Gene Ontology Biological process Cellular process Cellular physiolgical



Gene Ontology





Структура иерархии: сеть Biological process • Cellular process – Cellular physiolgical process • Cell division – Asymmetric cell division » Regulation of asymmetric cell division – Regulation of cell division » Regulation of asymmetric cell division • Regulation of cellular physiological process – Regulation of cell division » Regulation of assymmetric cell division • Physiological process – Cellular physiolocical process • … – Regulation of physiological process • …

Упражнение Нарисовать пути, ведущие к: (А-Д) (Е-К) (Л-Н) (О-П) (Р-С) (Т-Я) GO: 0045782 : positive regulation of cell budding GO: 0004612 : phosphoenolpyruvate carboxykinase (ATP) activity GO: 0019568 : arabinose catabolism GO: 0003726 : double-stranded RNA adenosine deaminase activity GO: 0030660 : Golgi vesicle membrane GO: 0030570 : pectate lyase activity GO: 0019319 : hexose biosynthesis GO: 0047689 : aspartate racemase activity GO: 0006068 : ethanol catabolism GO: 0004129 : cytochrome-c oxidase activity GO: 0030334 : regulation of cell migration GO: 0003705 : RNA polymerase II transcription factor activity, enhancer binding используя Ami. GO http: //www. geneontology. org Ami. Go http: //www. godatabase. org/cgi-bin/amigo/go. cgi? search_constraint=terms&action=replace_tree&session_id=7922 b 11 25244220
BLAST home page


Similarity ≠ homology • BLAST e-value is a measure of nonrandomness of sequence similarity • Possible causes of similarity: – homology – domain homology – low complexity, coiled-coil, transmembrane and other types of regions with non-standard amino acid composition • Homology ≠ same function. Normally: – similar (general) function (e. g. enzymatic activity) – maybe different specificity
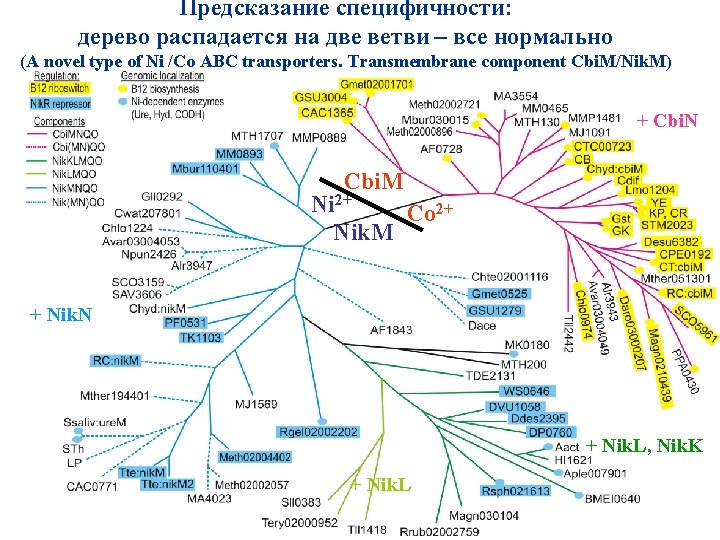
Предсказание специфичности: дерево распадается на две ветви – все нормально (A novel type of Ni /Co ABC transporters. Transmembrane component Cbi. M/Nik. M) + Cbi. N Cbi. M Ni 2+ Co 2+ Nik. M + Nik. N + Nik. L, Nik. K + Nik. L
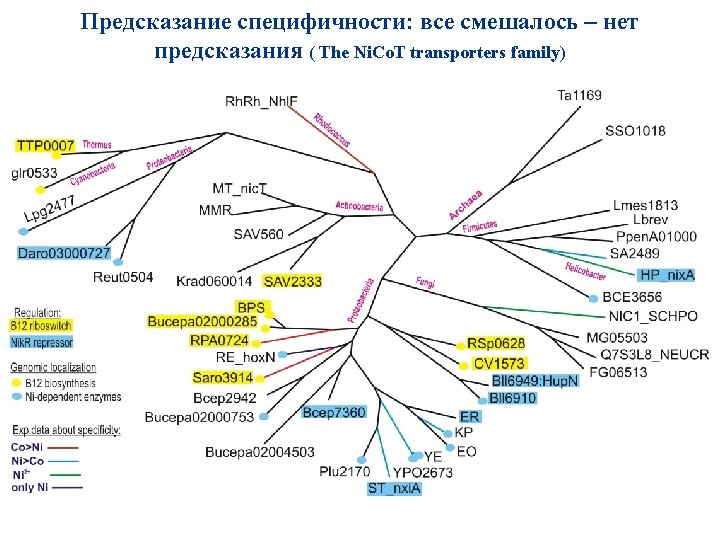
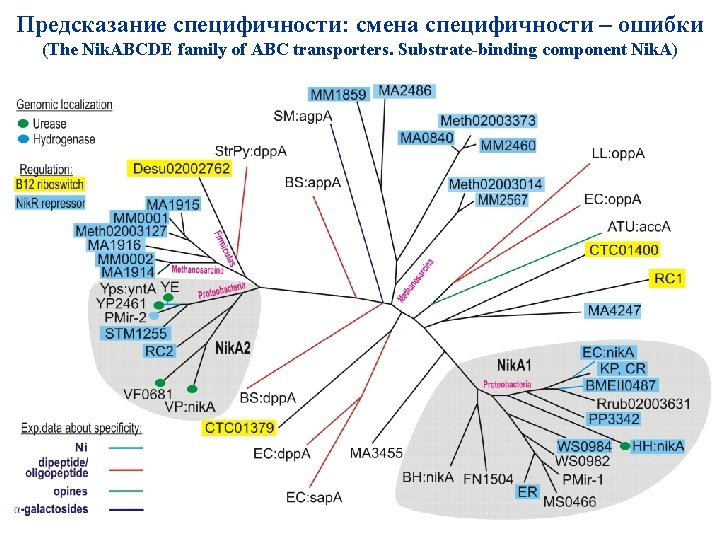
Предсказание специфичности: смена специфичности – ошибки (The Nik. ABCDE family of ABC transporters. Substrate-binding component Nik. A)

Noradrenaline transporter in an archaeon? SOURCE ORGANISM FEATURES source Protein Methanococcus jannaschii Archaea; Euryarchaeota; Methanococcales; Methanococcaceae; Methanococcus. Location/Qualifiers 1. . 492 /organism=" Methanococcus jannaschii " /db_ xref="taxon: 2190" 1. . 492 /product="sodium-dependent noradrenaline transporter" CDS 1. . 492 /gene="MJ 1319" /note="similar to EGAD: HI 0736 percent identity: 38. 5; identified by sequence similarity; putative" /coded_by="U 67572: 71. . 1549" /transl_table=11 Now corrected: Hypothetical sodium-dependent transporter MJ 1319.

Lesson(s) 1. Avoid overprediction (homology does not necessarily mean same cellular role or specificity)
Similarity to hypothetical proteins: somebody else’s errors… The only correct annotation!

Genes with curious functional assignments • C 75604: Probable head morphogenesis protein, Deinococcus radiodurans • O 05360: Automembrane protein H, Yersinia enterocolitica • Q 8 TID 9: Benzodiazepine (valium) receptor Tsp. O, Methanosarcina acetivorans • NP_069403: DR-beta chain MHC class II, Archaeoglobus fulgidus

Errors in experimental papers Swiss. Prot: DEFINITION Hypothetical 43. 6 k. Da protein. ACCESSION. . . KEYWORDS SOURCE ORGANISM P 48012 Hypothetical protein. Debaryomyces occidentalis Eukaryota; Fungi; Ascomycota; Saccharomycotina; Saccharomycetes; Saccharomycetales ; Saccharomycetaceae; Debaryomyces. [CAUTION] Was originally (Ref. 1) thought to be 3 -isopropylmalate dehydrogenase (LEU 2). PIR: DEFINITION 3 -isopropylmalate dehydrogenase ACCESSION KEYWORDS - yeast(Schwanniomyces occidentalis ). S 55845 oxidoreductase. (EC 1. 1. 1. 85)

Swiss. Prot entry DSDX_ECOLI -!- CAUTION: An ORF called dsd. C was originally (Ref. 3) assigned to the wrong DNA strand thought to be a D-serine deaminase activator, it was then resequenced by Ref. 2 and still thought to be "dsd. C", but this time to function as a D -serine permease. It is Ref. 1 that showed that dsd. C is another gene and that this sequence should be called dsd. X. It should also be noted that the C-terminal part of dsd. X (from 338 onward) was also sequenced (Ref. 6 and Ref. 7) and was thought to be a separate ORF (don't worry, we also had difficulties understanding what happened!).

Lesson(s) 1. Avoid overprediction (homology does not necessarily mean same cellular role or specificity) 2. Check carefully the source(s) of annotations in the list of homologs

mastermind protein of Drosophila

Filtering of low-complexity segments • often insufficient • may lose non-trivial information

Lesson(s) 1. Avoid overprediction (homology does not necessarily mean same cellular role or specificity) 2. Check the source(s) of annotations in the list of homologs 3. Beware of similarity in low-complexity regions, non-globular domains, transmembrane segments
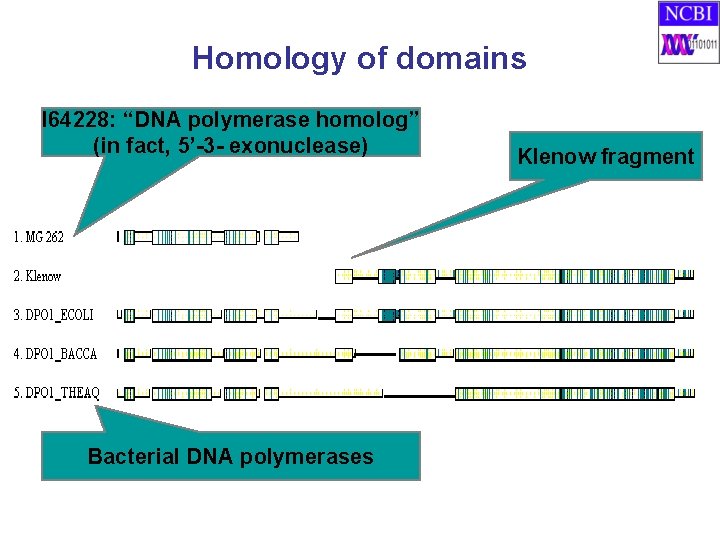
Homology of domains I 64228: “DNA polymerase homolog” (in fact, 5’-3 - exonuclease) Bacterial DNA polymerases Klenow fragment

BLAST domains page
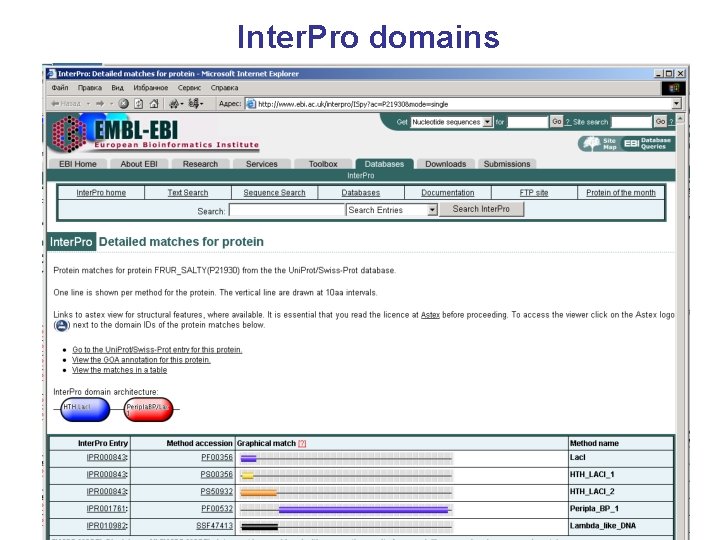
Inter. Pro domains

Lesson(s) 1. Avoid overprediction (homology does not necessarily mean same cellular role or specificity) 2. Check the source(s) of annotations in the list of homologs 3. Beware of similarity in low-complexity regions, non-globular domains, transmembrane segments 4. Do not extend domain homology to annotation of the whole protein


caspases/paracaspases/metacaspases


Lesson(s) 1. Avoid overprediction (homology does not necessarily mean same cellular role or specificity) 2. Check the source(s) of annotations in the list of homologs 3. Beware of similarity in low-complexity regions, non-globular domains, transmembrane segments 4. Do not extend domain homology to annotation of the whole protein 5. Правильный паттерн должен сохраняться у (близких) ортологов; должны сохраняться основные каталитические остатки

- Slides: 35