Gene Expression Overview By Salwa Hassan Teama Roche
Gene Expression Overview By Salwa Hassan Teama Roche 1

Gene Expression Overview By Salwa Hassan Teama M. D. N. C. I. Cairo University, Egypt 2

Contents n n n The Gene Housekeeping Gene Steps of Gene Expression Overview Transcription RNA Processing Protein Synthesis (Translation) Requirement for translation Phases of Protein Synthesis Eukaryotic Vs Prokaryotic Control of Gene Expression Molecular Biology Glossary online http: //seqcore. brcf. med. umich. edu/doc/educ/dnapr/mbglossary/mbgloss. html 3

The Gene n As genes are located, their functions need to be determined, and studies of gene expression become the focus. n Gene; It is a segment within a very long stand of DNA located on a chromosome. n In molecular terms, a gene defined as nucleic acid sequence that is necessary for synthesis for the functional polypeptide. n The DNA of a normal human cell contains an estimated 35 to 40, 000 genes, but only a fraction of these are used (or “expressed”) in any particular cell at any given time. 4

The Gene n The gene is the fundamental unit of inheritance and the ultimate determinant of all phenotypes. n The average gene contains 7 -10 exons spread over 10 -16 kb of DNA. The immediate product is the primary transcript. n Housekeeping genes are expressed in all cells all the time, are responsible for the routine metabolic functions common to all cells. 5
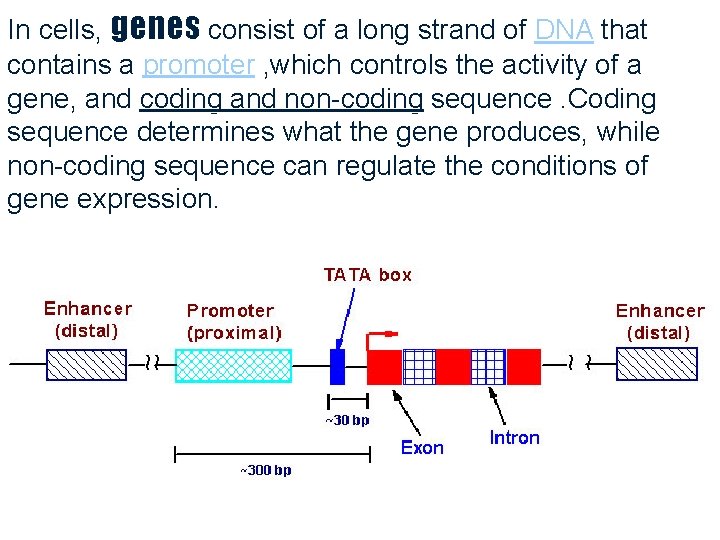
In cells, genes consist of a long strand of DNA that contains a promoter , which controls the activity of a gene, and coding and non-coding sequence. Coding sequence determines what the gene produces, while non-coding sequence can regulate the conditions of gene expression. 6
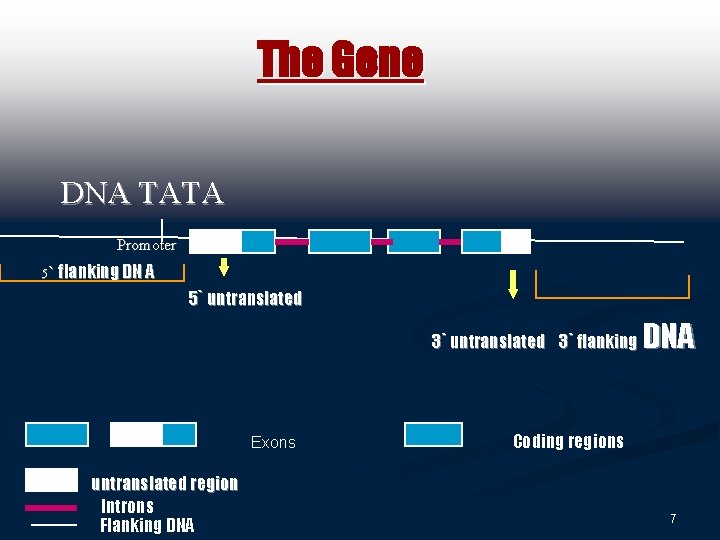
The Gene DNA TATA Promoter 5` flanking DN A 5` untranslated 3` flanking Exons untranslated region Introns Flanking DNA Coding regions 7

Housekeeping genes n Some are expressed all the time e. g. a plasma cell expresses continuously the gene for the antibody it synthesizes. n Some are expressed as a cell enters a particular pathway of differentiation. n Some are expressed only in the cell change. e. g. the arrival of a hormone may turn on (or off) certain genes in that cell. 8
9

Gene Expression n n Gene expression is the process by which a gene's information is converted into the structures and functions of a cell by which a gene's DNA sequence is converted into the functional proteins of the cell. Non-protein coding genes (e. g. r. RNA genes, t. RNA genes) are not translated into protein. When a gene is active, the coding and non-coding sequence is copied in a process called transcription , producing an RNA copy of the gene's information. This RNA can then direct the synthesis of proteins via the genetic code. However, RNAs can also be used directly, for example as part of the ribosome. These molecules resulting from gene expression, whether RNA or protein , are known as gene products. 10

Gene expression is a multi-step process that begins with: Transcription DNA → RNA and * Translation RNA → protein Followed by: Folding Post-translational modification and Targeting 11

Gene expression: Steps : n Transcription: A DNA strand is used as a template to synthesize a complementary RNA strand, which is called the primary transcript. n RNA processing: This step involves modifications of the primary transcript to generate a mature m. RNA (for protein genes) or a functional t. RNA or r. RNA. n Nuclear transport: m. RNA has to be transported from the nucleus to the cytoplasm for protein synthesis. 12

Gene expression: Steps : n Protein synthesis: In the cytoplasm, m. RNA binds to ribosomes, which can synthesize a polypeptide based on the sequence of m. RNA. n For RNA genes (t. RNA and r. RNA), the expression is complete after a functional t. RNA or r. RNA is generated. However, protein genes require additional steps. n The amount of protein that a cell expresses depends on: the tissue, the developmental stage of the organism and the metabolic or physiologic state of the cell. 13
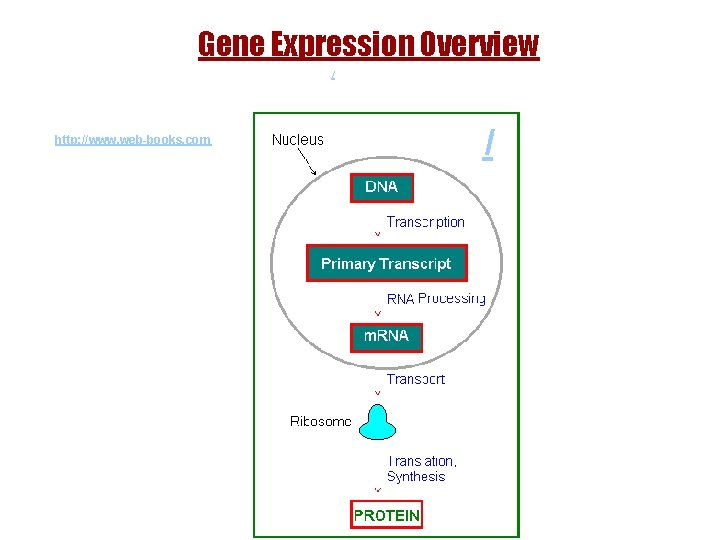
Gene Expression Overview / http: //www. web-books. com / 14

*Transcription* n Transcription is the process through which a DNA n Transcription proceeds in the 5' → 3' direction. n Transcription is divided into 3 stages : Initiation Elongation and Termination. n n n sequence is enzymatically copied by an RNA polymerase to produce a complementary RNA. Or, in other words, the transfer of genetic information from DNA into RNA. 15

Transcription n In the case of protein-encoding DNA, transcription is the beginning of the process that ultimately leads to the translation of the genetic code )via the m. RNA intermediate (into a functional peptide or protein. n The stretch of DNA that is transcribed into an RNA molecule is called transcription unit. 16
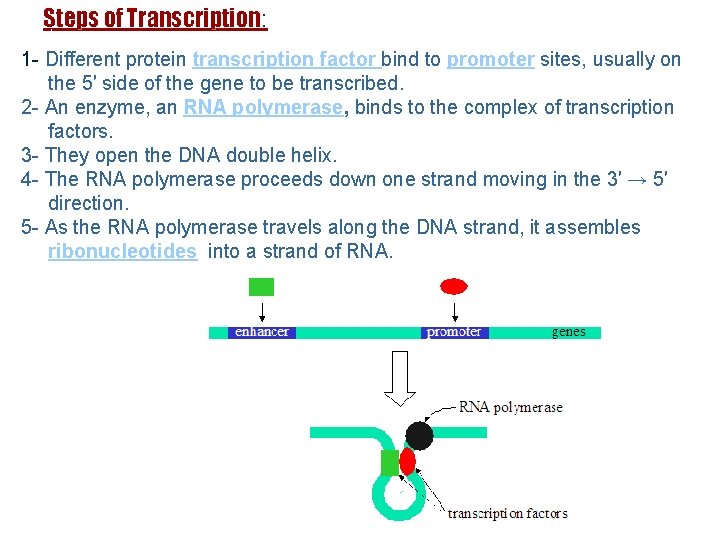
Steps of Transcription: 1 - Different protein transcription factor bind to promoter sites, usually on the 5′ side of the gene to be transcribed. 2 - An enzyme, an RNA polymerase, binds to the complex of transcription factors. 3 - They open the DNA double helix. 4 - The RNA polymerase proceeds down one strand moving in the 3′ → 5′ direction. 5 - As the RNA polymerase travels along the DNA strand, it assembles ribonucleotides into a strand of RNA. 17

Steps of Transcription: 6. Each ribonucleotide is inserted into the growing RNA strand following the rules of base pairing. Thus for each C on the DNA strand, a G is inserted in the RNA; for each G, a C; and for each T, an A. However, each A on the DNA guides the insertion of the pyrimidine uracil (U, from uridine triphosphate, UTP). There is no T in RNA. 7. Synthesis of the RNA proceeds in the 5′ → 3′ direction. 8. As each nucleoside triphosphate is brought in to add to the 3′ end of the growing strand, the two terminal phosphates are removed. 9. When transcription is complete, the transcript is released from the polymerase and, shortly thereafter, the polymerase is released from the DNA. 18

Transcription in eukaryotes n Transcription is complex process. Most genes in eukaryotic cells contain noncoding regions (Introns); present among the coding regions (exons) that actually code for the final protein. n To initiate transcription: RNA polymerase copies both the exons and the introns to form what is called precursor m. RNA or pre-m. RNA. n RNA Processing: pre-m. RNA → m. RNA All the primary transcripts produced in the nucleus must undergo processing steps to produce functional RNA molecules for export to the cytosol. 19

*RNA Processing* 1. Synthesis of the cap. This is a modified guanine (G) which is attached to the 5′ end of the pre-m. RNA. The cap protects the RNA from being degraded by enzymes that degrade RNA from the 5′ end. 2. Step-by-step removal of introns present in the pre-m. RNA and splicing of the remaining exons. 3. The removal of introns and splicing of exons is done with the spliceosome. This is a complex of several sn. RNA molecules and some 145 different proteins. 4. Synthesis of the poly (A) tail. This is a stretch of adenine (A) nucleotides. This completes the m. RNA molecule, which is now ready for export to the cytosol. (The remainder of the transcript is degraded, and the RNA polymerase leaves the DNA). 20

Transcription can be measured and detected in a variety of ways: n Northern blot n RNase protection assay n RT-PCR n In vitro transcription n In situ hybridization n DNA microarrays 21
22

*Protein synthesis* Proteins are made of long stretches of fundamental molecules called amino acids. A protein might be composed of a few or many thousands of amino acids. Amino acids are represented by codons, which are 3 nucleotide RNA sequences. Protein Synthesis is the process in which cells build proteins. The term is sometimes used to refer only to protein translation but more often it refers to a multistep process, beginning with amino acid synthesis and transcription which are then used for translation. 23

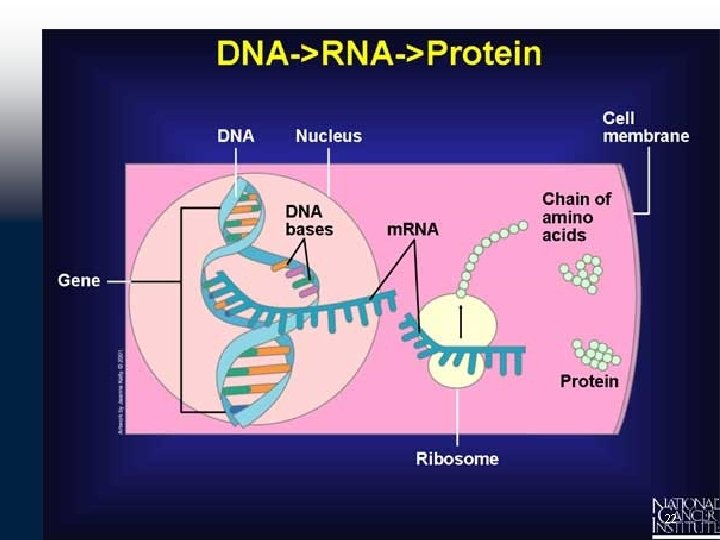
Protein Synthesis DNA Transcription : Step 1 Protein synthesis in eukaryotes is similar to that in bacteria. However, eukaryotic protein synthesis entails more protein components than does prokaryotic protein synthesis, and some steps are different. Protein synthesis begins in the cell's nucleus when the gene encoding a protein is copied into RNA. The RNA copy of the gene is called the m. RNA molecules copies the information stored in the nuclear DNA. After transcription, the m. RNA is transported out of the cell's nucleus through nuclear pores to go to the site of translation, the rough endoplasmic reticulum (discussed before). 24

Protein Synthesis Translation : Step 2 The translation of the information residing in m. RNA nucleotide sequence into the amino acid sequence of a protein, It must be through an intermediate adaptor molecules. These adaptor molecules that translate the codons into amino acid sequences of a protein are called t. RNA. This process involves about 64 codons of 3 nucleotides each to direct one amino acid synthesis which requires more than one codon. There are 3 codons whose job is only to signal termination of protein synthesis and are called stop codon. 25

Protein Synthesis Translation : Step 2 www. expasy. org/people/personal/nguex/ribosome. jpg//: http To convert the m. RNA into protein, t. RNA is used to read the m. RNA sequence, 3 nucleotides at a time. The m. RNA sequence is matched three nucleotides at a time to a complementary set of three nucleotides in the anticodon region of the corresponding t. RNA molecule. Here the translation occurs in the ribosomes where an enzyme catalyzes the formation of a peptide bond between the amino acid bound to t. RNA resulting in a growing protein chain. Reapeatedly, the specific small t-RNA anticodon base pairs with the next m. RNA codon and the enzymes of the ribosomes contine to peptide bonding for protein chain with reapted release of t. RNA giving place to the next t. RNA and so on. . The ribosome moves along the m. RNA until the process is terminated immediately when it meets a stop codon. Here the ribosome and protein products dissociate from m. RNA. 26
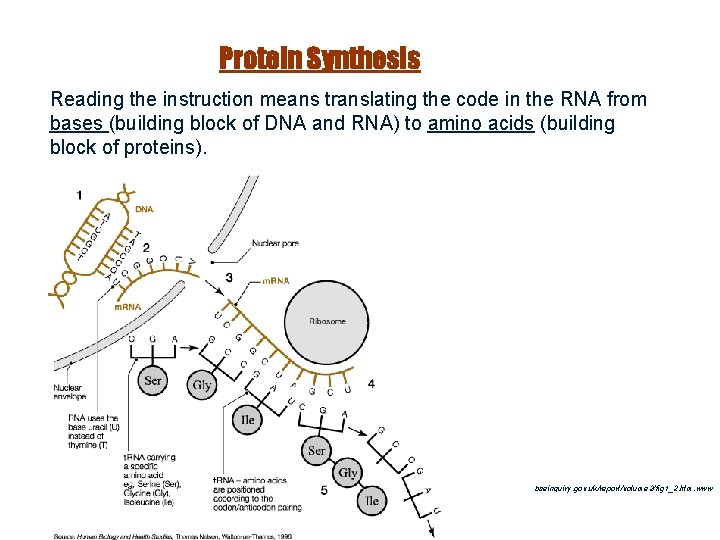
Protein Synthesis Reading the instruction means translating the code in the RNA from bases (building block of DNA and RNA) to amino acids (building block of proteins). www. bseinquiry. gov. uk/report/volume 2/fig 1_2. htm. www 27

Requirement for translation n n n n Ribosomes Amino Acids t. RNA m. RNA template Initiation factors Elongation factors Termination factors Aminoacyl t. RNA synthetase enzymes Energy source 28

Requirement for translation Ribosomes : Eukaryotic ribosomes are larger. They consist of a 60 S large subunit and a 40 S small subunit, which come together to form an 80 S particle having a mass of 4200 kd, compared with 2700 kd for the prokaryotic 70 S ribosome. The 40 S subunit contains an 18 S RNA that is homologous to the prokaryotic 16 S RNA. The 60 S subunit contains three RNAs : the 5 S and 28 S RNAs are the counterparts of the prokaryotic 5 S and 23 S molecules; its 5. 8 S RNA is unique to eukaryotes. n Ribosomes, factory for protein synthesis. Ribosomes are molecular machines that crunch along the m. RNA, and adding appropriate amino acids together to create proteins. The specific amino acids are recognized, and brought to the ribosomes by transfer RNA (t. RNAs. ( 29

Requirement for translation n Amino acids are the monomers which are polymerized to produce proteins. Amino acid synthesis is the set of biochemical processes (metabolic pathways )which build the amino acids from carbon sources like glucose. Not all amino acids may be synthesized by every organism, for example adult humans have to obtain 8 of the 20 amino acids from their diet. The amino acids are then loaded onto t. RNA molecules for use in the process of translation. 30

Requirement for translation t. RNA n Brings amino acids to the ribosome. n Has site for attachment of amino acid. n Has cloverleaf structure with base-paired arms, one variable arm. n Has anticodon sequence to base pair with the codon along the m. RNA. n t. RNA aligns with codon and interprets )translates) information. 33

Requirement for translation m. RNA n Source of coding information for the protein synthesis system. n Contains start and stop signals for translation. n Eukaryotic m. RNA is capped. This is used as the recognition feature for ribosome binding. n The site at which protein synthesis begins on the m. RNA is especially crucial, since it sets the reading frame for the whole length of the message. An error of one nucleotide either way at this stage would cause every subsequent codon in the message to be misread, so that a nonfunctional protein would result, the rate of initiation thus determines the rate at which the protein is synthesized. 34

Requirement for translation n Initiation factors help the ribosome, initiator t. RNA, and other components assemble the at the correct location on the m. RNA and ensure that protein synthesis starts in the correct reading frame. n Elongation factors are responsible for moving the ribosome along the m. RNA and maintain the correct reading frame. Facilitate removal of "used" t. RNAs and bringing in "new" t. RNAs. Termination factors recognize the stop codons and release proteins and ribosomes. n n Aminoacyl t. RNA synthetase enzymes: Each amino acid is activated and linked to a specific transfer RNA by an enzyme called an aminoacyl-t. RNA synthetase. n Energy source: ATP or GTP which are synthesized in the 35 mitochondria

Sequence of events of protein synthesis: 1. 2. Synthesis proceeds from the N-terminus to the C-terminus of the protein. The ribosomes "read" the m. RNA in the 5' to 3' direction. 3. Active translation occurs on polyribosomes (also termed polysomes). This means that more than one ribosome can be bound to and translate a given m. RNA at any one time. 4. Chain elongation occurs by sequential addition of amino acids to the C-terminal end of the ribosome bound polypeptide. 36

Phases of Protein Synthesis The protein synthesis occur in 3 phases: 1 - Accurate and efficient initiation occurs. 2 - Chain elongation. 3 - Accurate and efficient termination. 37

Phases of Protein Synthesis n The initiation phase of protein synthesis requires over 10 eukaryotic Initiation Factors (e. IFs). n The eukaryotic elongation phase closely resembles that in prokaryotes. The corresponding elongation factors are: n e. EF-1 a = EF-Tu n e. EF-1 bg = EF-Ts n e. EF-2 = EF-G n Eukaryotes use a single release factor, e. RF, to recognize all three stop codons. 38

Phases of Protein Synthesis Pre-initiation n Dissociation of the ribosome into 40 S and 60 S subunits. n Formation of 40 S pre-initiation complex: by binding met-t RNA, elf-2, GTP to 40 S ribosome then become stabilized by elf-3 and elf-1 A. 39

Phases of Protein Synthesis: Initiation n n Preparing and checking the m. RNA. Binding the initiator Met-t. RNAi. Met to the 40 S small subunit of the ribosome. Association of the m. RNA complex with the small subunit complex and scanning to find the start codon. Association of the 60 S large subunit of the ribosome, dissociation and recycling of the initiation factors. 40

I- Preparing and checking the m. RNA: This step involves the e. IF 4 factors -which, as a family of factors, play a role in binding to the m. RNA n The 5' end of the m. RNA binds (e. IF 4 E) and with (EIF 4 G) which may recognize secondary structure elements downstream of the 5'-end. Poly A-binding protein, already bound at the 3' end of the m. RNA, interacts with e. IF 4 G. The initiation factors e. IF 4 A and e. IF 4 B join the complex. e. IF 4 A is an RNA helicase which will remove secondary structure from the m. RNA ; e. IF 4 B is an RNA binding protein required for its activity. 41

II- Binding the initiator Met-t. RNA: This step involves e. IF 1, e. IF 2, e. IF 3 and e. IF 5. n e. IF 1 is a general stimulator of initiation n e. IF 2 binds with Met-t. RNAi. Met n e. IF 3 has a general role in binding to the 40 S small subunit and n e. IF 5 stimulates the GTPase activity of e. IF 2/GTP binds with Met-t. RNAi. Met This complex is bound by e. IF 1 -e. IF 3 -e. IF 5 Finally all are bound to the 40 S small subunit of the ribosome. 42

III- m. RNA binding and scanning: The two complexes ( the m. RNA complex and the t. RNA-40 S complex ) now associate with each other. The m. RNA to be correctly positioned for translation: I- The start codon must be located and positioned correctly in the P site of the riobosome. II- The initiator t. RNA must be positioned correctly in the same site. The AUG start codon is located by an ATP-dependent reaction by which helicases (e. IF 4 A and e. IF 4 B) move the m. RNA with respect to the small subunit of the ribosome. Positioning of the initiator t. RNA is checked by a codon-anticodon base pairing mechanism. 43

IV- Association of the 60 S large subunit Once the m. RNA and initiator t. RNA are correctly bound, the 60 S large subunit binds to form 80 s intiation complex with release of the e. IF factors. However, e. IF 2 now has GDP bound a guanine nucleotide release protein is required to exchange GTP for GDP. e. IF 2 B is the guanine nucleotide release protein that carries this out. 44

Elongation: Cyclic process catalyzed by the proteins called eukaryotic elongation factor (e. EFs) At the end of one complete round of amino acid addition the A site will be empty and ready to accept the incoming aminoacylt. RNA dictated by the next codon of the m. RNA. So not only the incoming amino acid need to be attached to the peptide chain but the ribosome must move down the m. RNA to the next codon. Elongation: three distinct steps: 1 - Transfer of proper aminoacyl-t. RNA from cytoplasm to A-site of ribosome. 2 -Peptide bond formation; Covalent linkage of new amino acid to growing polypeptide chain. 3 - Movement of t. RNA from A-site to P-site (translocation). Occurs multiple times per polypeptide. 45

Elongation I- Transfer of proper aminoacyl-t. RNA from cytoplasm to A-site of ribosome. After the formation of complete 80 S ribosome, the A site is free. After proper codon recognition the eukaryotic elongation factor e. EF 1 forms a complex with both GTP and the aminoacyl t. RNA. This complex allows the aminoacyl t. RNA to enter the free A site. II- Peptide bond formation; Covalent linkage of new amino acid to growing polypeptide chain. t. RNA molecule with the matching anticodon for the codon lined up with the A site. Then, a peptide bond forms between the two amino acids which are being held adjacent to one another. This bond formation requires energy from ATP. III- Translocation The peptidyl moiety is discharged from t-RNA in the P site and then t-RNA dissociates and P site is free. Then translocation of the new peptidyl t-RNA in the A site into the free P site occurs by the action of e. EF-2 and GTP. Now the A site is free for another cycle of aminoacyl t-RNA codon recognition and elongation. 46

Termination Like initiation and elongation, translational termination requires specific protein factors identified as releasing factors, RFs in E. coli and e. RFs in eukaryotes. There are 2 RFs in E. coli and one in eukaryotes. The signals for termination are the same in both prokaryotes and eukaryotes. These signals are termination codons present in the m. RNA. There are 3 termination codons, UAG, UAA and UGA. n After multiple cycles of elongation and polymerization of specific amino acids into protein molecules, a nonsense codon= termination codon of m. RNA appear in the A site. The is recognized as terminal signal by eukaryotic releasing factors (e. EF). Reading of final m. RNA codon (STOP codon) (UAA, UAG, and UGA) and dissociation of polypeptide from ribosome. n The e. RF plus GTP plus peptidyl transferase cause hydrolysis of the bond between the peptide and t-RNA in P site giving water molecule instead of amino acid, hence the separated protein from t-RNA is released from the P site, the 80 S ribosome dissociates again into 40 S and 60 S subunits to be recycled again. The m. RNA is released from the dissociated ribosome (into 40 S& 60 S) for another cycle. n 47

RNA Codon GAC UUA t. RNA Amino Acids After process of transcription and translation are complete we left with protein synthesis Protein consist of chain of Aspartic Acid, Leucine……………. . Hundred or thousand of Amino Acids 48
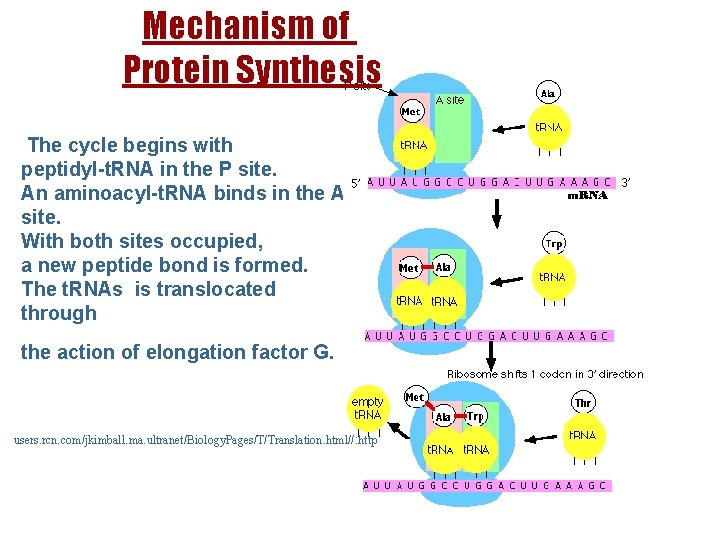
Mechanism of Proteinof Synthesis Mechanism Protein Synthesis. http//: users. rcn. com/jkimball. ma. ultranet/Biology. Pages/T/Translation. html The cycle begins with peptidyl-t. RNA in in the P Psite. peptidyl-t. RNA the site. An aminoacyl-t. RNA bindsininthe A A An aminoacyl-t. RNA binds site. both sites occupied, With both sites occupied, a new peptide bond formed. a new peptide bond isisformed. The t. RNAs is translocated through The t. RNAs is translocated the action of elongation factor G. through the action of elongation factor G. users. rcn. com/jkimball. ma. ultranet/Biology. Pages/T/Translation. html//: http 49

Prokaryotic vs. eukaryotic n Major difference between eukaryotes and prokaryotes is that, in a typical eukaryotic cell, protein synthesis takes place in the cytoplasm while transcription and RNA processing take place in the nucleus. In bacteria, these two processes can be coupled so that protein synthesis can start even before transcription has finished. n Eukaryote genes are not grouped in operons as are prokaryote genes. Each eukaryote gene is transcribed separately, with separate transcriptional controls on each gene. 50

Prokaryotic vs. eukaryotic n To initiate transcription in bacteria, sigma factors bind to RNA polymerases/ sigma factors complex can then bind to promoter about 40 deoxyribonucleotide bases prior to the coding region of the gene. When the complex recognizes the correct promoter, the sigma factor released from the RNA polymerase and the enzyme begins to open the helix of the DNA. n RNA polymerase moves down the DNA, till "stop" signal is reached. This causes the completed m. RNA to drop off the gene. n Once the RNA polymerase leave the promotor region, a new molecule of RNA polymerase can bind to the promotor and begins a new transcription process. So, a single gene can be transcribed multiple times. 51

Prokaryotic vs. eukaryotic n Prokaryotes have one type of RNA polymerase for all types of RNA, eukaryotes have a separate RNA polymerase for each type of RNA. One enzyme for m. RNA-coding genes such as structural proteins, second enzyme for large r. RNAs, third enzyme for smaller r. RNAs and t. RNAs. n The existence of introns in prokaryotes is extremely rare. Essentially all humans' genes contain introns. A notable exception is the histone genes which are intronless. In many genes the presence of introns separates exons into coding regions exhibiting distinct functional domains. 52

Control of Gene Expression Types of control in Eukaryotes: Transcriptional, prevent transcription, prevent m. RNA from being synthesized. Posttranscriptional, control m. RNA after it has been produced. Translational, prevent translation; involve protein factors needed for translation. Posttranslational, after the protein has been produced. 53

Control of Gene Expression Transcriptional Control: 1 -The transcription factors are necessary for RNA polymerase to attach. 2 - Hormones activate transcription factors. Post Transcriptional Control: 1 - Removal of introns, Differential removal of introns enables a gene to code for more than one different protein. 2 -Presence of ribonuclease, enzymes that destroy m. RNA. 54

Control of Gene Expression Translational Control: 1 - Leader Proteins and other molecules can bind to the leader which can enhance or restrict ribosome binding (and thus translation). 2 - Initiation factors are proteins that enable ribosomes to attach to m. RNA. Post Translational Control: Protein Activation Protein modification through: * Folding * Enzymatic cleavage or * Bond formation. * Chemical modification: Addition of: Methyl group, Phosphoryl group or Glycosyl group. 55

References & Online Further Reading n n n Innis MA, Gelfand DH. PCR Protocols: A quide to methods and application. San Diego: Academic Press, 1990 Mullis KB, Ferre F eds. The polymerase chain reaction. Boston, MA: Brikhauser Press, 1994. Daiel HFarkas. DNA Simplified. The Hitchhiker`s guide to DNA AACC Press. Washintgton D. C. Aboul-Enein. Molecular biology, NCI, Cairo University Chapter 26, Translation, pages 867 - 883 STRYER Chapter 34, Protein Synthesis, pages 903 -906 56 LEHNINGER Chapter 26, Protein Metabolism, pages 917 - 924 Chapter 26, Protein Metabolism, pages 927 - 928 TAMARIN Chapter 12, pages 277 – 288 29. 5. Eukaryotic Protein Synthesis Differs from Prokaryotic Protein Synthesis Primarily in Translation Initiation in Biochemistry, fifth edition eds (Geremy M. Berg, John. L. Tymoczko, and Lubert Stryer) http: //en. wikipedia. org/wiki/Protein_biosynthesis http: //ntbiouser. unibe. ch/trachsel/teaching/translation/initi ation. Online. html 56

References & Online Further Reading n n n n n http: //www. accessexcellence. org/RC/VL/GG/control_Express. html http: //www. cc. ndsu. nodak. edu/instruct/mcclean/plsc 431/geneexpres http: //www. biology. arizona. edu/cell_bio/tutorials/pev/page 3. html% Prokaryotes and Euokaryotes: http: www. biology. arizona. edu/cell_bio/tutorials/pev/ Prokaryotes: en. wikipedia. org/wiki/Prokaryote http: //www. indstate. edu/thcme/mwking/genehttp: //www. emc. maricopa. edu/faculty/farabee Geneti. CCode: en. wikipedia. org/wiki/Genetic_code Genetic Code: www. accessexcellence. org/AB/GG/genetic. html http: /www. med. uottawa. ca/patho/devel/index. html Protein Synthesis: http: //users. rcn. com/jkimball. ma. ultranet/Biology. Pages/C/Codons. ht mlhttp: //www. emc. maricopa. edu/faculty/farabee/biobk/Bio. Book. PROTSYn. html 57
- Slides: 55