Gene Expression II 1 Transcription Factor Binding Sites

Gene. Expression II: 1. Transcription Factor Binding Sites 2. Microarrays 26 th May, 2010 Karsten Hokamp Genetics Department Gene. Expression II BI 2010 1

TFBS prediction - Overview • Introduction • Methods • Implementations • Analyse 2 kb upstream of eve Gene. Expression II BI 2010 2

TFBS prediction - Introduction • TFBS = DNA motifs = 5 – 20 bp long = variable = multiple occurrences/sites per gene = combination of activators and repressors • cis-regulatory regions = clusters of TFBS -20 kb – first intron Gene. Expression II BI 2010 3
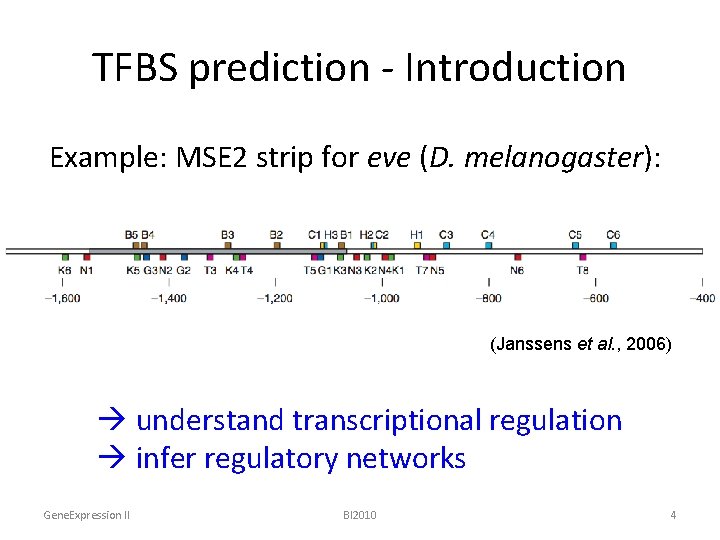
TFBS prediction - Introduction Example: MSE 2 strip for eve (D. melanogaster): (Janssens et al. , 2006) understand transcriptional regulation infer regulatory networks Gene. Expression II BI 2010 4

TFBS prediction - Methods • De novo motif prediction (overrepresentation) • Searching for known motifs • Phylogenetic Footprinting/Shadowing • Clustering of TFBSs • Integration of external data sources (co-expression, structure) Gene. Expression II BI 2010 5
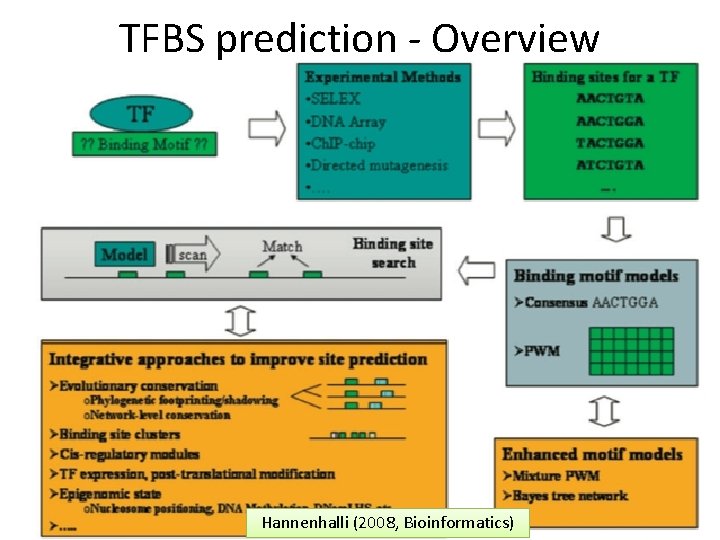
TFBS prediction - Overview Gene. Expression II BI 2010 Hannenhalli (2008, Bioinformatics) 6

De novo motif prediction • Search for over-represented motifs • Frequency count • Works well for yeast and prokaryotes • Not so successful in higher organisms Gene. Expression II BI 2010 7

Using motif databases • Search for known motifs • Position specific scoring matrix (PSSM) or Position weight matrix (PWM) • Databases: – Transfac – Jasper Gene. Expression II BI 2010 8

Phylogenetic-based methods • Search for islands of highly conserved regions • Footprinting: elements conserved across distant species • Shadowing: elements conserved between closely related species • Pros: increases specificity • Cons: conservation is not sufficient nor necessary Gene. Expression II BI 2010 9

Practical: • Try some tools on 2 kp upstream sequence of D. melanogaster eve and compare with published results. – Alibaba (de novo) – Match (Tranfac) – Meme (de novo) – Promo (Tranfac) – Weeder. H (phylogenetic footprinting) Gene. Expression II BI 2010 10

Other tools: • Many more tools available for download: – Sombrero – Foot. Printer – Phylo. Gibbs • Other Web-tools for groups of co-regulated genes: – RSAT – Nested. MICA – Web. MOTIFS Gene. Expression II BI 2010 11

TFBS prediction - Conclusion: • No single tool gives accurate results • Combination of predictions from multiple tools might increase specificity • Incorporate additional information for greater precision Gene. Expression II BI 2010 12

Microarrays - Overview • • • Gene. Expression II Introduction Data Generation Data Characteristics Diagnostic Plots Preprocessing Statistical Analysis BI 2010 13

What is a microarray? • A solid support onto which the sequences from thousands of different genes are immobilized • Different array supports - glass slide - nylon membrane - silicon chip • Different probe types - short oligonucleotides - long oligonucleotides - c. DNA • Each probe measures the expression of a single transcript Gene. Expression II BI 2010 14
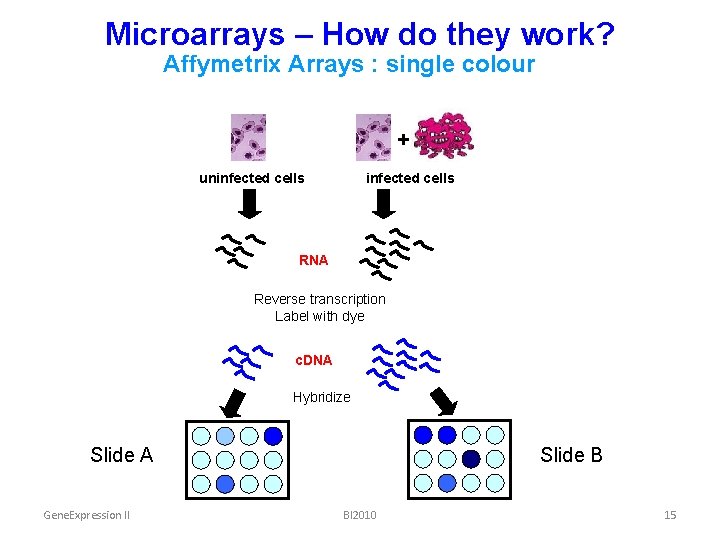
Microarrays – How do they work? Affymetrix Arrays : single colour + uninfected cells RNA Reverse transcription Label with dye c. DNA Hybridize Slide A Gene. Expression II Slide B BI 2010 15
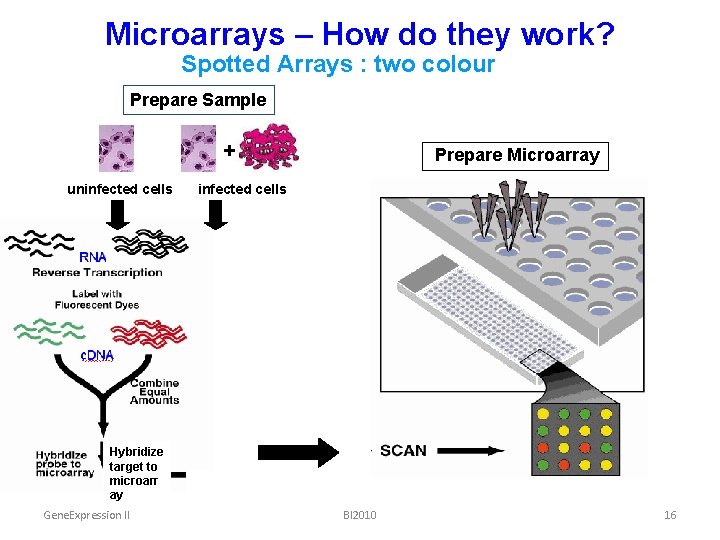
Microarrays – How do they work? Spotted Arrays : two colour Prepare Sample + uninfected cells Prepare Microarray infected cells Hybridize target to microarr ay Gene. Expression II BI 2010 16

Microarray: Subgrids • One pin per subgrid (print. Tip group, stratus) Gene. Expression II BI 2010 17

Microarrays – Data Extraction • How to get data from the slides into the computer? Gene. Expression II BI 2010 18

Data Extraction – Scanning Slide Scanner Images (TIFF) PRMS 02 -001 -S 100 CF 010 Gene. Expression II settings: - laser power - sensitivity - focus BI 2010 channel 1 (green) channel 2 (red) composite (green, yellow, red) 19

Data Extraction – Quantification align grid, tag unreliable spots Software: -Ima. Gene -Gene. Pix -Scan. Alyze. . . Gene. Expression II foreground (FG) background (BG) BI 2010 Data File Spot ID FG CH 1 BG CH 1 FG CH 2 BG CH 2 FL GFP 1241 6707 713 1 PA 0080 570 495 599 384 0 PA 0080 691 632 667 651 0 PA 0122 703 610 653 619 0 PA 0122 708 598 695 602 0 . . … … … program assigns numbers representing intensity of spot 20

Quantification: Intensity Range - area composed of pixel - value range: 0 – 216 - 1 - value range: 0 – 65535 - saturation possible - low intensities = noise Gene. Expression II BI 2010 21

Data Generation – Summary • • RNA labelling and hybridization Array Scanning One image per channel Load into quantification software Flag flawed spots Extract values Text file with FG and BG intensities (per probe) Gene. Expression II BI 2010 22
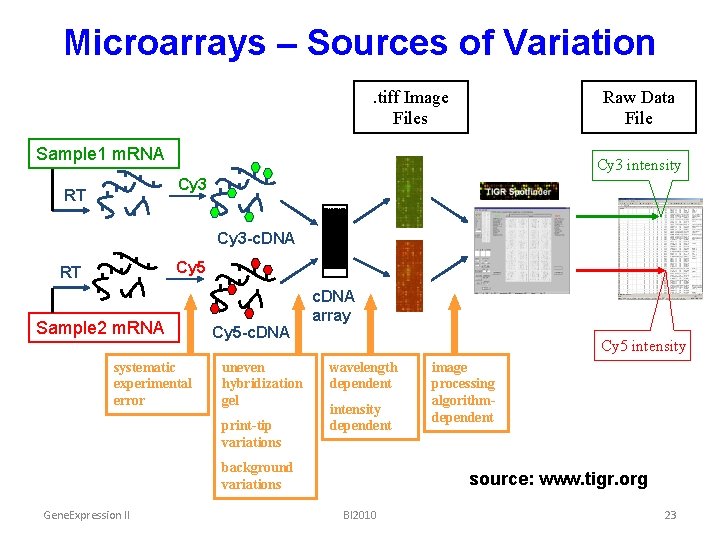
Microarrays – Sources of Variation. tiff Image Files Raw Data File Sample 1 m. RNA Cy 3 intensity Cy 3 RT Cy 3 -c. DNA Cy 5 RT Sample 2 m. RNA systematic experimental error Cy 5 -c. DNA uneven hybridization gel print-tip variations c. DNA array Cy 5 intensity wavelength dependent intensity dependent background variations Gene. Expression II image processing algorithmdependent source: www. tigr. org BI 2010 23

Microarrays – Sources of Variation • Technical: – labelling – hybridization – slide quality – scanning – print-tip effect – quantification – experimenter Gene. Expression II • Biological: – individual/strain/sample – environment – time point BI 2010 24

Microarrays – Data Characteristics • Intensities vs. ratios • Natural scale vs. log scale Gene. Expression II BI 2010 25
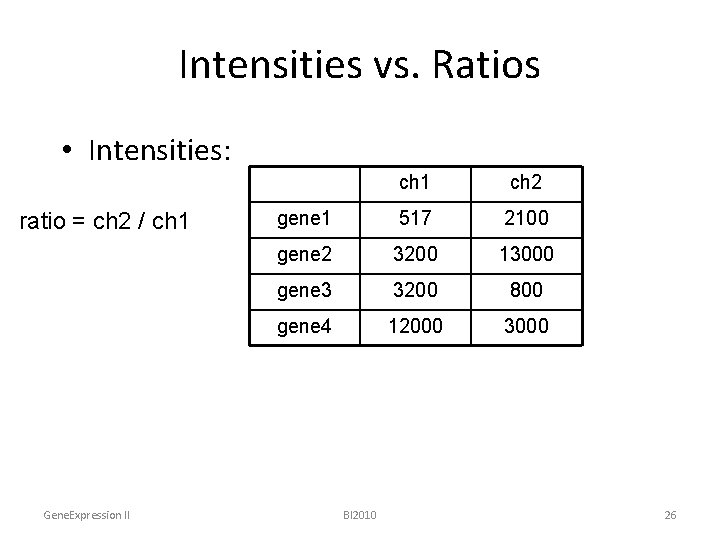
Intensities vs. Ratios • Intensities: ratio = ch 2 / ch 1 Gene. Expression II ch 1 ch 2 gene 1 517 2100 gene 2 3200 13000 gene 3 3200 800 gene 4 12000 3000 BI 2010 26
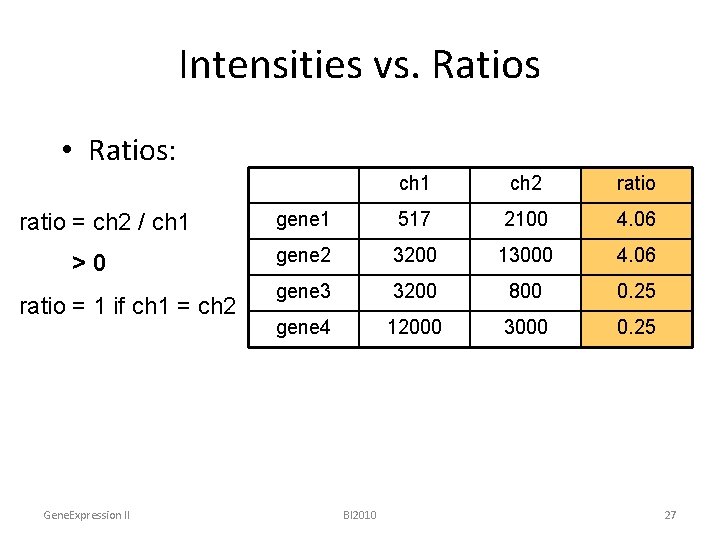
Intensities vs. Ratios • Ratios: ratio = ch 2 / ch 1 >0 ratio = 1 if ch 1 = ch 2 Gene. Expression II ch 1 ch 2 ratio gene 1 517 2100 4. 06 gene 2 3200 13000 4. 06 gene 3 3200 800 0. 25 gene 4 12000 3000 0. 25 BI 2010 27

Intensities vs. Ratios • Ratios – convey expression changes – hide base level differences • But: absolute changes can be important, too! Gene. Expression II BI 2010 28
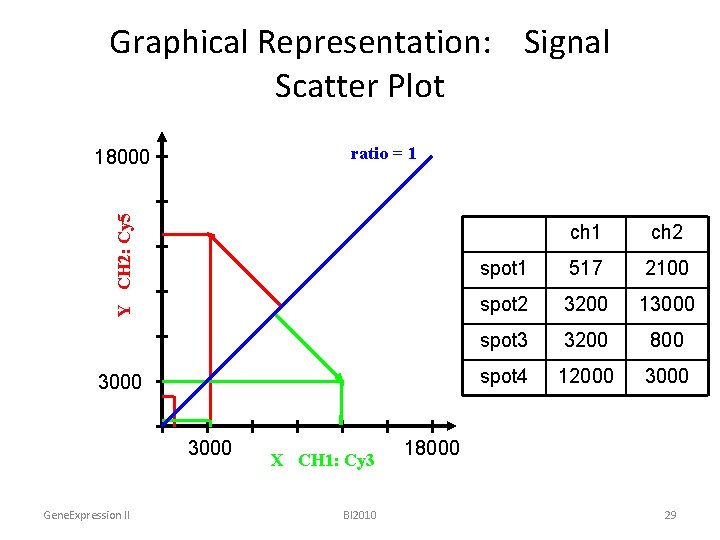
Graphical Representation: Signal Scatter Plot ratio = 1 Y CH 2: Cy 5 18000 3000 Gene. Expression II X CH 1: Cy 3 BI 2010 ch 1 ch 2 spot 1 517 2100 spot 2 3200 13000 spot 3 3200 800 spot 4 12000 3000 18000 29
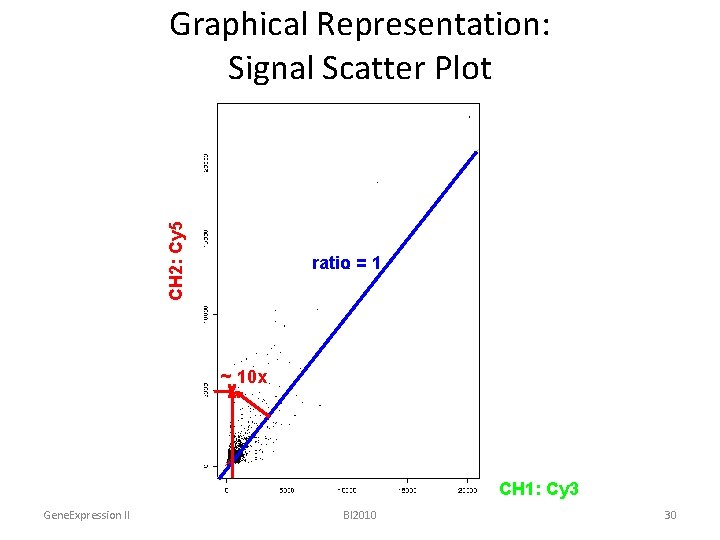
CH 2: Cy 5 Graphical Representation: Signal Scatter Plot ratio = 1 ~ 10 x CH 1: Cy 3 Gene. Expression II BI 2010 30
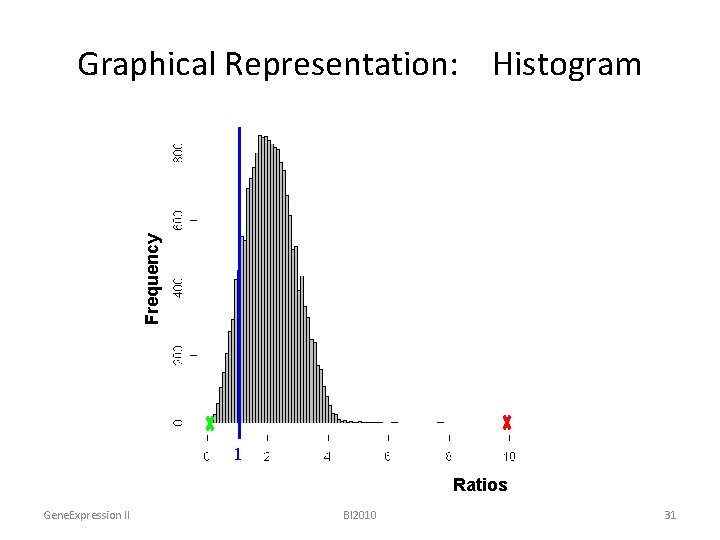
Frequency Graphical Representation: Histogram ratios 1 Ratios Gene. Expression II BI 2010 31

Raw vs. Log ratios x = 2 y • Log transformation ratios x = basey raw log 8 = 23 0. 1 -3. 3 0. 125 = 2 -3 0. 5 -1 1 0 2 1 10 3. 3 y undefined for x <= 0 Gene. Expression II BI 2010 32
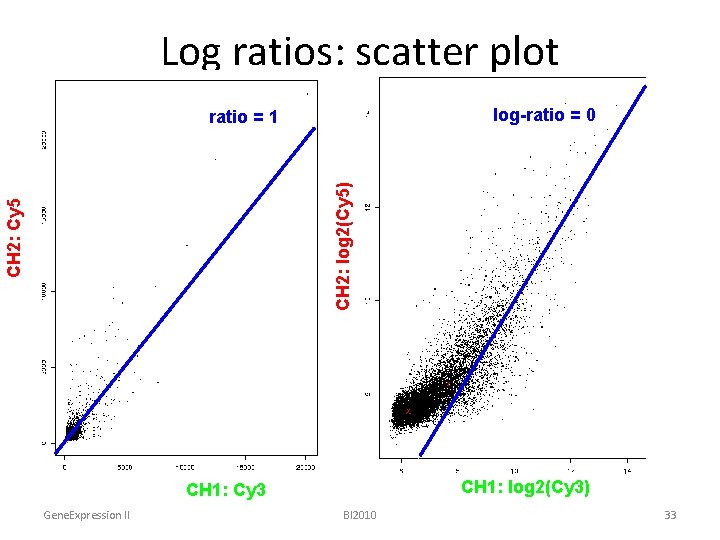
Log ratios: scatter plot log-ratio = 0 CH 2: Cy 5 CH 2: log 2(Cy 5) ratio = 1 CH 1: log 2(Cy 3) CH 1: Cy 3 Gene. Expression II BI 2010 33
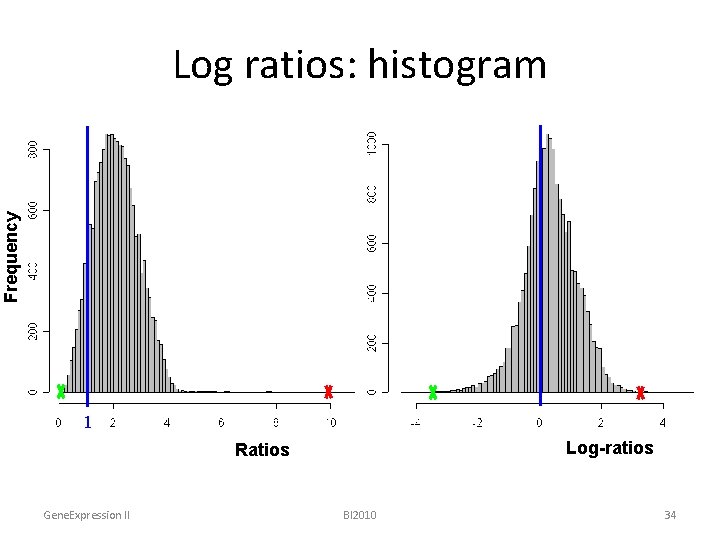
Frequency Log ratios: histogram ratios 1 Log-ratios Ratios Gene. Expression II BI 2010 34

Microarrays – Data Characteristics • ratios vs. intensities – convey expression changes – hide base level differences • log ratios vs. raw ratios – reduce spread – provide symmetry Gene. Expression II BI 2010 35

Diagnostic plots • • • Gene. Expression II histogram scatter plot box plot MA plot chip visualization BI 2010 36
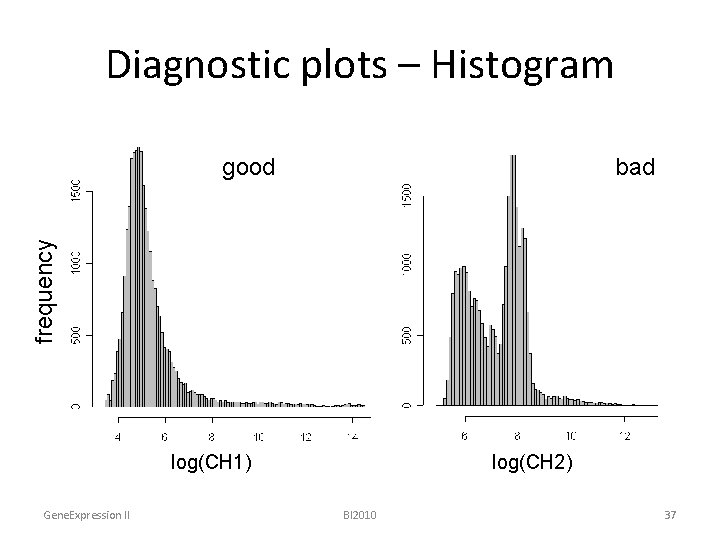
Diagnostic plots – Histogram bad frequency good log(CH 1) Gene. Expression II log(CH 2) BI 2010 37
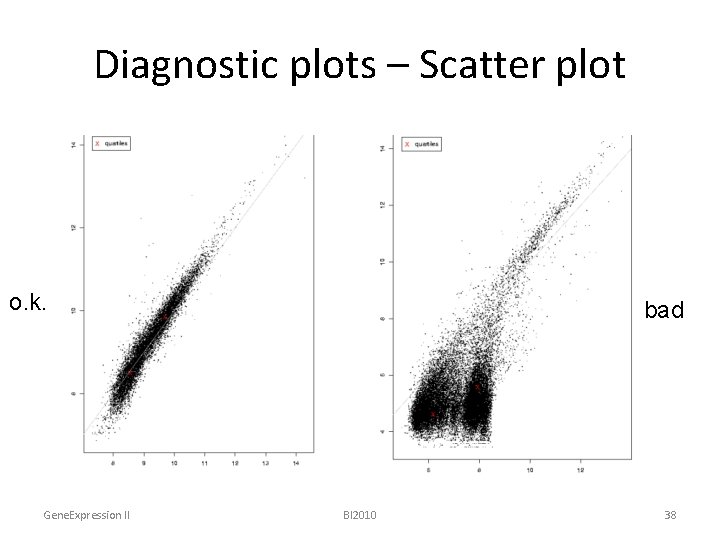
Diagnostic plots – Scatter plot o. k. Gene. Expression II bad BI 2010 38
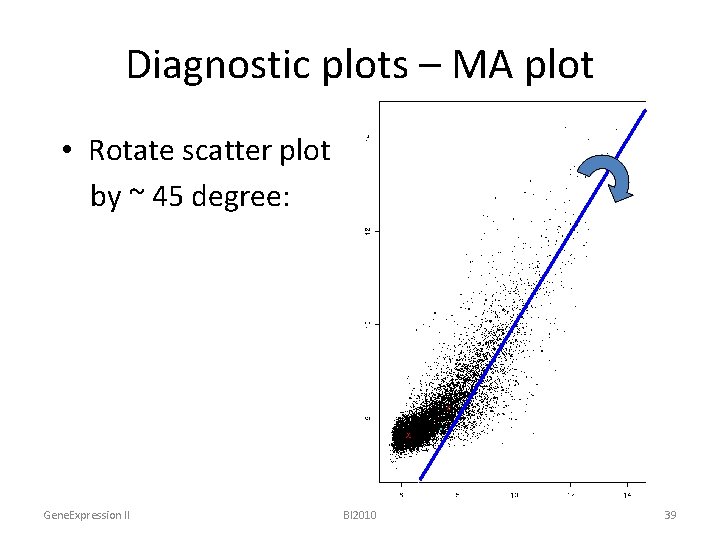
Diagnostic plots – MA plot • Rotate scatter plot by ~ 45 degree: Gene. Expression II BI 2010 39
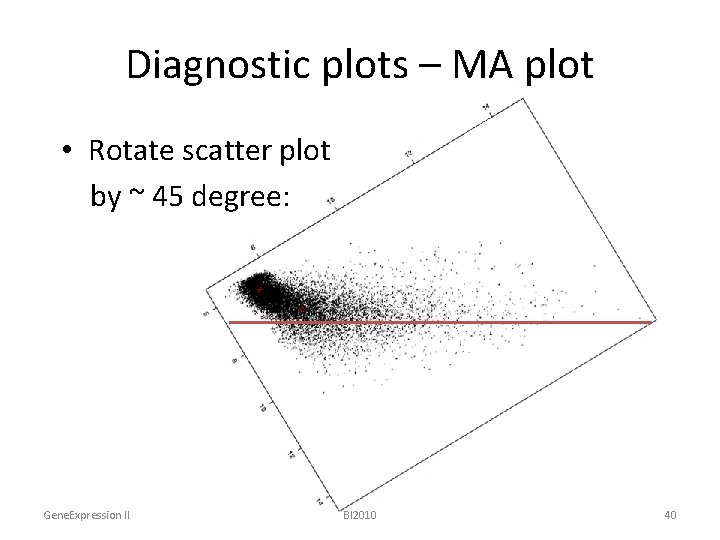
Diagnostic plots – MA plot • Rotate scatter plot by ~ 45 degree: Gene. Expression II BI 2010 40

Diagnostic plots – MA plot • Mathematically: Minus = log 2(R) – log 2(G) = 0. 5 * ( log 2(R) + log 2(G) ) Addition Gene. Expression II BI 2010 41
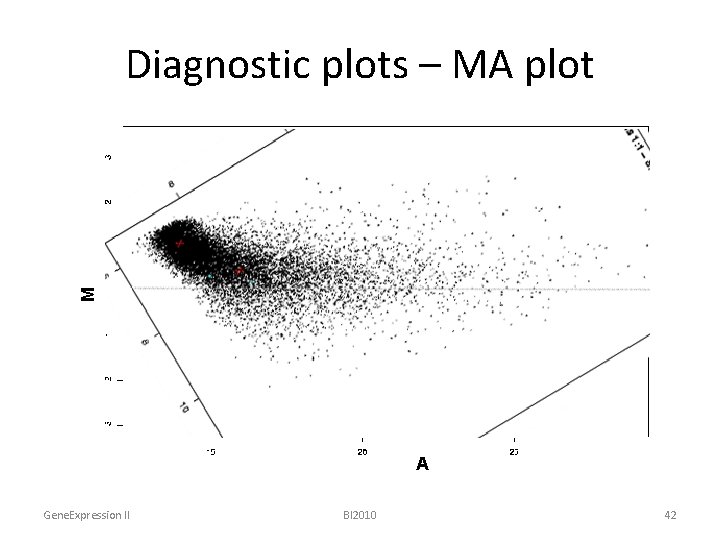
M Diagnostic plots – MA plot A Gene. Expression II BI 2010 42
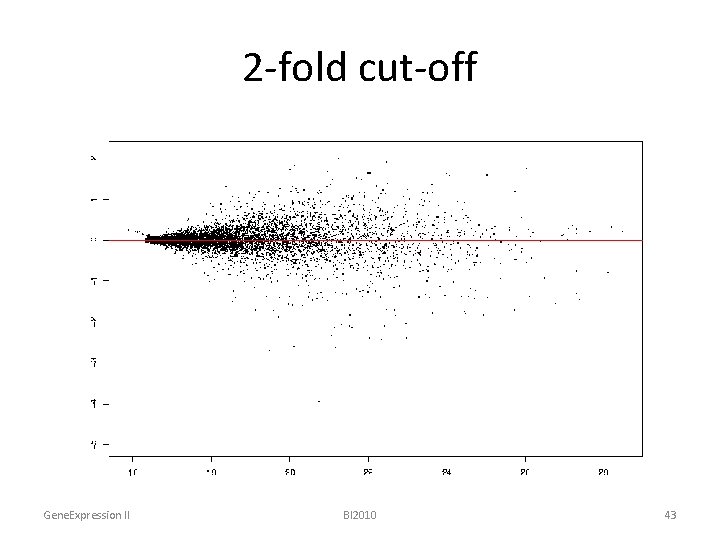
2 -fold cut-off Gene. Expression II BI 2010 43
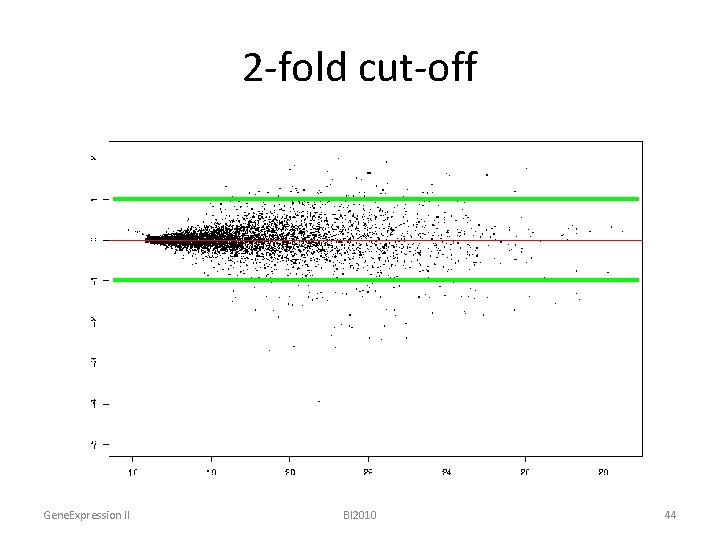
2 -fold cut-off Gene. Expression II BI 2010 44
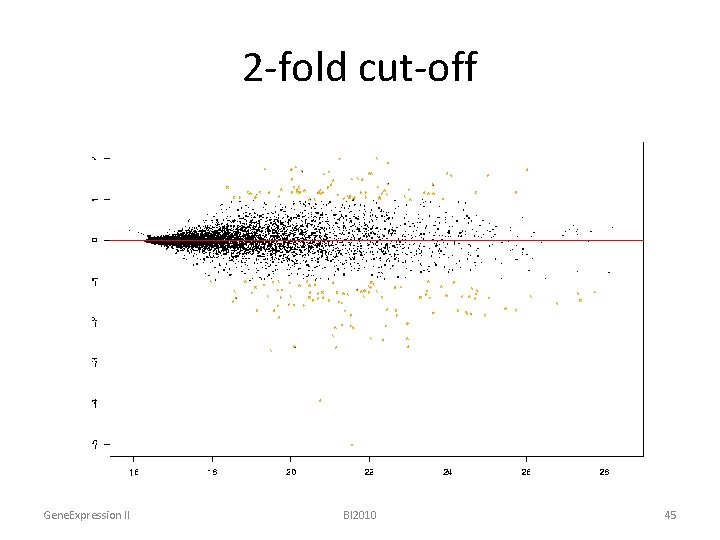
2 -fold cut-off Gene. Expression II BI 2010 45
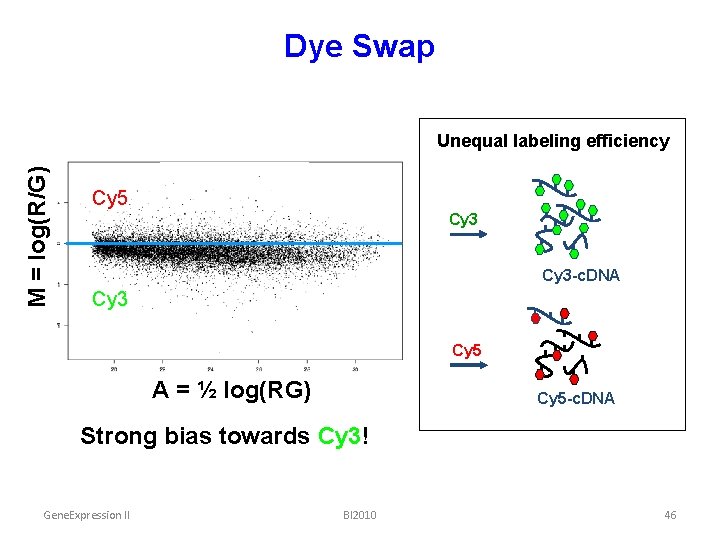
Dye Swap M = log(R/G) Unequal labeling efficiency Cy 5 Cy 3 -c. DNA Cy 3 Cy 5 A = ½ log(RG) Cy 5 -c. DNA Strong bias towards Cy 3! Gene. Expression II BI 2010 46
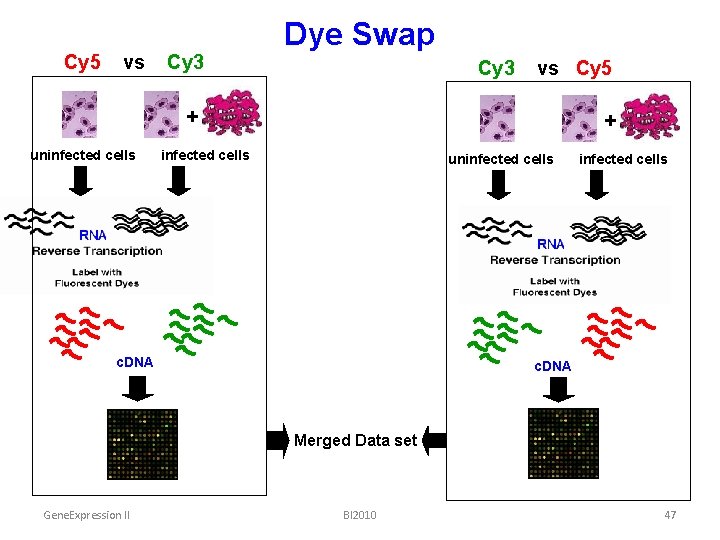
Cy 5 vs Cy 3 Dye Swap Cy 3 vs Cy 5 + uninfected cells + infected cells uninfected cells c. DNA Merged Data set Gene. Expression II BI 2010 47
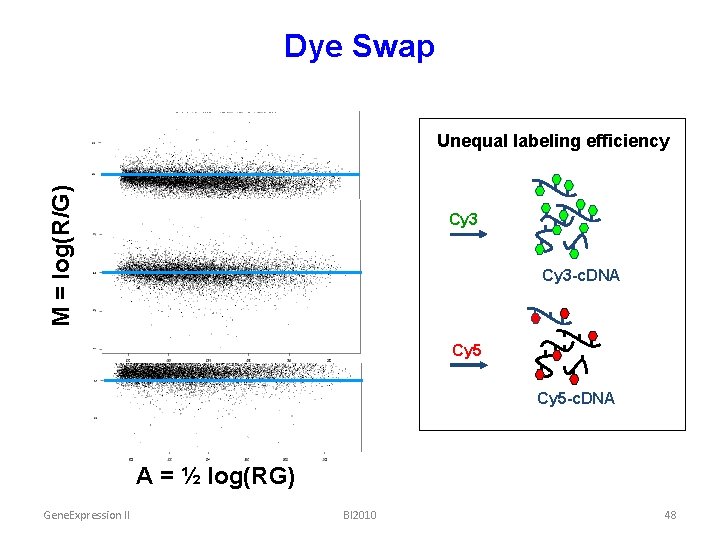
Dye Swap M = log(R/G) Unequal labeling efficiency Cy 3 -c. DNA A = ½ log(RG) Cy 5 -c. DNA A = ½ log(RG) Gene. Expression II BI 2010 48
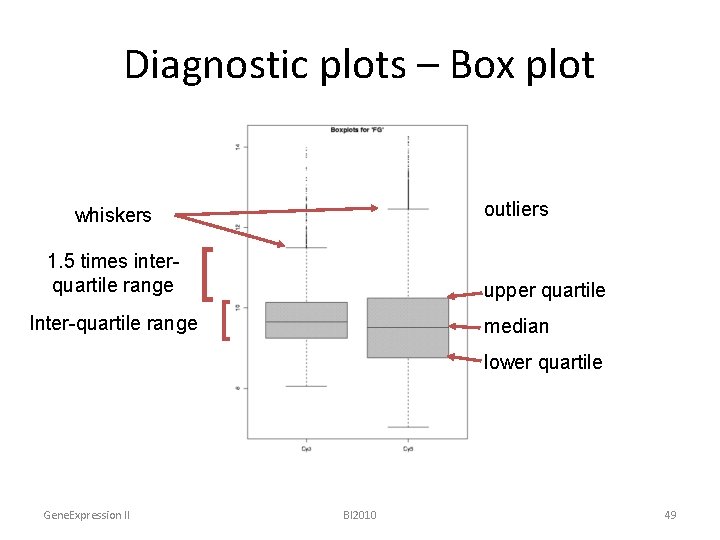
Diagnostic plots – Box plot outliers whiskers 1. 5 times interquartile range Inter-quartile range [ upper quartile [ median lower quartile Gene. Expression II BI 2010 49
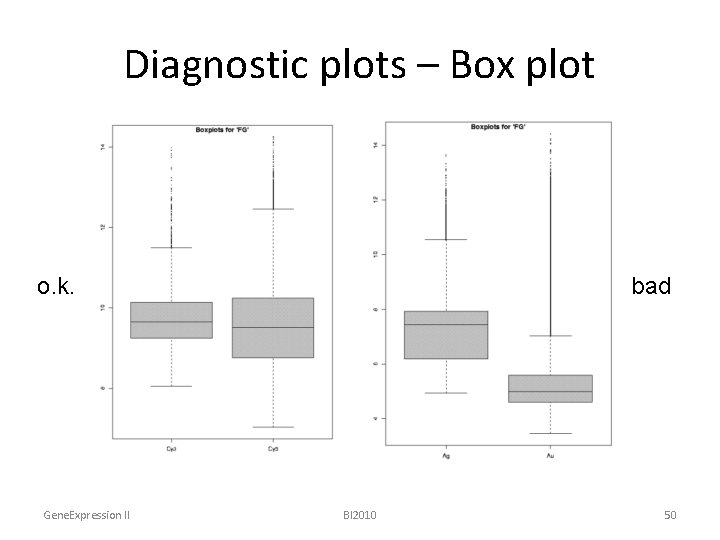
Diagnostic plots – Box plot o. k. Gene. Expression II bad BI 2010 50
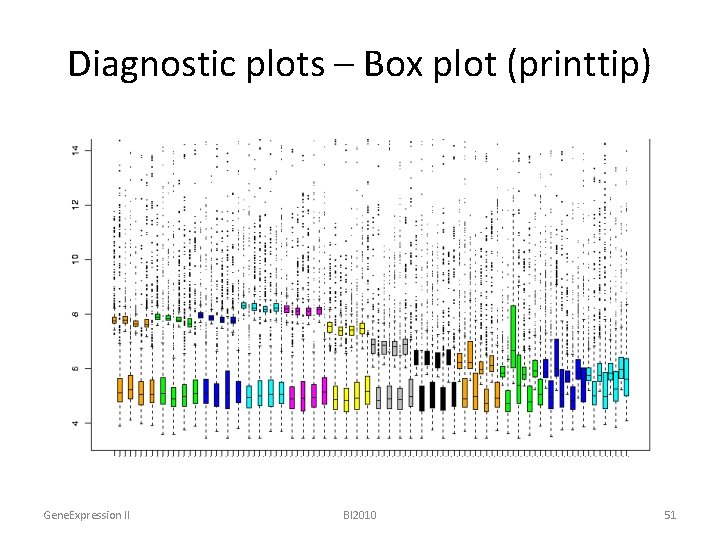
Diagnostic plots – Box plot (printtip) Gene. Expression II BI 2010 51
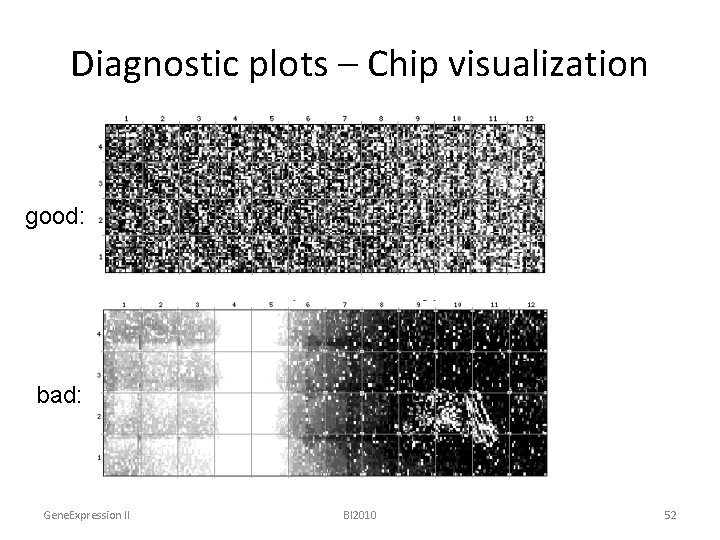
Diagnostic plots – Chip visualization good: bad: Gene. Expression II BI 2010 52

Diagnostic plots: Summary • histogram – data distribution (intensities, ratios) • scatter plot – dye effect, print-tip effect • box plot – equal average ratio and distribution, print-tip effect • MA plot – dye effect and intensity-dependant ratio • chip visualization – spatial bias, scratches, bubbles, smears Gene. Expression II BI 2010 53

Microarrays – Preprocessing • • Flagging Background correction Normalization Flawed slides: Discard and repeat Gene. Expression II BI 2010 54
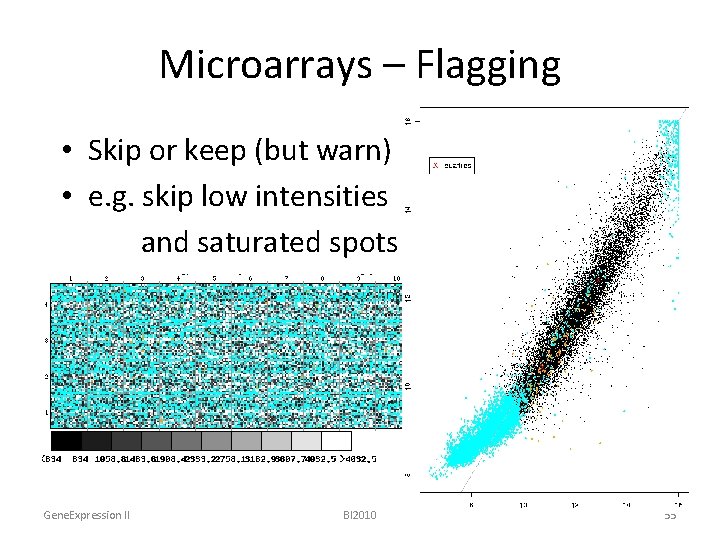
Microarrays – Flagging • Skip or keep (but warn) • e. g. skip low intensities and saturated spots Gene. Expression II BI 2010 55

Microarrays – Background correction • Subtract background measurements from foreground intensities • Brings intensities lower to zero, increases ratios: example spot with five fold upregulation: 500 / 100 = 5 subtract background (50) from both channels 450 / 50 = 9 • Additional source of variance! Gene. Expression II BI 2010 56

Microarrays – Normalization • Remove effect from intensities, dye bias, spatial bias or print-tip variations: – Global mean, median – Loess, lowess – Print-tip loess – 2 D loess – Variance stabilazation (VSN) Gene. Expression II BI 2010 57
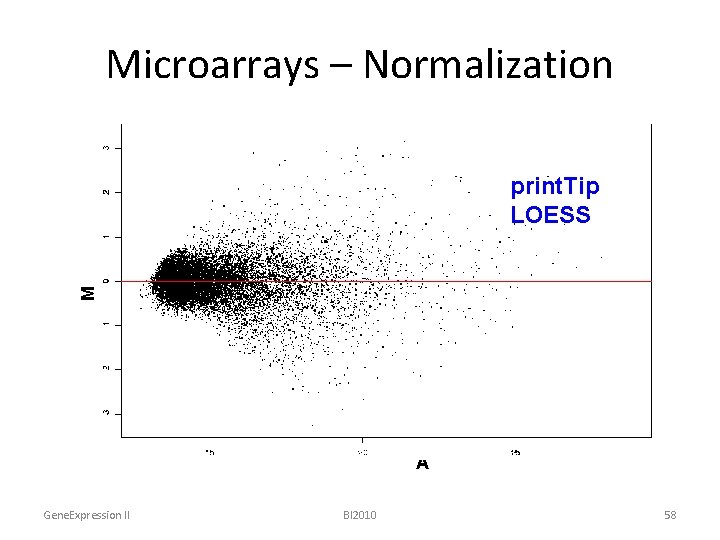
Microarrays – Normalization M Global rawmean LOESS print. Tip LOESS A Gene. Expression II BI 2010 58
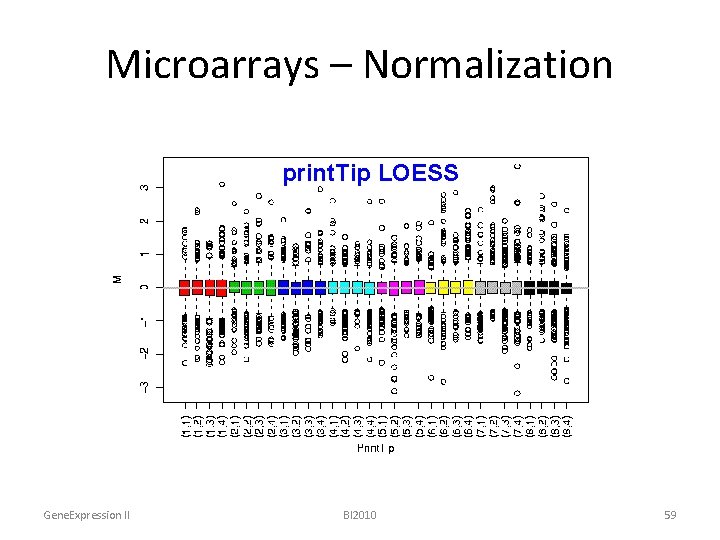
Microarrays – Normalization print. Tip global LOESS raw LOESS mean Gene. Expression II BI 2010 59

Microarrays – Discard and repeat • Some slides turn out to be uncorrectable and need to be repeated (unless a sufficient number of replicates remains). • Remember: bad data in = bad data out! Gene. Expression II BI 2010 60

Microarrays – Statistical Analysis • • • Gene. Expression II Replicates Variation t-tests multiple-testing correction� gene lists BI 2010 61

Statistical Analysis – Replicates • Two types of repeats • Technical: – multiple copies of probes on array – multiple repeats of hybridiztion (same RNA) • Biological: – multiple hybridizations with RNA from multiple extractions Need replicates to measure variation! Gene. Expression II BI 2010 62

Statistical Analysis – Variation • Biological variation different from technical • Statistically incorrect to mix • Important consideration for repeats: High confidence in results for a) one sample/patient/colony b) group of samples/patients/colonies Prioritise biological repeats! Gene. Expression II BI 2010 63

Statistical Analysis – t-tests Different classes of samples: - find genes that are affected by a treatment - p-value = degree of evidence - H 0: expression does not change - t-test requires at least 2 replicates provides p-value for each gene Gene. Expression II BI 2010 64

Statistical Analysis – multiple-testing correction Carrying out t-tests on 10, 000 genes average of 500 will have p-value <= 0. 05 Methods for multiple testing: Bonferroni (very strict) Benjamini-Hochberg false-discovery rate (FDR) Gene. Expression II BI 2010 65

Statistical Analysis – Gene lists • List of good candidate genes to follow up • FP vs FN • Fold-change vs p-value Choice depends on downstream analysis Input for downstream analysis: Clustering, pathway analysis, enrichment, etc. Gene. Expression II BI 2010 66

Analysis tools • Stand-alone tools: – – – R Bio. Conductor Array. Norm TM 4 Gene. Spring (commercial) • Web-based tools – – – Gene. Expression II Array. Pipe Express. Yourself Gene. Publisher GEPAS Gene. Traffic (commercial) BI 2010 67

Public Repositories • Array. Express – EBI, MIAME-compliant • Gene Expression Omnibus (GEO) – NCBI – „world‘s first write-only database“ Gene. Expression II BI 2010 68

Summary • Many sources of variance • Large numbers of replicates required for reliable results • Data: be aware of flaws/bias • Flagging/discarding results in data loss • Correction often possible but can insert artifacts • However: Microarrays can still help making great discoveries! Gene. Expression II BI 2010 69

END Gene. Expression II BI 2010 70
- Slides: 70