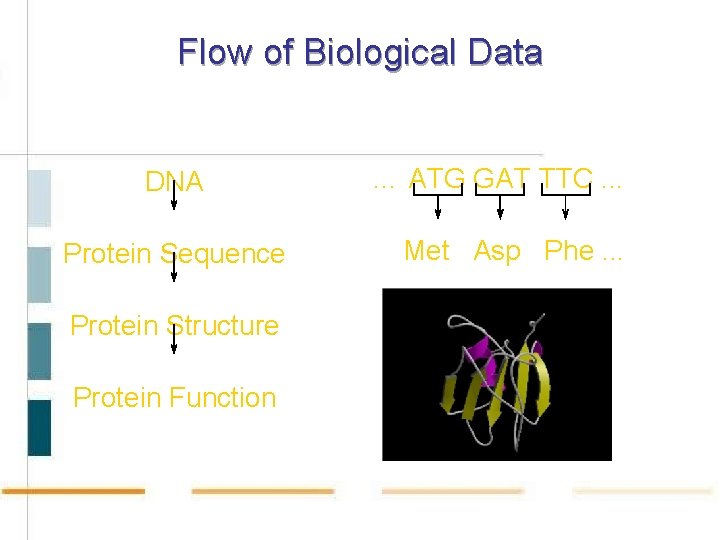
Flow of Biological Data DNA Protein Sequence Protein Structure Protein Function … ATG GAT TTC. . . Met Asp Phe. . .

Data Analysis Lots of primary data -- need to discover gene function. n Scan databases for similar sequences n Collect matching sequences and alignments n Infer function from annotations of matched proteins. n Analysis by range of existing programs. n Interpret results. Additional factors: n some programs/results available over WWW/email; n continual updates of primary databases -- need for reassessment.
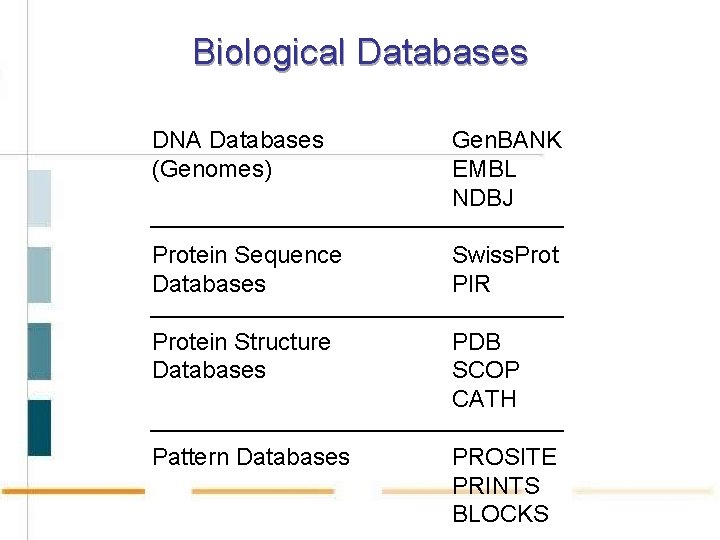
Biological Databases DNA Databases (Genomes) Gen. BANK EMBL NDBJ Protein Sequence Databases Swiss. Prot PIR Protein Structure Databases PDB SCOP CATH Pattern Databases PROSITE PRINTS BLOCKS

Swiss. Prot Entry ID PRIO_BOVIN STANDARD; PRT; 264 AA. AC P 10279; DT 01 -MAR-1989 (Rel. 10, Created) DT 01 -NOV-1991 (Rel. 20, Last sequence update) DT 15 -JUL-1998 (Rel. 36, Last annotation update) DE MAJOR PRION PROTEIN 1 PRECURSOR (PRP) (MAJOR SCRAPIE-ASSOCIATED FIBRIL DE PROTEIN 1). GN PRNP. OS Bos taurus (Bovine). OC Eukaryota; Metazoa; Chordata; Craniata; Vertebrata; Mammalia; . . . CC -!- FUNCTION: THE FUNCTION OF PRP IS NOT KNOWN. PRP IS ENCODED IN THE CC HOST GENOME AND IS EXPRESSED BOTH IN NORMAL AND INFECTED CELLS. CC -!- SUBUNIT: PRP HAS A TENDENCY TO AGGREGATE YIELDING POLYMERS CALLED CC "RODS". CC -!- SUBCELLULAR LOCATION: ATTACHED TO THE MEMBRANE BY A GPI-ANCHOR. CC ------------------------------------CC This SWISS-PROT entry is copyright. It is produced through a collaboration CC between the Swiss Institute of Bioinformatics and the EMBL outstation. . . SQ SEQUENCE 264 AA; 28614 MW; DEA 01 B 4 E CRC 32; MVKSHIGSWI LVLFVAMWSD VGLCKKRPKP GGGWNTGGSR YPGQGSPGGN RYPPQGGGGW GQPHGGGWGQPH GGGWGQPHGGGG WGQGGTHGQW NKPSKPKTNM KHVAGAAAAG AVVGGLGGYM LGSAMSRPLI HFGSDYEDRY YRENMHRYPN QVYYRPVDQY
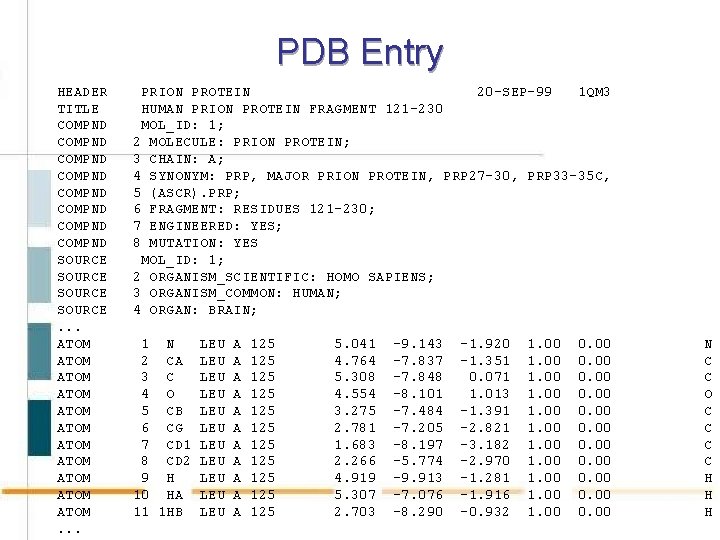
PDB Entry HEADER TITLE COMPND COMPND SOURCE. . . ATOM ATOM ATOM. . . PRION PROTEIN 20 -SEP-99 1 QM 3 HUMAN PRION PROTEIN FRAGMENT 121 -230 MOL_ID: 1; 2 MOLECULE: PRION PROTEIN; 3 CHAIN: A; 4 SYNONYM: PRP, MAJOR PRION PROTEIN, PRP 27 -30, PRP 33 -35 C, 5 (ASCR). PRP; 6 FRAGMENT: RESIDUES 121 -230; 7 ENGINEERED: YES; 8 MUTATION: YES MOL_ID: 1; 2 ORGANISM_SCIENTIFIC: HOMO SAPIENS; 3 ORGANISM_COMMON: HUMAN; 4 ORGAN: BRAIN; 1 N 2 CA 3 C 4 O 5 CB 6 CG 7 CD 1 8 CD 2 9 H 10 HA 11 1 HB LEU LEU LEU A A A 125 125 125 5. 041 4. 764 5. 308 4. 554 3. 275 2. 781 1. 683 2. 266 4. 919 5. 307 2. 703 -9. 143 -7. 837 -7. 848 -8. 101 -7. 484 -7. 205 -8. 197 -5. 774 -9. 913 -7. 076 -8. 290 -1. 920 -1. 351 0. 071 1. 013 -1. 391 -2. 821 -3. 182 -2. 970 -1. 281 -1. 916 -0. 932 1. 00 1. 00 0. 00 N C C O C C H H H
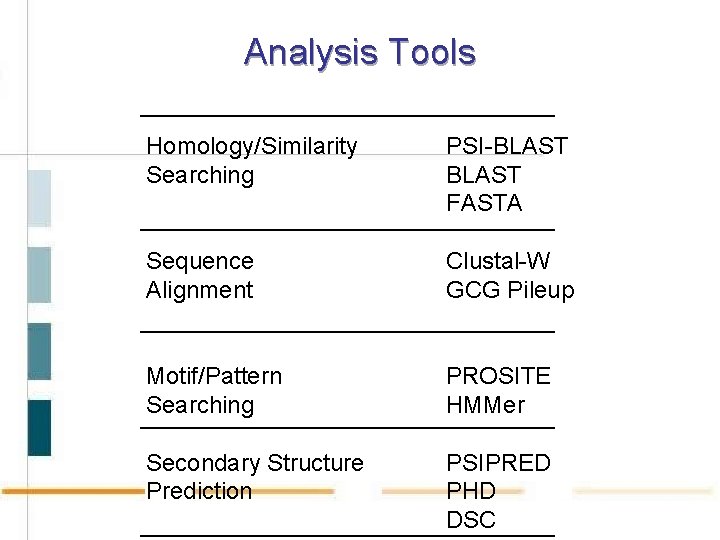
Analysis Tools Homology/Similarity Searching PSI-BLAST FASTA Sequence Alignment Clustal-W GCG Pileup Motif/Pattern Searching PROSITE HMMer Secondary Structure Prediction PSIPRED PHD DSC

BLAST Output. . . Database: pdb_seq 14, 442 sequences; 3, 011, 261 total letters Searching. . . done Sequences producing significant alignments: Score (bits) pdb|1 NBD|1 cftrfragment: nbd 1, first (or n-terminal) nucleotide-. . . pdb|1 WAI|1 DNA polymerase(t 4 gp 43)DNA substrate (tttt)DNA … 79 28 >pdb|1 NBD|1 cftrfragment: nbd 1, first (or n-terminal) nucleotide-binding domain; (cftr nbd 1, cystic fibrosis transmembrane conductance regulator nucleotide-binding domain 1) Length = 214 Score = 78. 8 bits (191), Expect = 6 e-16 Identities = 37/40 (92%), Positives = 39/40 (97%) Query: 4 TTLLVTSKMEHLKKADKILILHEGSSYFYGTFSELQNLRP 43 T +LVTSKMEHLKKADKILILHEGSSYFYGTFSELQNL+P Sbjct: 175 TRILVTSKMEHLKKADKILILHEGSSYFYGTFSELQNLQP 214. . . E Valu 6 e-1 1. 0

Alignment Output *>>>TRANSGELIN : TRANSGELIN SEQUENCE *P 1; A 60598 : actin-associated protein p 27 - mouse *>>>SM 22_RAT : SMOOTH MUSCLE PROTEIN 22 -ALPHA (SM 22 -ALPHA). . MANKGPSYGMSREVQSKIEKKYDEELEERLVEWIIVQCGPDVGRPDRGRLGFQVWLKNGVILSKLVNSLYPDGSKPVKVP MANKGPSYGMSREVQSKIEKKYDEELEERLVEWIVVQCGPDVGAPDRGRLGFQVWLKNGVILSKLVNSLYPEGSKPVKVP ANKGPSYGMSREVQSKIEKKYDEELEERLVEWIVMQCGPDVGRPDRGRLGFQVWLKNGVILSKLVNSLYPEGSKPVKVP MANKGPSYGMSREVQSKIEKKYDEELEERLVEWIVMQCGPDVGRPDRGRLGFQVWLKNGVILSKLVNSLYPEGSKPVKVP ANKGPAYGMSRDVQSKIEKKYDDELEDRLVEWIVAQCGSSVGRPDRGRLGFQVWLKNGIVLSQLVNSLYPDGSKPVKIP MANKGPAYGMSRDVQSKIEKKYDDELEDRLVEWIVAQCGSSVGRPDRGRLGFQVWLKNGIVLSQLVNSLYPDGSKPVKIP ANKGPSYGMSREVQSKIEKKYDEELEERLVEWIIVQCGPDVGRPDRGPLGFQVWLKNGVILSKLVNSLYPDGSKPVKVP MANKGPSYGMSREVQSKIEKKYDEELEERLVEWIIVQCGPDVGRPDRGRLGFQVWLKNGVILSKLVNSLYPEGSKPVKVP MANRGPAYGLSREVQQKIEKQYDADLEQILIQWITTQCRKDVGRPQPGRENFQNWLKDGTVLCELINALYPEGQAPVKKI MANRGPSYGLSREVQEKIEQKYDADLENKLVDWIILQCAEDIEHPPPGRTHFQKWLMDGTVLCKLINSLYPPGQEPIPKI MSLERAVRAKIAGKRNPEMDKEAQEWIEAIIAEKFPAGQS YEDVLKDGQVLCKLINVLSPNA VPKV EFPPSGLSYQVKKKLEGKRDKDQENEALEWIEALTGLKLDRSKL YEDILKDGTVLCKLMNSIKPGC IKKI MELWRQCTHWLIQCRVLPPSHRVTWDGAQVCELAQALRDGVLLCQLLNNLLPHAINLREVN MELWRQCTHWLIQCRVLPPSHRVTWEGAQVCELAQALRDGVLLCQLLNNLLPQAINLREVN MSMEGISYTNSNPSATPNMEDTLLTFSMGILPITMDCDPVTQLSQLFQQGAPLCILFNSVKPQF KLP ENPPSMVFKQMEQVAQFLKAA. . . EDYGVTKTDMFQTVDLFEGKDMAAVQRTVMALGSLAVTKNDGHYRGDPNWFMKKAQEH EDYGVIKTDMFQTVDLYEGKDMAAVQRTLMALGSLAVTKNDGNYRGDPNWFMKKAQEH EDYGVTKTDMFQTVDLFEGKDMAAVQRTVMALGSLAVTKNDGHYRGDPNWFMKKAQEH
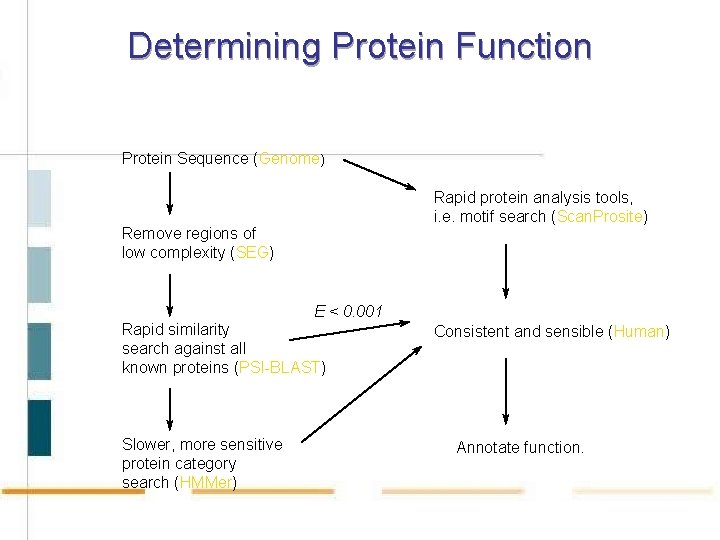
Determining Protein Function Protein Sequence (Genome) Rapid protein analysis tools, i. e. motif search (Scan. Prosite) Remove regions of low complexity (SEG) E < 0. 001 Rapid similarity search against all known proteins (PSI-BLAST) Slower, more sensitive protein category search (HMMer) Consistent and sensible (Human) Annotate function.

Agent Classes Primary database agents manage remote primary sequence databases, providing up-to-date data in various common formats. Non-redundant database agents filter and combine data from various primary database agents into non-redundant data sources. Calculation agents encapsulate pre-existing methods or tools for the analysis of data to determine function. Genome agents manage genome information for a particular organism and use other agents to derive annotations. Broker agents provide information about agents registered within the agent community.
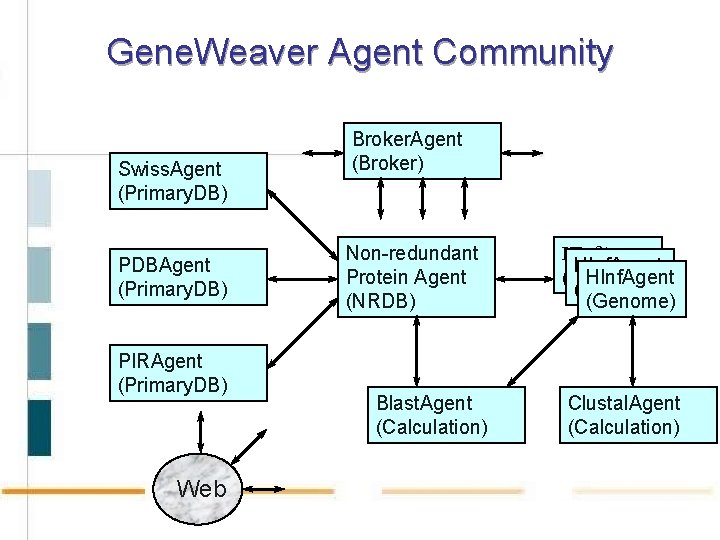
Gene. Weaver Agent Community Swiss. Agent (Primary. DB) PDBAgent (Primary. DB) PIRAgent (Primary. DB) Web Broker. Agent (Broker) Non-redundant Protein Agent (NRDB) Blast. Agent (Calculation) HInf. Agent (Genome) Clustal. Agent (Calculation)

BAL Performatives register Register with a broker. unregister Cancel a registration with a broker. ask Ask about data. derive Request an agent to derive particular data. tell Inform another agent about data. deny Inform another agent about lack of data. subscribe Obtain regular updates of certain data. unsubscribe Stop receiving regular updates of data. ok Indicates success. sorry Indicates failure on the agent’s part. error Indicates problem with protocol or other error.
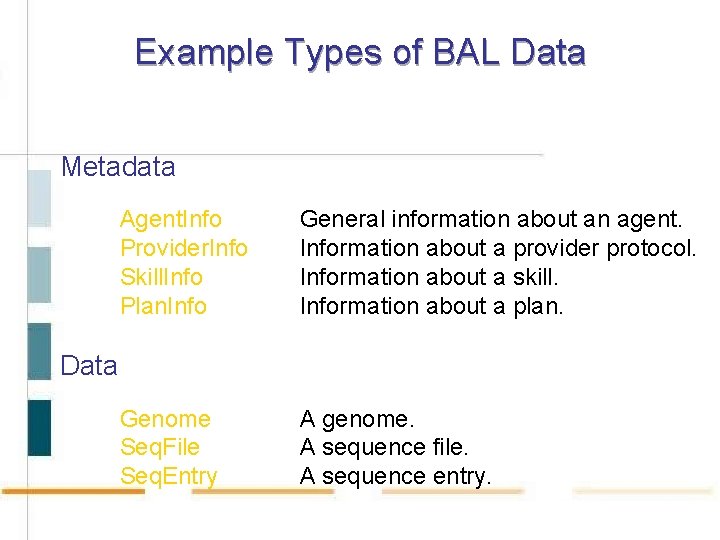
Example Types of BAL Data Metadata Agent. Info Provider. Info Skill. Info Plan. Info General information about an agent. Information about a provider protocol. Information about a skill. Information about a plan. Genome Seq. File Seq. Entry A genome. A sequence file. A sequence entry. Data

BAL Message Example Sender: //localhost. localdomain/7/HInf Receiver: //localhost. localdomain/0/Broker Transport: rmi Language: bal Perform: register Ref: hinf 77 f 001_0 Content: Agent. Info( TYPE = Genome, OWNER = hinf 77 f 001, UPD_TIME = 962601420367, MOD_TIME = 962601420367, ID = HInf, DESCRIPTION = "H. Influenzae Genome Agent")
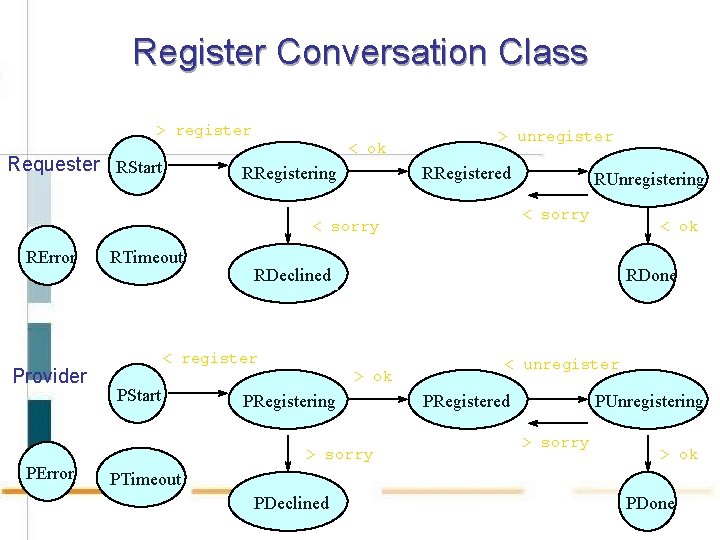
Register Conversation Class > register Requester RStart < ok RRegistering > unregister RRegistered < sorry RError Provider PError RTimeout < ok RDeclined < register PStart RUnregistering RDone > ok PRegistering > sorry < unregister PRegistered PUnregistering > sorry > ok PTimeout PDeclined PDone
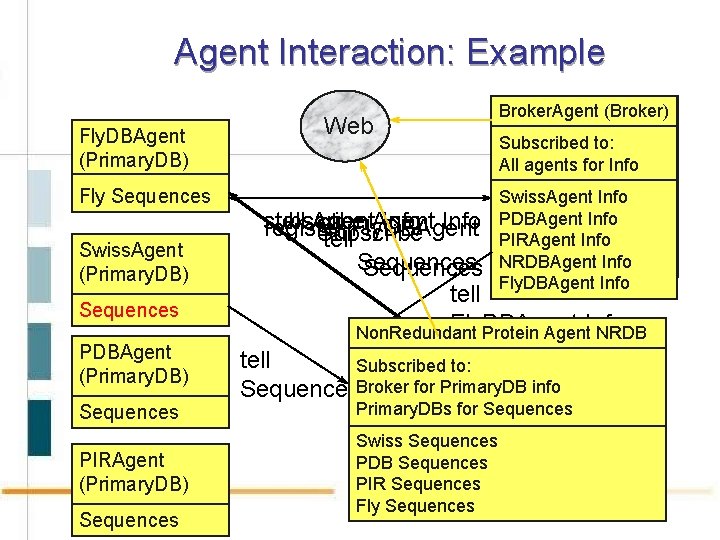
Agent Interaction: Example Fly. DBAgent (Primary. DB) Fly Sequences Swiss. Agent (Primary. DB) Sequences PDBAgent (Primary. DB) Sequences PIRAgent (Primary. DB) Sequences Web Broker. Agent (Broker) Subscribed to: All agents for Info Swiss. Agent Info PDBAgent Info PIRAgent Info NRDBAgent Info Fly. DBAgent Info subscribe tell Agent Info ok Fly. DBAgent register Subscribe tell Sequences tell Fly. DBAgent Non. Redundant Protein Agent. Info NRDB Non. Redundant Protein Agent NRDB tell Subscribed to: Broker for Primary. DB info Sequences. Subscribed Broker for Primary. DB info Primary. DBs for Sequences Swiss Sequences PDB Sequences PIR Sequences Fly Sequences

Goals Higher Level Goals Derive. Goal Agent should try to derive data with particular properties. Update. Goal Agent should try to update data matching a given template. Relation. Goal Agent should attempt to establish the given type of relationship with another agent. Lower Level Goals Do. Goal, Query. Goal, Tell. Goal.
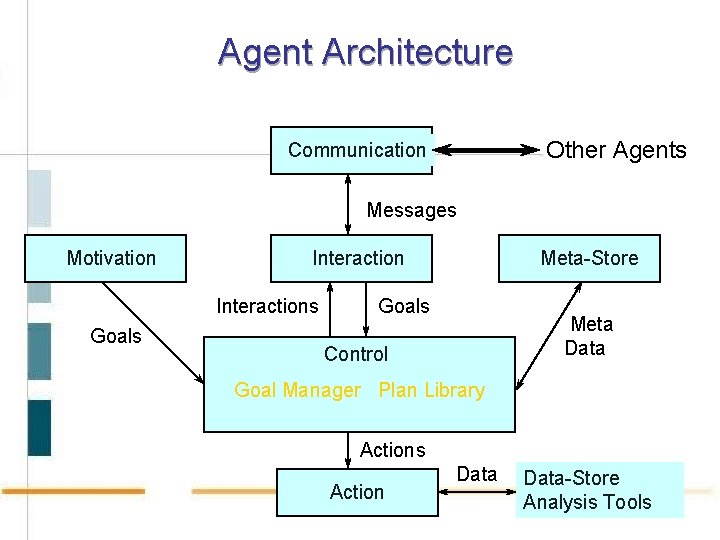
Agent Architecture Other Agents Communication Messages Motivation Interactions Goals Meta-Store Goals Meta Data Control Goal Manager Plan Library Actions Action Data-Store Analysis Tools

Annotate Function Example Genome Agent 1 In response to a higher level motivation, Derive. Goal(Seq. Function) is created to annotate any sequences with annotated function confidence < 0. 5. 2 Using a plan from the plan library, Derive. Goal(Seq. Function) is decomposed into Relational. Goal(derive), Derive. Goal(Homologue) and function assignment using the homologue if confidence > 0. 5. 3 A suitable agent with Derive. Provider and ‘homology’ skill is located. 4 Derive requester interaction used to accomplish Relation. Goal(derive). Blast Agent 5 Skill used to satisfy Do. Goal(Homologue).

Summary n Applications: bioinformatics n n Tensions between biological sciences and computer science Work remaining n n n problem not created by the technologies used to solve it practical developments to inform conceptual infrastructure Consolidation of existing prototype Inclusion of multiple calculation agents Evaluation of implementation infrastructure Staged and full deployment Future work: agent marketplace with calculation agents competing?
- Slides: 23