Gene Finding in Eukaryotic Genomes Center for Biologisk

”Gene Finding in Eukaryotic Genomes” Center for Biologisk Sekvensanalyse DTU course #27803 Fall 2003 Nikolaj Blom Center for Biological Sequence Analysis Bio. Centrum. DTU Technical University of Denmark nikob@cbs. dtu. dk

Center for Biologisk Sekvensanalyse Human Genome Published HUGO: Nature, 15. feb. 2001 Celera: Science, 16. feb. 2001

Center for Biologisk Sekvensanalyse We Have the Human Genome Sequence. . . now what? So, what is the problem? • Well. . . • We don’t know how many genes there are! • We don’t know where they are! • We don’t know what they do!
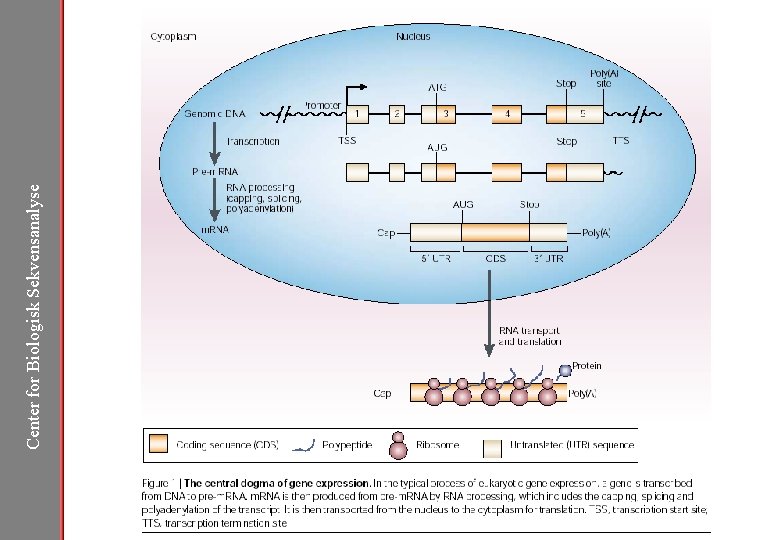
Center for Biologisk Sekvensanalyse
Center for Biologisk Sekvensanalyse The cellular machinery recognize genes without access to Gen. Bank, Swiss. Prot or computers – can we?

Center for Biologisk Sekvensanalyse Needles Hiding in Genome Haystacks. . . Genes are embedded in the genome sequence Coding regions constitute only 2% of human genome Can we distinguish the gene features from the background?

Can U spot ’Spot’? Center for Biologisk Sekvensanalyse

Center for Biologisk Sekvensanalyse TTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTATGCTGAACAGGCCAGAGAATTCATCTAAATAGCCTAAGCAGGCTGGG TGGCTCACCTGTAATCCCAACACTTGGGAGGCCGAGGTGGGCAGATCACCTGAGGTCAGGAGTTCAAGGCTAGCCAACATGACAAAACC TCTACTAAAAAAATACAAAAATTAGCCAGGCATAGTGGCGCCTATAGTTCCAGCTACTTGGGGGCTGAGGTAGGAAGATCGCTAGAGCCTGGGA AGGCTGCGGTGAGCTGTGATTGTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTT TTTTTTTTTTTTTTTTTTTTTTTTTTTTTGGACTCTTCTGTTTGATGGTGGTCTTCCTCATCCTCT ATGTGAAGCTCCATGGAGATCACCTACCCATACCTGCTTCTGTGACCTCATTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTG CCTGGTGTTGGAATGTGCCAAGGTTTGCCATTAAACACACATTTCTCATTTCATATATATATATGTGTGTGTT ATGCGTGTGTGTTATATAAAATATATAGGAAGAGGCACCAGAGAGCTCTCTGCATAGTCACAGAGGAAAGGTCATGTGAGG CAGAAGGCAGATGTCACAAGCCTCACCAGCAACCTACCATACCCTGCTTGTACCTCCATCCTGGAAGTCCAGCTTCTAAAATTAGAAGAAAATA GTGTGTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTGTGGCTCGCACCTATAATCCCAGCACTTTGGGAGGCTGATGT GATCATTTGAGGTCAAGAGTTTGAAACCAGCCTAGGCAACATAGGGAGACCCTGTCTTTAAAATTTTTGTTTTAATTAGCTGGGTG GTGCACACCTGAGTCCTAGCTACTTGGGAGGCTGAGGTAGGAGGATCCCCTGAGCCCAGGGAAGTGGAGGCTGCAGTGAGCCATGATCACACCT TTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTT TTTTTTTTTTTTTTTTTTCCTTGTCAGGTTTTCACCCCATGCTCCTCCATTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTGGGCTAGTCTGCTCTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTGCTTCCCGTCTTACTGGAAGA AGCATTTGACAGAGTTGGTCACTCTCTCCTCCTTGGACACCTTTTCTTCACTTGGTTTCCAGAACAGCATTATCTCCTGCTTATTGTCTTCCTC

Center for Biologisk Sekvensanalyse

Center for Biologisk Sekvensanalyse TAATTAAAGCTAAATGAAGTTGTAAGAGTGGCCCTATCGCATAGGACTAGTGTCCCTATAAGAACACGAAGAAATCACCTTAGAAAGGCTGAG GCTGCAGGGCAGTGGGAGTGCAGACTGAAAGATGCAGACCACTGGGCTTCTACTTCTGTTTCCATTTCTGATCCGGCCTGCATCTGCCTCCTTC CAGGCCAGAGAATTCATCTAAATAGCCTAAGCAGGCTGGGTGCTGTGGCTCACCTGTAATCCCAACACTTGGGAGGCCGAGGTGGGCAGATCAC GTCAGGAGTTCAAGGCTAGCCAACATGACAAAACCCCATCTCTACTAAAAAAATACAAAAATTAGCCAGGCATAGTGGCGCCTATAGTT TACTTGGGGGCTGAGGTAGGAAGATCGCTAGAGCCTGGGAGGTTAAGGCTGCGGTGAGCTGTGATTGTGCCACTGCACTCCAGCCTGGGTGACA AGACCCTGCCTCAAAAATAAATAAATAAAAATAAGAGTGCTTGGCAGCTTGATCAAGCTATGCCAGGAACCCATCTCTCAAG GCTCTTCTCCTGTGCCATTGTCAGCTTTGTCCTGTCTGAGTCCATGGGACTCTTCTGTTTGATGGTGGTCTTCCTCATCCTCTTCATCATGTGA CATGGAGATCACCTACCCATACCTGCTTCTGTGACCTCATGCCATTCCTGGTGTTGGAATGTGCCAAGGTTTGCCATTAAACACACATTTCTCA TAATTTCATATTATATGTGTGTGTTTATATATGCGTGTGTGTGTGTGTGTATATATATATAAAATATATAGGAAGAGGCACCAGAGAGCTCTCTGCATAGTCACAGAGGAAAGGTCATGTGAGGACAGCCA CAGATGTCACAAGCCTCACCAGCAACCTACCATACCCTGCTTGTACCTCCATCCTGGAAGTCCAGCTTCTAAAATTAGAAGAAAATAGTCGGGT GGCTCGCACCTATAATCCCAGCACTTTGGGAGGCTGATGTGGGAGGATCATTTGAGGTCAAGAGTTTGAAACCAGCCTAGGCAACATAGGGAGA TCTTTAAAATTTTTGTTTTAATTAGCTGGGTGTGATGGTGCACACCTGAGTCCTAGCTACTTGGGAGGCTGAGGTAGGAGGATCCC CCCAGGGAAGTGGAGGCTGCAGTGAGCCATGATCACACCACTGCAATACAGCCTGGGTGACAGAGCAAGACCTTATCTCAAAATAAACAA AAAGATGACAAAATGTCTGTCGTTTAAGTCACCCATTCTGTGATATCTTGTTACGGCAGCCTGAACTGACCAATACACTTCCTCACCCAG ATTCCATGCTCAATCATAATCAGCCATTGCAATTACCCTCAACTGTATTATCAACCCTCAATTTGTATTAGTTGCTTGGCAAAACCCT AATCCAGTTCTTCTATATCTACATCGATGCTGCCGAATATGGCTGAAGAAAAGCAACTGTGTTGACTGCTTTAAATTCATGACCACTTA AGTGGGCACTTAACTTCCTGGCAATTATTCTACATTTTTCTAGTCCATTAACTCTCCTCCTCTCTGAGTTAATTATTTCACAGCTTTTCCTCCC ATACATGTTCCATCCTAACTCTCTGCTGATGACCTTGTTTCTTATTTCACTAATGGAGGCCACCAGGAGAGAACTCCCACAGCCATCAAATTCA CCAACAGCATCCTTACACAAATCCTCTGCCTTCTCTCTGGGCTGTGCCCTCTCTTTGCTCCTGCAATTTCCCTAACTCTCCTATACTGTT TCACTCTCCAGTGGATAATCACCATCAGGATGCAAAGATGCTGTACTAGCTTCTGAACTCTCCAAAAACCCAGGAAACAAAAAGGCTA TTTCTTATTCCCCCTTATATACATATATAGTAGGCACTCAATAAACATTCACTGAATGAACAGTAATGCTCACTTGCCCATAAA GTACCTCATCTTTTACCACAAAGGGTATTTGTAAATATTTAGGTTGTTTCTACCCAGATTATGGCTTGGTAATTCTTTTTTTCTAATT TTTTTTTCTAGGGACAGGGTCTCACTATGTTGCCCAGGATGGTCTTGAACTCCTGGGCTCAAGCATTCTGCCTTGGCCTCCTAAAGTGCT TACAGGCATGAGCCACCGTGCCTTCATGTTTTTAGAACACAGAGAAAATGTGTTCTAAATGTGCTCATTGCTCAGCAATGAGCAAA ATGCAGTCACCACCAATCAAAAACTTTTTTTTGAGACAAGATCTTGCTCTGTTGCCCAGGCTGGAGTGCAGTGGCAGGATCATAGC GCAGTCTTGACCTCATAGGCCTAAATCATCCTCCCACCTCAGCCTCACAAGTAGCTAAGACCACAGGTACAAGCCACCGTATCTAGCTAACTTT TTTTTTGAATTTTTAAAAATTTTGAGGCCAGGCTGGCCTCAAACTCCTGAGCTCAAGCAATCCTCCCACCTTGGCTTCCCAAAGTGCT TATAGGCGTGAGCAACTGTACCTGGCAAAAACTTTTTAAGAGCTTCCAGATTTAGGTTGTTTCTACCCAGATTATGGCTTGGTAATTCT TTTTTTTCTAATTTTTTTTCTAGGGACAGGGTCTCACTATGTTGCCCAGGATGGTCTTGAACTCCTGGGCTCAAGCATTCTGCC CTCCTAAAGTGCTGAGATTACAGGCATGAGCCACCGTGCCTTCATGTTTTTAGAACACAGAGAAAATGTGTTCTAAATGTGCTCAT AGCAATGAGCAAAGGCTTATGCAGTCACCACCAATCAAAAACTTTTTTTTGAGACAAGATCTTGCTCTGTTGCCCAGGCTGGAGTG GCAGGATCATAGCAAGCTGCAGTCTTGACCTCATAGGCCTAAATCATCCTCCCACCTCAGCCTCACAAGTAGCTAAGACCACAGGTACAAGCCA Can. UUspotthe the. Gene? Gin? Can Ooops

Center for Biologisk Sekvensanalyse TAATTAAAGCTAAATGAAGTTGTAAGAGTGGCCCTATCGCATAGGACTAGTGTCCCTATAAGAACACGAAGAAATCACCTTAGAAAGGCTGAG GCTGCAGGGCAGTGGGAGTGCAGACTGAAAGATGCAGACCACTGGGCTTCTACTTCTGTTTCCATTTCTGATCCGGCCTGCATCTGCCTCCTTC CAGGCCAGAGAATTCATCTAAATAGCCTAAGCAGGCTGGGTGCTGTGGCTCACCTGTAATCCCAACACTTGGGAGGCCGAGGTGGGCAGATCAC GTCAGGAGTTCAAGGCTAGCCAACATGACAAAACCCCATCTCTACTAAAAAAATACAAAAATTAGCCAGGCATAGTGGCGCCTATAGTT TACTTGGGGGCTGAGGTAGGAAGATCGCTAGAGCCTGGGAGGTTAAGGCTGCGGTGAGCTGTGATTGTGCCACTGCACTCCAGCCTGGGTGACA AGACCCTGCCTCAAAAATAAATAAATAAAAATAAGAGTGCTTGGCAGCTTGATCAAGCTATGCCAGGAACCCATCTCTCAAG GCTCTTCTCCTGTGCCATTGTCAGCTTTGTCCTGTCTGAGTCCATGGGACTCTTCTGTTTGATGGTGGTCTTCCTCATCCTCTTCATCATGTGA CATGGAGATCACCTACCCATACCTGCTTCTGTGACCTCATGCCATTCCTGGTGTTGGAATGTGCCAAGGTTTGCCATTAAACACACATTTCTCA TAATTTCATATTATATGTGTGTGTTTATATATGCGTGTGTGTGTGTGTGTATATATATATAAAATATATAGGAAGAGGCACCAGAGAGCTCTCTGCATAGTCACAGAGGAAAGGTCATGTGAGGACAGCCA CAGATGTCACAAGCCTCACCAGCAACCTACCATACCCTGCTTGTACCTCCATCCTGGAAGTCCAGCTTCTAAAATTAGAAGAAAATAGTCGGGT GGCTCGCACCTATAATCCCAGCACTTTGGGAGGCTGATGTGGGAGGATCATTTGAGGTCAAGAGTTTGAAACCAGCCTAGGCAACATAGGGAGA TCTTTAAAATTTTTGTTTTAATTAGCTGGGTGTGATGGTGCACACCTGAGTCCTAGCTACTTGGGAGGCTGAGGTAGGAGGATCCC CCCAGGGAAGTGGAGGCTGCAGTGAGCCATGATCACACCACTGCAATACAGCCTGGGTGACAGAGCAAGACCTTATCTCAAAATAAACAA AAAGATGACAAAATGTCTGTCGTTTAAGTCACCCATTCTGTGATATCTTGTTACGGCAGCCTGAACTGACCAATACACTTCCTCACCCAG ATTCCATGCTCAATCATAATCAGCCATTGCAATTACCCTCAACTGTATTATCAACCCTCAATTTGTATTAGTTGCTTGGCAAAACCCT AATCCAGTTCTTCTATATCTACATCGATGCTGCCGAATATGGCTGAAGAAAAGCAACTGTGTTGACTGCTTTAAATTCATGACCACTTA AGTGGGCACTTAACTTCCTGGCAATTATTCTACATTTTTCTAGTCCATTAACTCTCCTCCTCTCTGAGTTAATTATTTCACAGCTTTTCCTCCC ATACATGTTCCATCCTAACTCTCTGCTGATGACCTTGTTTCTTATTTCACTAATGGAGGCCACCAGGAGAGAACTCCCACAGCCATCAAATTCA CCAACAGCATCCTTACACAAATCCTCTGCCTTCTCTCTGGGCTGTGCCCTCTCTTTGCTCCTGCAATTTCCCTAACTCTCCTATACTGTT TCACTCTCCAGTGGATAATCACCATCAGGATGCAAAGATGCTGTACTAGCTTCTGAACTCTCCAAAAACCCAGGAAACAAAAAGGCTA TTTCTTATTCCCCCTTATATACATATATAGTAGGCACTCAATAAACATTCACTGAATGAACAGTAATGCTCACTTGCCCATAAA GTACCTCATCTTTTACCACAAAGGGTATTTGTAAATATTTAGGTTGTTTCTACCCAGATTATGGCTTGGTAATTCTTTTTTTCTAATT TTTTTTTCTAGGGACAGGGTCTCACTATGTTGCCCAGGATGGTCTTGAACTCCTGGGCTCAAGCATTCTGCCTTGGCCTCCTAAAGTGCT TACAGGCATGAGCCACCGTGCCTTCATGTTTTTAGAACACAGAGAAAATGTGTTCTAAATGTGCTCATTGCTCAGCAATGAGCAAA ATGCAGTCACCACCAATCAAAAACTTTTTTTTGAGACAAGATCTTGCTCTGTTGCCCAGGCTGGAGTGCAGTGGCAGGATCATAGC GCAGTCTTGACCTCATAGGCCTAAATCATCCTCCCACCTCAGCCTCACAAGTAGCTAAGACCACAGGTACAAGCCACCGTATCTAGCTAACTTT TTTTTTGAATTTTTAAAAATTTTGAGGCCAGGCTGGCCTCAAACTCCTGAGCTCAAGCAATCCTCCCACCTTGGCTTCCCAAAGTGCT TATAGGCGTGAGCAACTGTACCTGGCAAAAACTTTTTAAGAGCTTCCAGATTTAGGTTGTTTCTACCCAGATTATGGCTTGGTAATTCT TTTTTTTCTAATTTTTTTTCTAGGGACAGGGTCTCACTATGTTGCCCAGGATGGTCTTGAACTCCTGGGCTCAAGCATTCTGCC CTCCTAAAGTGCTGAGATTACAGGCATGAGCCACCGTGCCTTCATGTTTTTAGAACACAGAGAAAATGTGTTCTAAATGTGCTCAT AGCAATGAGCAAAGGCTTATGCAGTCACCACCAATCAAAAACTTTTTTTTGAGACAAGATCTTGCTCTGTTGCCCAGGCTGGAGTG GCAGGATCATAGCAAGCTGCAGTCTTGACCTCATAGGCCTAAATCATCCTCCCACCTCAGCCTCACAAGTAGCTAAGACCACAGGTACAAGCCA

Center for Biologisk Sekvensanalyse Needles Hiding in Genome Haystacks. . . Intron-exon structure of genes • Large introns (average 3365 bp ) • Small exons (average 145 bp) • Long genes (average 27 kb)
![Manual Genefinding Start codon: ATG Stop codons: TAA, TAG, TGA Donor splice site: ^GT[AG]AG Manual Genefinding Start codon: ATG Stop codons: TAA, TAG, TGA Donor splice site: ^GT[AG]AG](http://slidetodoc.com/presentation_image/c94c96acc891c871d8a0968913baa50d/image-13.jpg)
Manual Genefinding Start codon: ATG Stop codons: TAA, TAG, TGA Donor splice site: ^GT[AG]AG Acceptor splice site: [CT]AG^ Center for Biologisk Sekvensanalyse >U 70368 (950 bp) 351 401 451 501 551 601 651 701 751 801 851 901 951 1001 1051 1101 1151 1201 1251 CTCCCTTAGA GTGGTTTCTC GTCTTGTGTG CATAAGGACA TGCTCTGCCC TGCATGTGTA ACTGACGAGA CTACCATGAG AATATCTCTC GGTGAGTGTG TGGTGT TGGTCATTAG AGTAATAACA TTATGTTCAT TGGGGAGGGG CATTTCCCTG GGTAAAARAT CCCCTAGCAG GGATGAAGCT AGACTCCAGC TTTCCCCCAGCA GATTGAGTGT TTCTAAGAGG GTGCCAGGAA TGCAATCACT CCACTGTCTA AGGGTAAGAG CAAGTATCTA TGGGAGTCAT GGAGGGGGCA AGACACTGGT GTTTG GTCGGTTAAC AGGTGAAAGT TTGGCCACTG ATCCAGCCCT CAGGTAGTGG AAGTTATTTG AGAAGGTAAG GCCTTGTCAT GCTGGGATAG TTGAGTCTCA AGAGTAGTCA GTGTGGACTT GCAGGGAGGC TTTACAGTAA ATTGGCTAGT AAGCTAAATG GATTATGAAC CTTGTTGGGG GGATGGGTTT TTGGAAAGTA TAGGTCAGGT GCATGCCCTA ATCATCTCCT TGGAGCTAAC AAGAGGTCTT TCTCACTGTA CTCCGGCTGC ACTTTTGTTG TCATAAGGCC TCCCCCAAAA TTTACCAGCT TTTGGGGATG GCAGCA TTTTGTGGCC TTTGCTTTCC CTAGGTTGCA GTATAACCTA TATGTGAGTG ATGCTAGGTA TTTCTAGAAT AAGTCTTTTT GGTGCCCAAC TGCCAATGAC TGGAGACATG AGGTCTTTAT CCTAGACCTG ACAAAGGGGC TTTTGCAGCT CCAGACAGGA AGCTAGGGCA GTGTGCCCCG GAGGGGTGTG TGTAACATAT TCTGCATTGG GATCTGTTGG GAGACTCGAT TTTTCTTTTT CTGTCCTGTT GGAGTCTGAA GTGTTCTTGT AGCTGCATCA AAGCCCAGTC
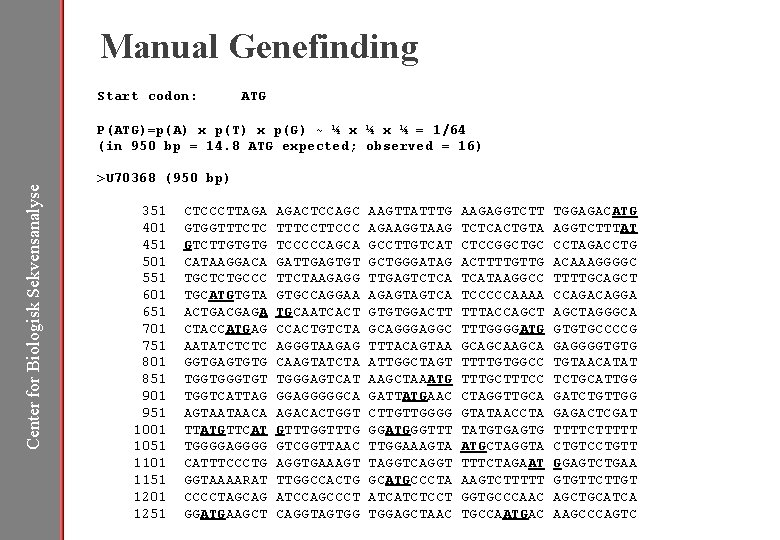
Manual Genefinding Start codon: ATG P(ATG)=p(A) x p(T) x p(G) ~ ¼ x ¼ = 1/64 (in 950 bp = 14. 8 ATG expected; observed = 16) Center for Biologisk Sekvensanalyse >U 70368 (950 bp) 351 401 451 501 551 601 651 701 751 801 851 901 951 1001 1051 1101 1151 1201 1251 CTCCCTTAGA GTGGTTTCTC GTCTTGTGTG CATAAGGACA TGCTCTGCCC TGCATGTGTA ACTGACGAGA CTACCATGAG AATATCTCTC GGTGAGTGTG TGGTGT TGGTCATTAG AGTAATAACA TTATGTTCAT TGGGGAGGGG CATTTCCCTG GGTAAAARAT CCCCTAGCAG GGATGAAGCT AGACTCCAGC TTTCCCCCAGCA GATTGAGTGT TTCTAAGAGG GTGCCAGGAA TGCAATCACT CCACTGTCTA AGGGTAAGAG CAAGTATCTA TGGGAGTCAT GGAGGGGGCA AGACACTGGT GTTTG GTCGGTTAAC AGGTGAAAGT TTGGCCACTG ATCCAGCCCT CAGGTAGTGG AAGTTATTTG AGAAGGTAAG GCCTTGTCAT GCTGGGATAG TTGAGTCTCA AGAGTAGTCA GTGTGGACTT GCAGGGAGGC TTTACAGTAA ATTGGCTAGT AAGCTAAATG GATTATGAAC CTTGTTGGGG GGATGGGTTT TTGGAAAGTA TAGGTCAGGT GCATGCCCTA ATCATCTCCT TGGAGCTAAC AAGAGGTCTT TCTCACTGTA CTCCGGCTGC ACTTTTGTTG TCATAAGGCC TCCCCCAAAA TTTACCAGCT TTTGGGGATG GCAGCA TTTTGTGGCC TTTGCTTTCC CTAGGTTGCA GTATAACCTA TATGTGAGTG ATGCTAGGTA TTTCTAGAAT AAGTCTTTTT GGTGCCCAAC TGCCAATGAC TGGAGACATG AGGTCTTTAT CCTAGACCTG ACAAAGGGGC TTTTGCAGCT CCAGACAGGA AGCTAGGGCA GTGTGCCCCG GAGGGGTGTG TGTAACATAT TCTGCATTGG GATCTGTTGG GAGACTCGAT TTTTCTTTTT CTGTCCTGTT GGAGTCTGAA GTGTTCTTGT AGCTGCATCA AAGCCCAGTC
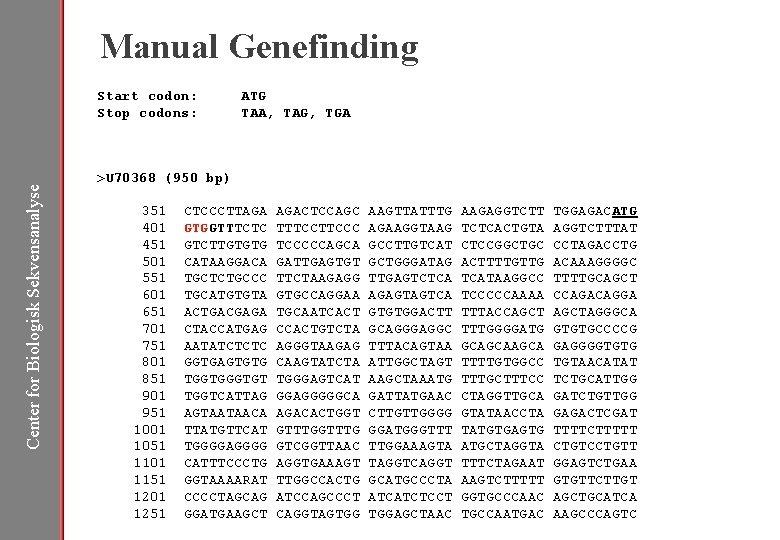
Manual Genefinding Start codon: Stop codons: ATG TAA, TAG, TGA Center for Biologisk Sekvensanalyse >U 70368 (950 bp) 351 401 451 501 551 601 651 701 751 801 851 901 951 1001 1051 1101 1151 1201 1251 CTCCCTTAGA AGACTCCAGC GTGGTTTCTC TTTCCC GTCTTGTGTG TCCCCCAGCA CATAAGGACA GATTGAGTGT TGCTCTGCCC TTCTAAGAGG TGCATGTGTA GTGCCAGGAA ACTGACGAGA TGCAATCACT CTACCATGAG CCACTGTCTA AATATCTCTC AGGGTAAGAG GGTGAGTGTG CAAGTATCTA TGGTGT TGGGAGTCAT TGGTCATTAG GGAGGGGGCA AGTAATAACA AGACACTGGT TTATGTTCAT GTTTG TGGGGAGGGG GTCGGTTAAC CATTTCCCTG AGGTGAAAGT GGTAAAARAT TTGGCCACTG CCCCTAGCAG ATCCAGCCCT GGATGAAGCT CAGGTAGTGG AAGTTATTTG AGAAGGTAAG GCCTTGTCAT GCTGGGATAG TTGAGTCTCA AGAGTAGTCA GTGTGGACTT GCAGGGAGGC TTTACAGTAA ATTGGCTAGT AAGCTAAATG GATTATGAAC CTTGTTGGGG GGATGGGTTT TTGGAAAGTA TAGGTCAGGT GCATGCCCTA ATCATCTCCT TGGAGCTAAC AAGAGGTCTT TCTCACTGTA CTCCGGCTGC ACTTTTGTTG TCATAAGGCC TCCCCCAAAA TTTACCAGCT TTTGGGGATG GCAGCA TTTTGTGGCC TTTGCTTTCC CTAGGTTGCA GTATAACCTA TATGTGAGTG ATGCTAGGTA TTTCTAGAAT AAGTCTTTTT GGTGCCCAAC TGCCAATGAC TGGAGACATG AGGTCTTTAT CCTAGACCTG ACAAAGGGGC TTTTGCAGCT CCAGACAGGA AGCTAGGGCA GTGTGCCCCG GAGGGGTGTG TGTAACATAT TCTGCATTGG GATCTGTTGG GAGACTCGAT TTTTCTTTTT CTGTCCTGTT GGAGTCTGAA GTGTTCTTGT AGCTGCATCA AAGCCCAGTC
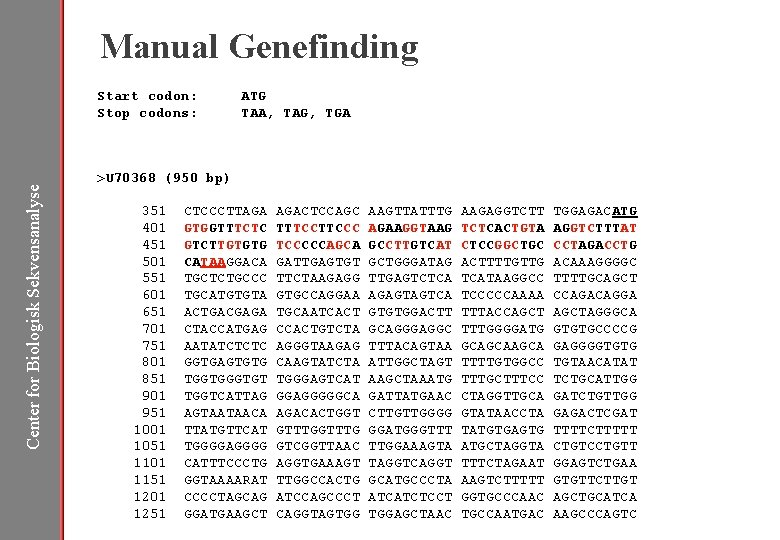
Manual Genefinding Start codon: Stop codons: ATG TAA, TAG, TGA Center for Biologisk Sekvensanalyse >U 70368 (950 bp) 351 401 451 501 551 601 651 701 751 801 851 901 951 1001 1051 1101 1151 1201 1251 CTCCCTTAGA AGACTCCAGC AAGTTATTTG AAGAGGTCTT TGGAGACATG GTGGTTTCTC TTTCCC AGAAGGTAAG TCTCACTGTA AGGTCTTTAT GTCTTGTGTG TCCCCCAGCA GCCTTGTCAT CTCCGGCTGC CCTAGACCTG CATAAGGACA GATTGAGTGT GCTGGGATAG ACTTTTGTTG ACAAAGGGGC TGCTCTGCCC TTCTAAGAGG TTGAGTCTCA TCATAAGGCC TTTTGCAGCT TGCATGTGTA GTGCCAGGAA AGAGTAGTCA TCCCCCAAAA CCAGACAGGA ACTGACGAGA TGCAATCACT GTGTGGACTT TTTACCAGCTAGGGCA CTACCATGAG CCACTGTCTA GCAGGGAGGC TTTGGGGATG GTGTGCCCCG AATATCTCTC AGGGTAAGAG TTTACAGTAA GCAGCA GAGGGGTGTG GGTGAGTGTG CAAGTATCTA ATTGGCTAGT TTTTGTGGCC TGTAACATAT TGGTGT TGGGAGTCAT AAGCTAAATG TTTGCTTTCC TCTGCATTGG TGGTCATTAG GGAGGGGGCA GATTATGAAC CTAGGTTGCA GATCTGTTGG AGTAATAACA AGACACTGGT CTTGTTGGGG GTATAACCTA GAGACTCGAT TTATGTTCAT GTTTG GGATGGGTTT TATGTGAGTG TTTTCTTTTT TGGGGAGGGG GTCGGTTAAC TTGGAAAGTA ATGCTAGGTA CTGTCCTGTT CATTTCCCTG AGGTGAAAGT TAGGTCAGGT TTTCTAGAAT GGAGTCTGAA GGTAAAARAT TTGGCCACTG GCATGCCCTA AAGTCTTTTT GTGTTCTTGT CCCCTAGCAG ATCCAGCCCT ATCATCTCCT GGTGCCCAAC AGCTGCATCA GGATGAAGCT CAGGTAGTGG TGGAGCTAAC TGCCAATGAC AAGCCCAGTC
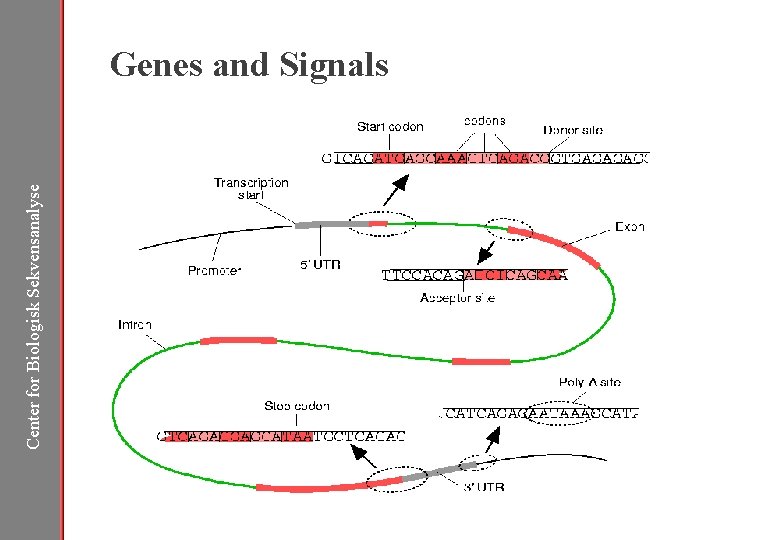
Center for Biologisk Sekvensanalyse Genes and Signals
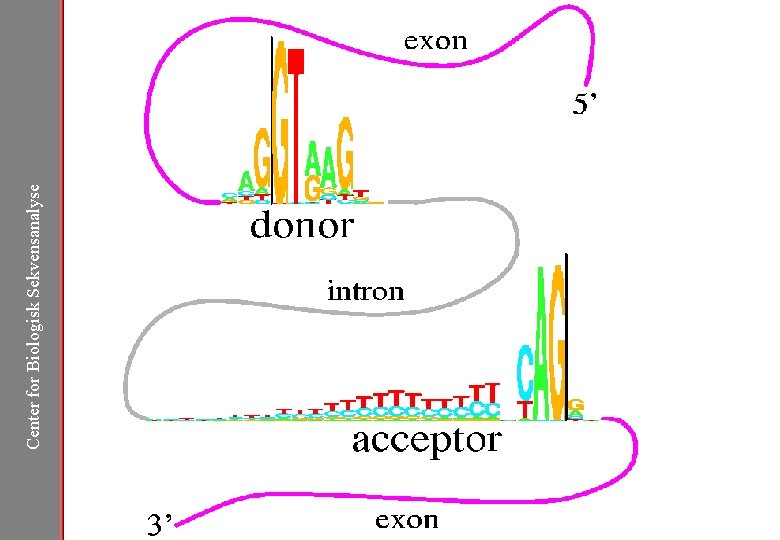
Center for Biologisk Sekvensanalyse
![Manual Genefinding Start codon: ATG Stop codons: TAA, TAG, TGA Donor splice site: ^GT[AG]AG Manual Genefinding Start codon: ATG Stop codons: TAA, TAG, TGA Donor splice site: ^GT[AG]AG](http://slidetodoc.com/presentation_image/c94c96acc891c871d8a0968913baa50d/image-19.jpg)
Manual Genefinding Start codon: ATG Stop codons: TAA, TAG, TGA Donor splice site: ^GT[AG]AG Acceptor splice site: [CT]AG^ Center for Biologisk Sekvensanalyse >U 70368 (950 bp) 351 401 451 501 551 601 651 701 751 801 851 901 951 1001 1051 1101 1151 1201 1251 CTCCCTTAGA AGACTCCAGC AAGTTATTTG AAGAGGTCTT TGGAGACATG GTGGTTTCTC TTTCCC AGAAGGTAAG TCTCACTGTA AGGTCTTTAT GTCTTGTGTG TCCCCCAGCA GCCTTGTCAT CTCCGGCTGC CCTAGACCTG CATAAGGACA GATTGAGTGT GCTGGGATAG ACTTTTGTTG ACAAAGGGGC TGCTCTGCCC TTCTAAGAGG TTGAGTCTCA TCATAAGGCC TTTTGCAGCT TGCATGTGTA GTGCCAGGAA AGAGTAGTCA TCCCCCAAAA CCAGACAGGA ACTGACGAGA TGCAATCACT GTGTGGACTT TTTACCAGCTAGGGCA CTACCATGAG CCACTGTCTA GCAGGGAGGC TTTGGGGATG GTGTGCCCCG AATATCTCTC AGGGTAAGAG TTTACAGTAA GCAGCA GAGGGGTGTG GGTGAGTGTG CAAGTATCTA ATTGGCTAGT TTTTGTGGCC TGTAACATAT TGGTGT TGGGAGTCAT AAGCTAAATG TTTGCTTTCC TCTGCATTGG TGGTCATTAG GGAGGGGGCA GATTATGAAC CTAGGTTGCA GATCTGTTGG AGTAATAACA AGACACTGGT CTTGTTGGGG GTATAACCTA GAGACTCGAT TTATGTTCAT GTTTG GGATGGGTTT TATGTGAGTG TTTTCTTTTT TGGGGAGGGG GTCGGTTAAC TTGGAAAGTA ATGCTAGGTA CTGTCCTGTT CATTTCCCTG AGGTGAAAGT TAGGTCAGGT TTTCTAGAAT GGAGTCTGAA GGTAAAARAT TTGGCCACTG GCATGCCCTA AAGTCTTTTT GTGTTCTTGT CCCCTAGCAG ATCCAGCCCT ATCATCTCCT GGTGCCCAAC AGCTGCATCA GGATGAAGCT CAGGTAGTGG TGGAGCTAAC TGCCAATGAC AAGCCCAGTC
![Manual Genefinding Start codon: ATG Stop codons: TAA, TAG, TGA Donor splice site: ^GT[AG]AG Manual Genefinding Start codon: ATG Stop codons: TAA, TAG, TGA Donor splice site: ^GT[AG]AG](http://slidetodoc.com/presentation_image/c94c96acc891c871d8a0968913baa50d/image-20.jpg)
Manual Genefinding Start codon: ATG Stop codons: TAA, TAG, TGA Donor splice site: ^GT[AG]AG Acceptor splice site: [CT]AG^ Center for Biologisk Sekvensanalyse >U 70368 (950 bp) 351 401 451 501 551 601 651 701 751 801 851 901 951 1001 1051 1101 1151 1201 1251 CTCCCTTAGA AGACTCCAGC AAGTTATTTG AAGAGGTCTT TGGAGACATG GTGGTTTCTC TTTCCC AGAAGGTAAG TCTCACTGTA AGGTCTTTAT GTCTTGTGTG TCCCCCAGCA GCCTTGTCAT CTCCGGCTGC CCTAGACCTG CATAAGGACA GATTGAGTGT GCTGGGATAG ACTTTTGTTG ACAAAGGGGC TGCTCTGCCC TTCTAAGAGG TTGAGTCTCA TCATAAGGCC TTTTGCAGCT TGCATGTGTA GTGCCAGGAA AGAGTAGTCA TCCCCCAAAA CCAGACAGGA ACTGACGAGA TGCAATCACT GTGTGGACTT TTTACCAGCTAGGGCA CTACCATGAG CCACTGTCTA GCAGGGAGGC TTTGGGGATG GTGTGCCCCG AATATCTCTC AGGGTAAGAG TTTACAGTAA GCAGCA GAGGGGTGTG GGTGAGTGTG CAAGTATCTA ATTGGCTAGT TTTTGTGGCC TGTAACATAT TGGTGT TGGGAGTCAT AAGCTAAATG TTTGCTTTCC TCTGCATTGG TGGTCATTAG GGAGGGGGCA GATTATGAAC CTAGGTTGCA GATCTGTTGG AGTAATAACA AGACACTGGT CTTGTTGGGG GTATAACCTA GAGACTCGAT TTATGTTCAT GTTTG GGATGGGTTT TATGTGAGTG TTTTCTTTTT TGGGGAGGGG GTCGGTTAAC TTGGAAAGTA ATGCTAGGTA CTGTCCTGTT CATTTCCCTG AGGTGAAAGT TAGGTCAGGT TTTCTAGAAT GGAGTCTGAA GGTAAAARAT TTGGCCACTG GCATGCCCTA AAGTCTTTTT GTGTTCTTGT CCCCTAGCAG ATCCAGCCCT ATCATCTCCT GGTGCCCAAC AGCTGCATCA GGATGAAGCT CAGGTAGTGG TGGAGCTAAC TGCCAATGAC AAGCCCAGTC

Gene Features Center for Biologisk Sekvensanalyse Codon frequency/bias • Organism dependent • Hexamer statistics Transcriptional • Promoters/enhancers Exon/introns • Length distributions • ORFs Splicing • Donor/acceptor sites • Branchpoints Translational • Start codon context
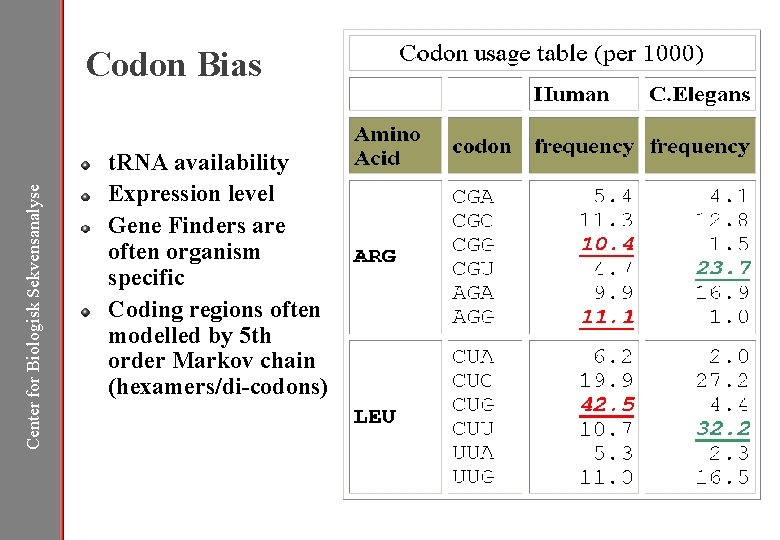
Center for Biologisk Sekvensanalyse Codon Bias t. RNA availability Expression level Gene Finders are often organism specific Coding regions often modelled by 5 th order Markov chain (hexamers/di-codons)
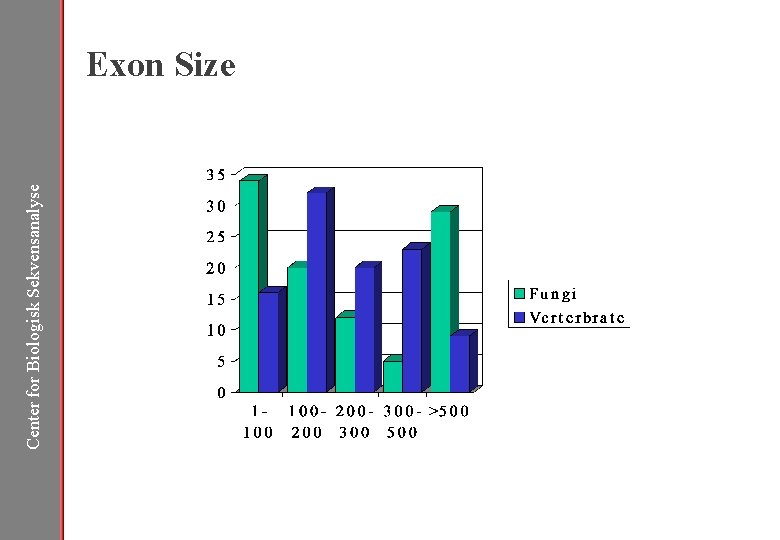
Center for Biologisk Sekvensanalyse Exon Size
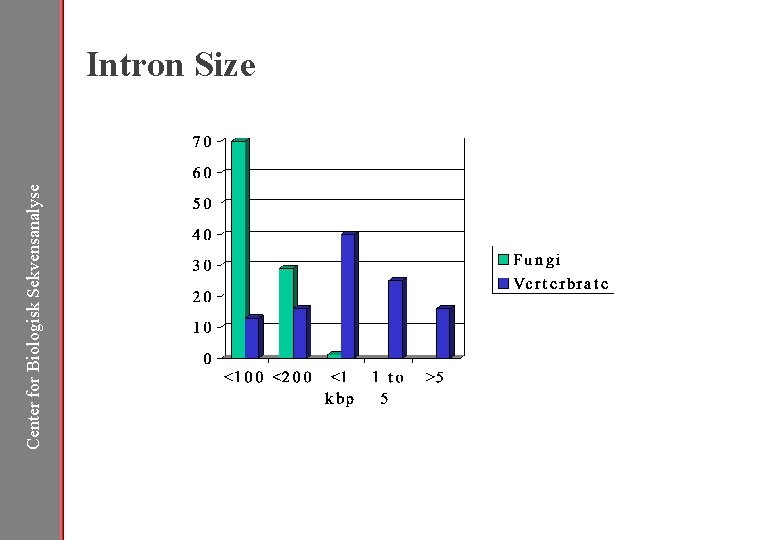
Center for Biologisk Sekvensanalyse Intron Size
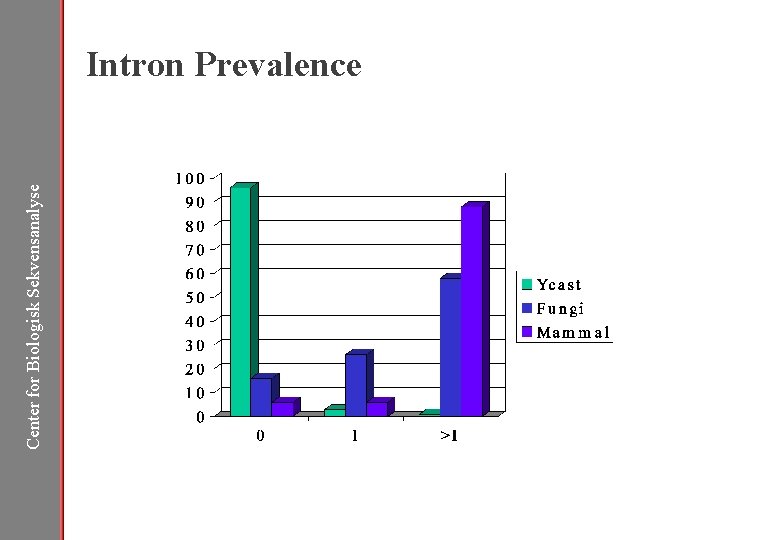
Center for Biologisk Sekvensanalyse Intron Prevalence
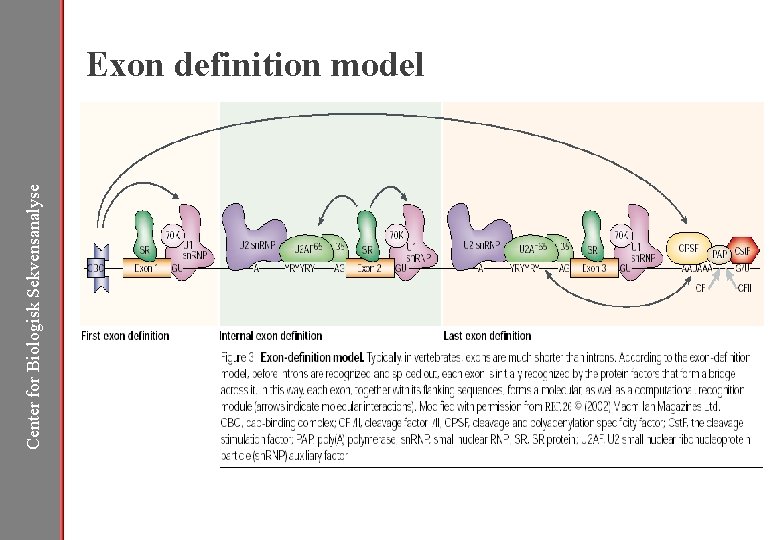
Center for Biologisk Sekvensanalyse Exon definition model

Gene Prediction Center for Biologisk Sekvensanalyse Eukaryotic Gene Prediction • Prediction relies on integration of several gene features • Each gene feature carries a low signal • E. g. ATG, splice sites, etc. • Combinatorial explosion • Some are mutually exclusive (e. g. reading frame) • Sensor based HMMs well suited for gene prediction

Gene Prediction Center for Biologisk Sekvensanalyse Take home messages • Human genome sequence is known • Number of human genes is unknown! • Before 2001: est. 30, 000 -140, 000 • Anno 2003: 25, 000 -40, 000 • Why? • Because gene structure prediction is hard! • Location, structure and function of many human genes is unknown! • Genes may be discovered by different means and methods • . . .
Center for Biologisk Sekvensanalyse The End

Center for Biologisk Sekvensanalyse
Gene Finding Challenges Center for Biologisk Sekvensanalyse Need the correct reading frame • Introns can interrupt an exon in mid-codon There is no hard and fast rule for identifying donor and acceptor splice sites • Signals are very weak

Center for Biologisk Sekvensanalyse Overpredicting Genes Easy to predict all exons Report all sequences flanked by. . AG and GT. . as exons Sensitivity = 100% Specificity ~ 0%

Center for Biologisk Sekvensanalyse Sensor-based methods Similarity searches misses some/many genes c. DNA/EST libraries are not perfect Ab initio Gene Finders • HMM-based (Hidden Markov Model) • Gen. Scan • HMMgene • Neural network-based • GRAIL • Net. Gene 2 (splice sites)

Gene Prediction Center for Biologisk Sekvensanalyse ”Isolated” methods • Predict individual features • E. g. splice sites, coding regions • Net. Gene (Neural network) – http: //www. cbs. dtu. dk/services/Net. Gene 2/ ”Integrated” methods • Predict genes in context • ”Grammar” of genes • Certain elements in specific order are required – HMMgene http: //www. cbs. dtu. dk/services/HMMgene/ – Gen. Scan (HMM-based) http: //genes. mit. edu/GENSCAN. html

Gene Grammar Center for Biologisk Sekvensanalyse Isolated features HAPPYEUGENEAWASGUYFINDER

Gene Grammar Center for Biologisk Sekvensanalyse Isolated features HAPPYEUGENEAWASGUYFINDER Intron 3’UTR Exon Promoter Exon RBS

Gene Grammar Center for Biologisk Sekvensanalyse Integrated features HAPPYEUGENEAWASGUYFINDER EUGENEFINDERWASAHAPPYGUY

Gene Grammar Center for Biologisk Sekvensanalyse Integrated features EUGENEFINDERWASAHAPPYGUY Prom RBS Exon Intron Exon 3’UTR

Gene Grammar Center for Biologisk Sekvensanalyse ”Isolated” methods (e. g. NN): HAPPYEUGENEAWASGUYFINDER ”Integrated” methods (e. g. HMM): EUGENEFINDERWASAHAPPYGUY
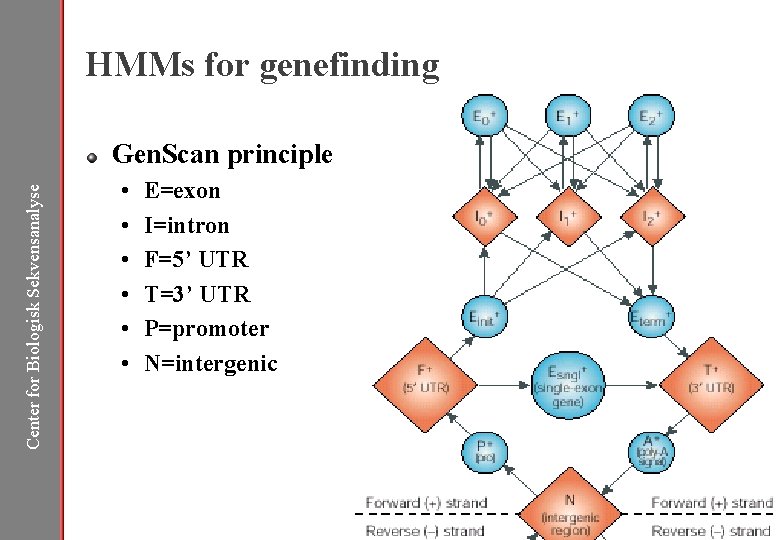
HMMs for genefinding Center for Biologisk Sekvensanalyse Gen. Scan principle • • • E=exon I=intron F=5’ UTR T=3’ UTR P=promoter N=intergenic
Center for Biologisk Sekvensanalyse Genscan http: //genes. mit. edu/GENSCAN. html
Center for Biologisk Sekvensanalyse Genscan
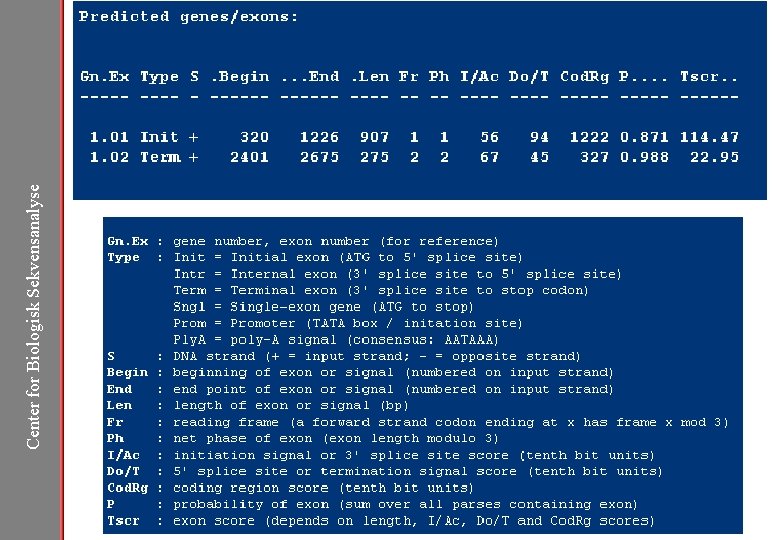
Center for Biologisk Sekvensanalyse Genscan http: //genes. mit. edu/GENSCAN. html
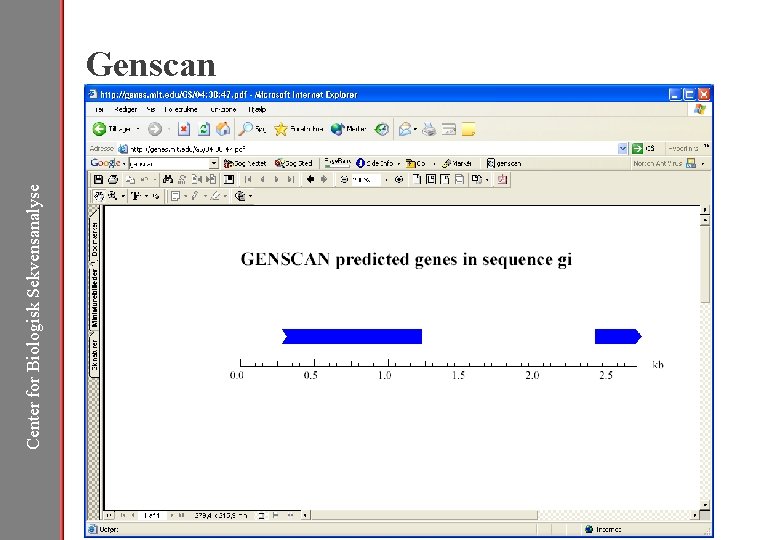
Center for Biologisk Sekvensanalyse Genscan
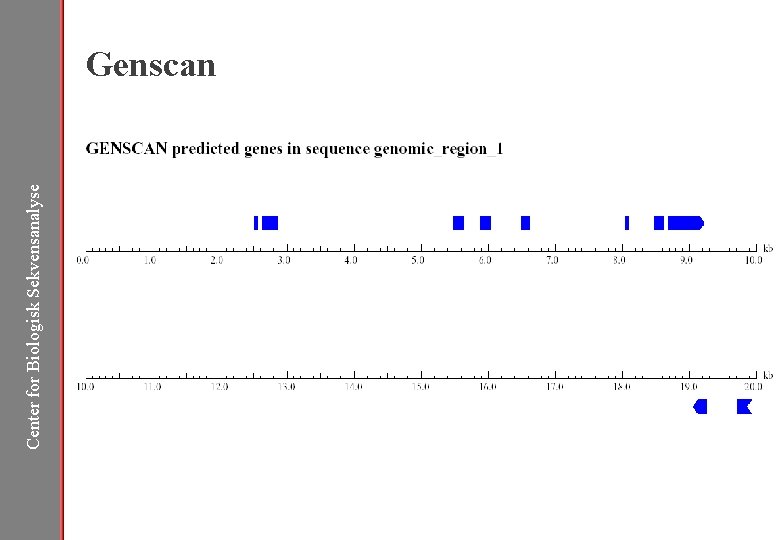
Center for Biologisk Sekvensanalyse Genscan
Center for Biologisk Sekvensanalyse HMMgene http: //www. cbs. dtu. dk/services/HMMgene/

Center for Biologisk Sekvensanalyse HMMgene http: //www. cbs. dtu. dk/services/HMMgene/ Columns 1. Sequence identifier 2. Program name 3. Prediction (see table below for the meaning). 4. Beginning 5. End 6. Score between 0 and 1 7. Strand: $+$ for direct and $-$ for complementary 8. Frame (for exons it is the position of the donor in the frame) 9. Group to which prediction belong. If several CDS's are found they will be called cds_1, cds_2, etc. `bestparse: ' is there because alternative predictions will also be available (see below). Name firstex exon_N lastex singleex CDS Meaning The coding part of the first coding exon starting with the first base of the start codon. The N'th predicted internal coding exon. The coding part of the last coding exon ending with the last base of the stop codon. The coding part of an exon in a gene with only one coding exon. Coding region composed of the exon predictions prior to this line.
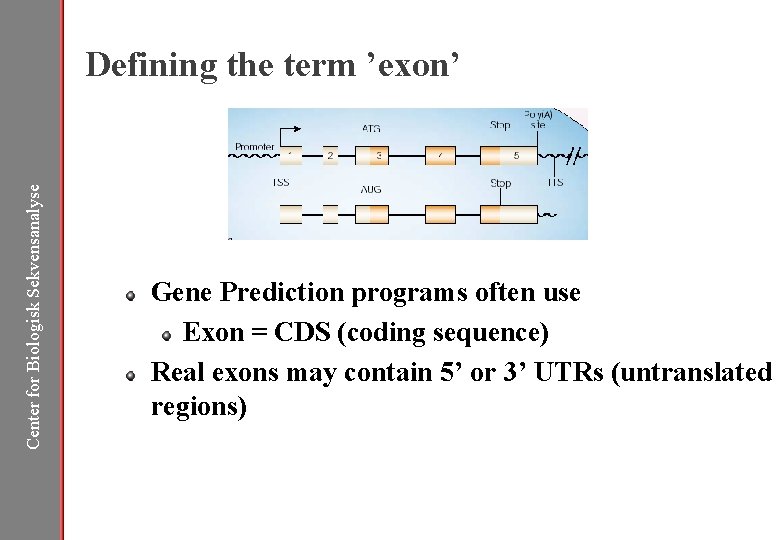
Center for Biologisk Sekvensanalyse Defining the term ’exon’ Gene Prediction programs often use Exon = CDS (coding sequence) Real exons may contain 5’ or 3’ UTRs (untranslated regions)
Center for Biologisk Sekvensanalyse Gene Prediction – Net. Gene 2
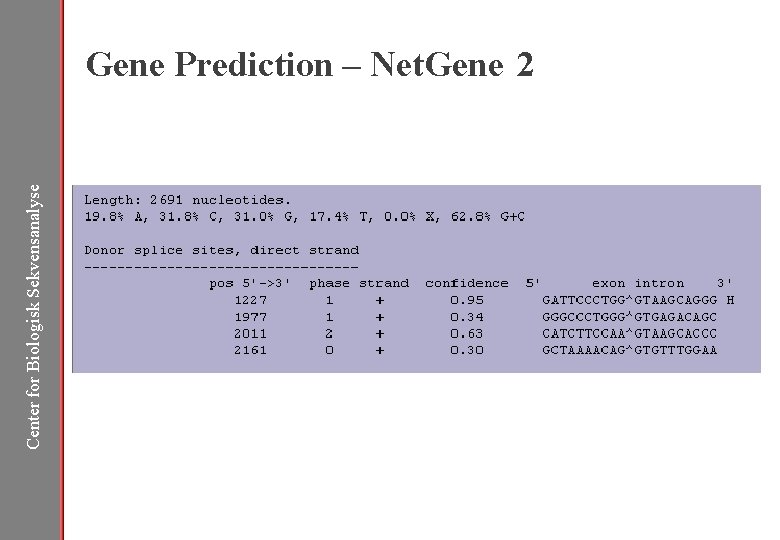
Center for Biologisk Sekvensanalyse Gene Prediction – Net. Gene 2
Center for Biologisk Sekvensanalyse Gene Prediction – Net. Gene 2
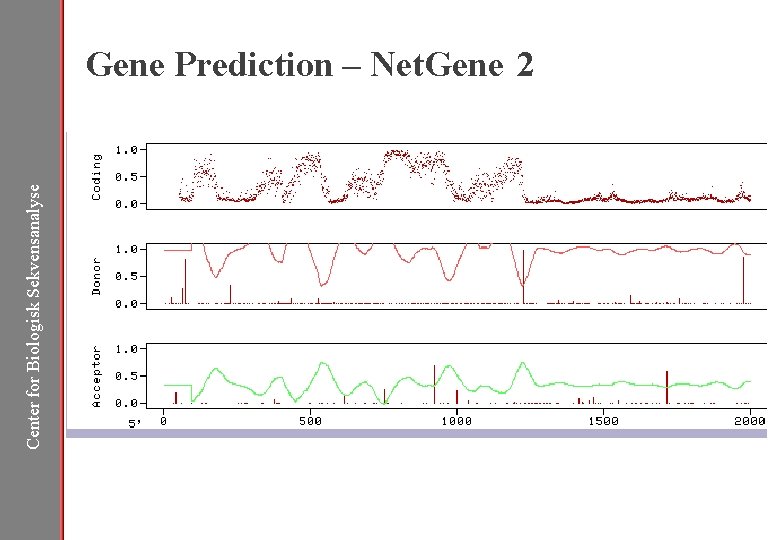
Center for Biologisk Sekvensanalyse Gene Prediction – Net. Gene 2
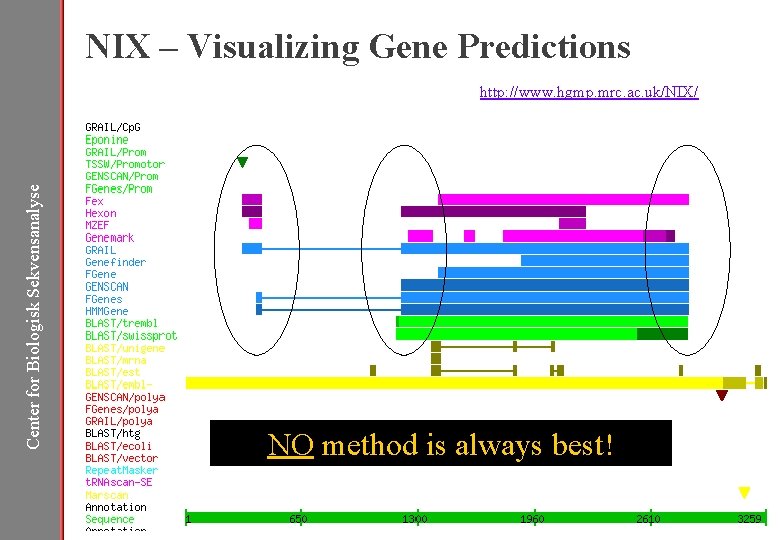
NIX – Visualizing Gene Predictions Center for Biologisk Sekvensanalyse http: //www. hgmp. mrc. ac. uk/NIX/ NO method is always best!
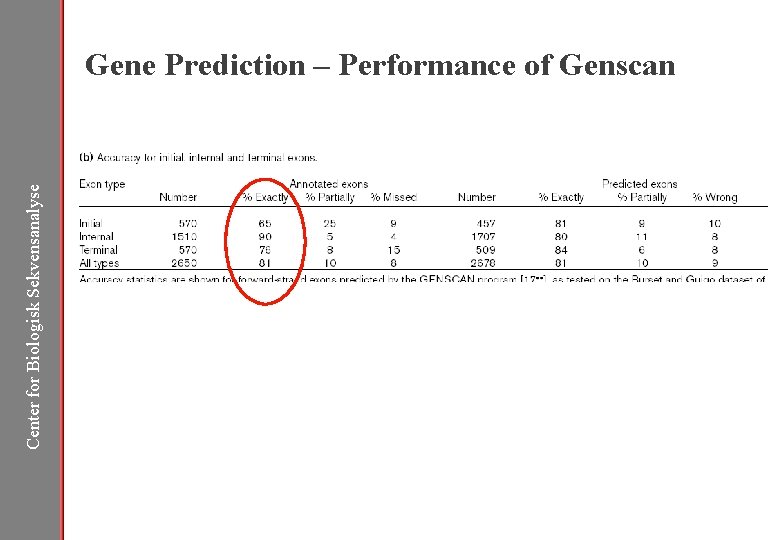
Center for Biologisk Sekvensanalyse Gene Prediction – Performance of Genscan
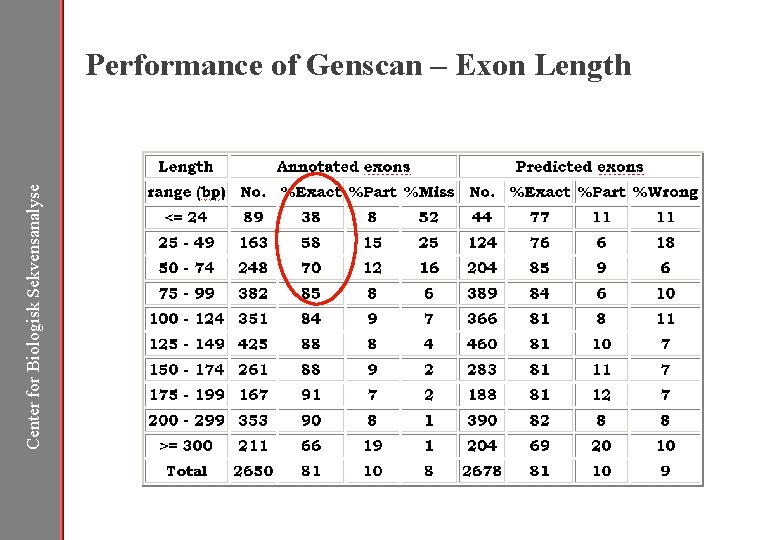
Center for Biologisk Sekvensanalyse Performance of Genscan – Exon Length

Center for Biologisk Sekvensanalyse Repeatmasker Repetitive sequences in human/eukaryotic genomes are a problem Run gene predictions on large genomic regions before and after masking of repetitive sequence: • http: //ftp. genome. washington. edu/cgi-bin/Repeat. Masker Up to 45% of human genomic sequence derived from transposable/repetitive elements

Center for Biologisk Sekvensanalyse Repeatmasker

Future Challenges Center for Biologisk Sekvensanalyse Bootstrapping: prediction improves as more genes become known • ’Extreme’ genes (long/short) still difficult • Initial and terminal exons are predicted with lower confidence Combine with Sequence Similarity Matches Non-coding RNAs • Most gene prediction programs only predict protein-coding genes • t. RNA and r. RNA genes are not predicted Predict alternatice splicing, enhancers and silencers Predict matrix- and scaffold-attachment regions, insulators and boundary elements

Gene Prediction Center for Biologisk Sekvensanalyse Take home messages • Human genome sequence is known • Number of human genes is unknown! • Before 2001: est. 30, 000 -140, 000 • Anno 2003: 25, 000 -40, 000 • Location, structure and function of many human genes is unknown! • Genes may be discovered by different means and methods • . . .

Gene Prediction Center for Biologisk Sekvensanalyse Take home messages • Genes may be predicted by computer programs • Masking of repetitive sequences may be required for large genomic sequences • ’Unusual’ genes are difficult (high GC%, short or terminal exons) • HMM-based gene prediction programs are suitable for “Gene Grammar” Prediction methods are not perfect!
Center for Biologisk Sekvensanalyse The End
- Slides: 61