Gene Expression and Mutations Bacterial control of gene
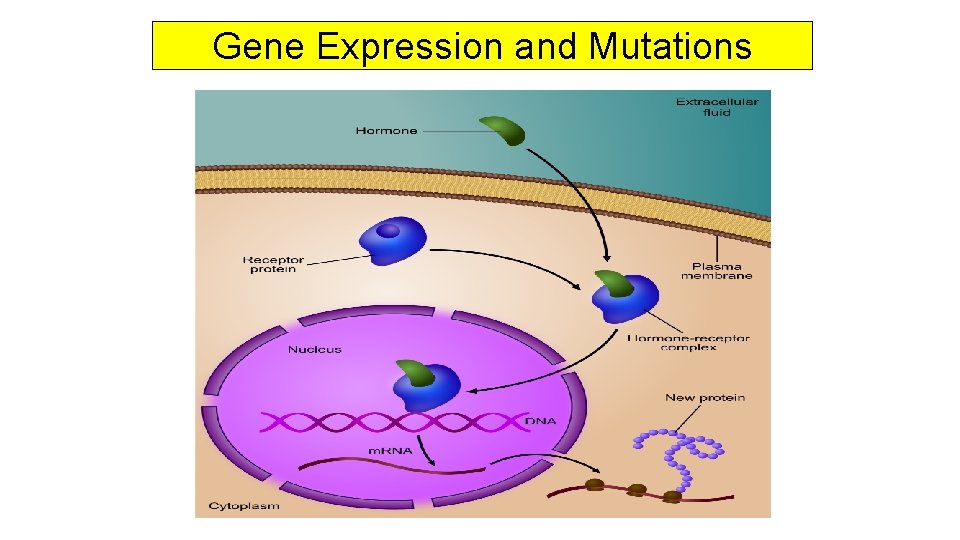
Gene Expression and Mutations
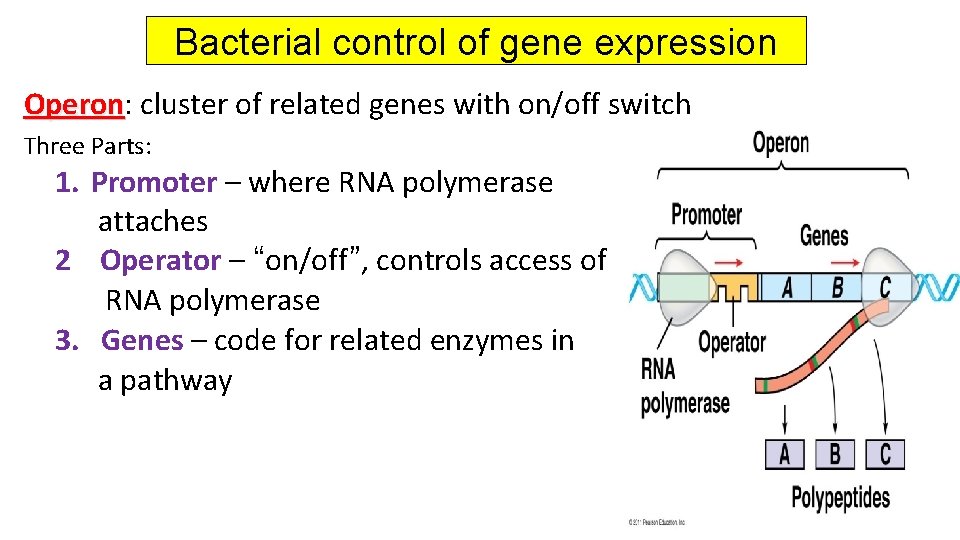
Bacterial control of gene expression Operon: Operon cluster of related genes with on/off switch Three Parts: 1. Promoter – where RNA polymerase attaches 2 Operator – “on/off”, controls access of RNA polymerase 3. Genes – code for related enzymes in a pathway
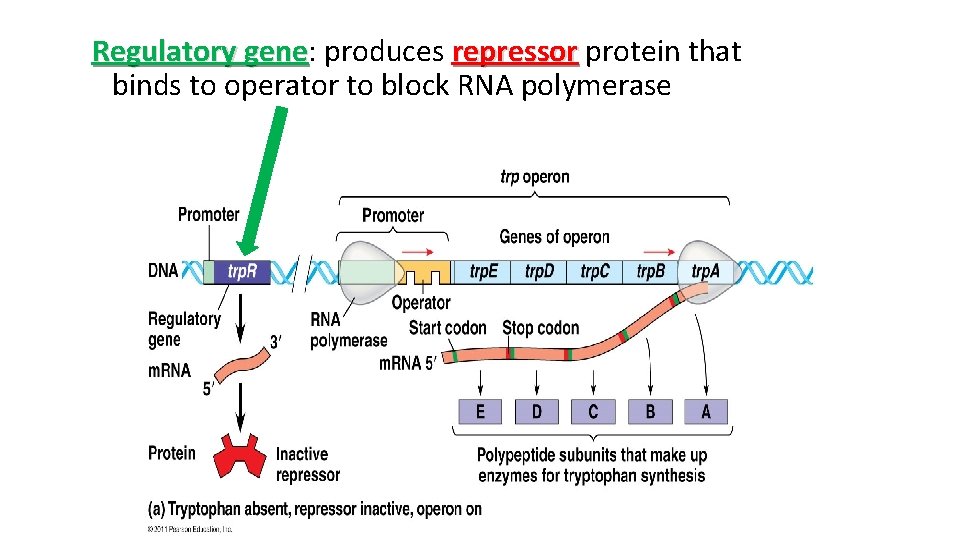
Regulatory gene: gene produces repressor protein that binds to operator to block RNA polymerase
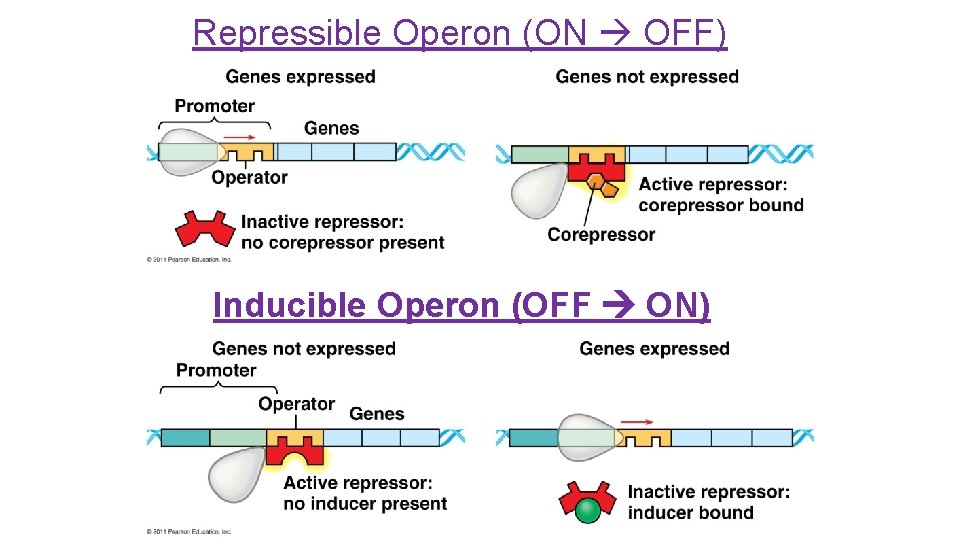
Repressible Operon (ON OFF) Inducible Operon (OFF ON)
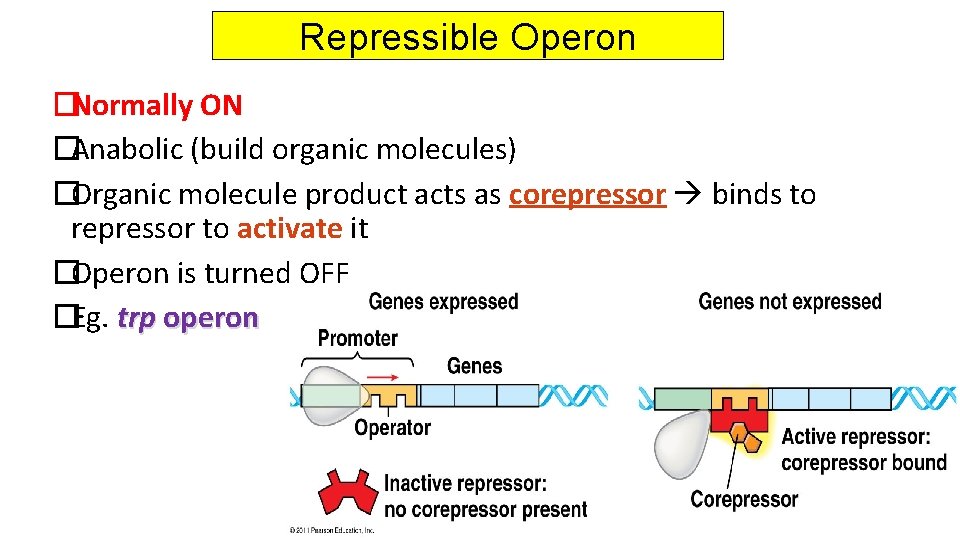
Repressible Operon �Normally ON �Anabolic (build organic molecules) �Organic molecule product acts as corepressor binds to repressor to activate it �Operon is turned OFF �Eg. trp operon
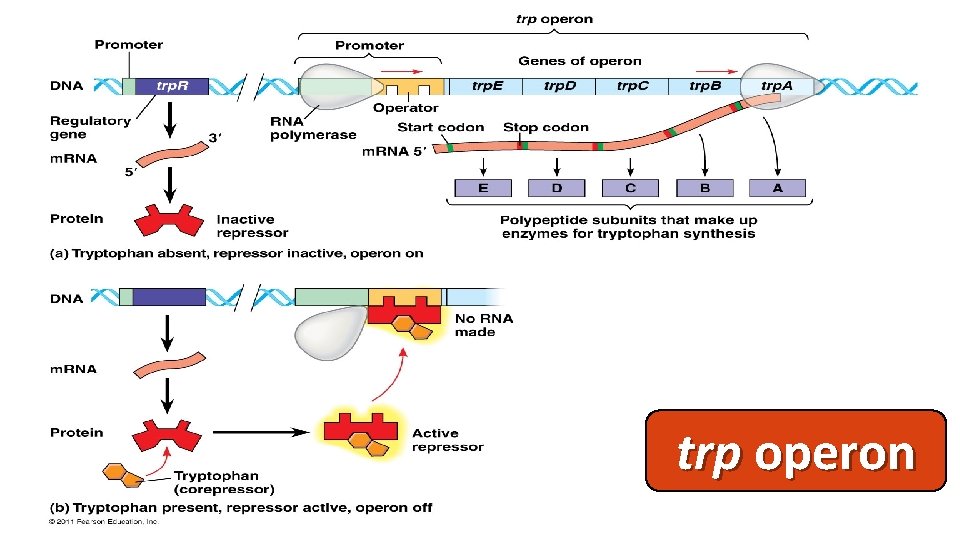
trp operon
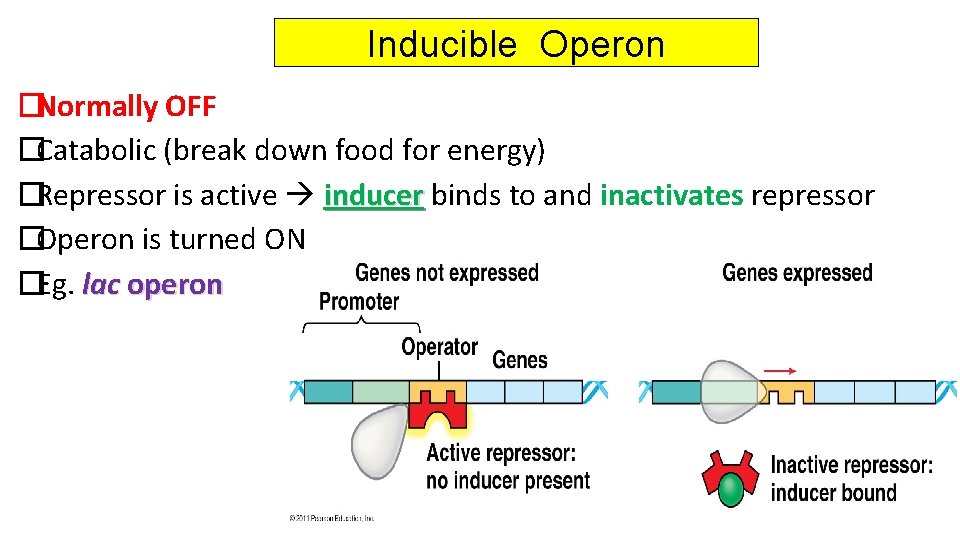
Inducible Operon �Normally OFF �Catabolic (break down food for energy) �Repressor is active inducer binds to and inactivates repressor �Operon is turned ON �Eg. lac operon
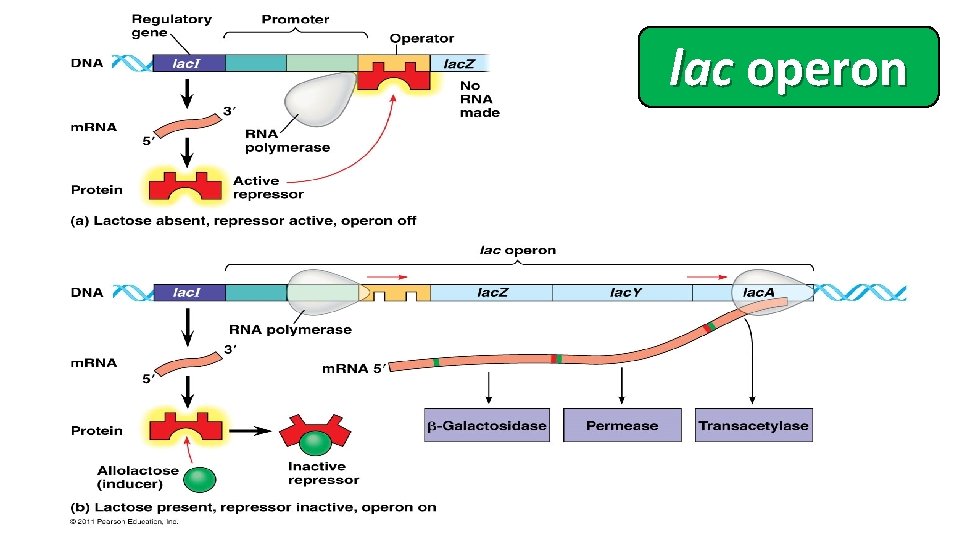
lac operon

Positive or Negative Control • Negative control: operons are switched off by active form of repressor protein • Eg. trp operon, lac operon • Positive control: regulatory protein interacts directly with genome to increase transcription • Eg. c. AMP & CAP
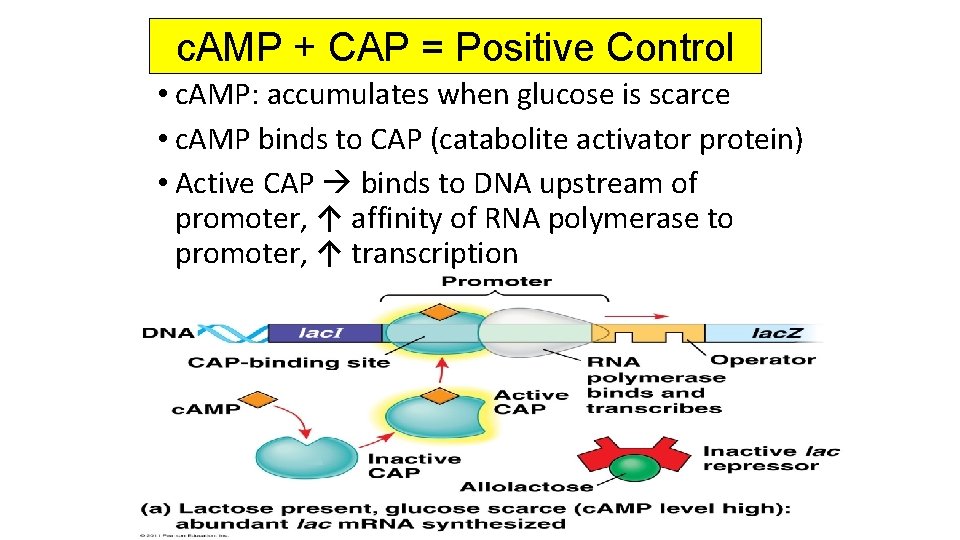
c. AMP + CAP = Positive Control • c. AMP: accumulates when glucose is scarce • c. AMP binds to CAP (catabolite activator protein) • Active CAP binds to DNA upstream of promoter, ↑ affinity of RNA polymerase to promoter, ↑ transcription

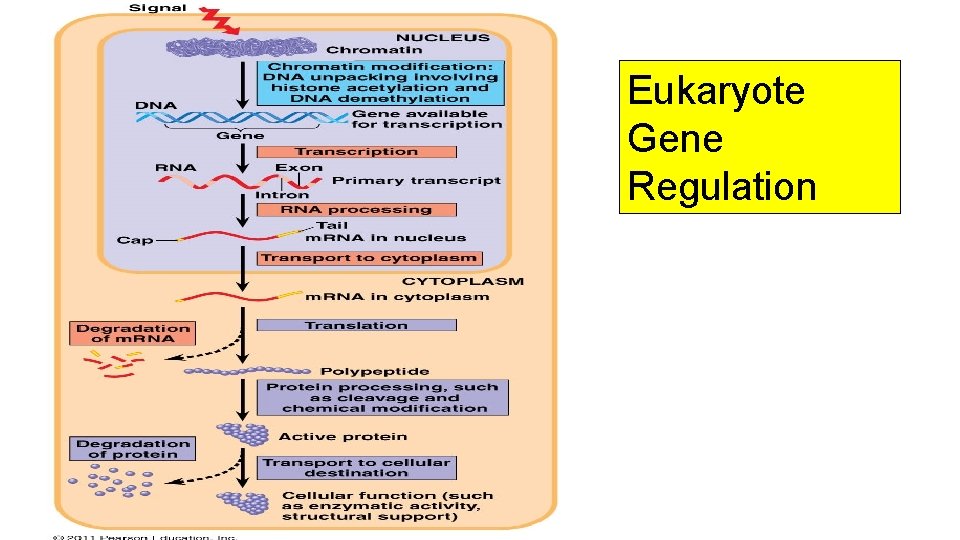
Eukaryote Gene Regulation

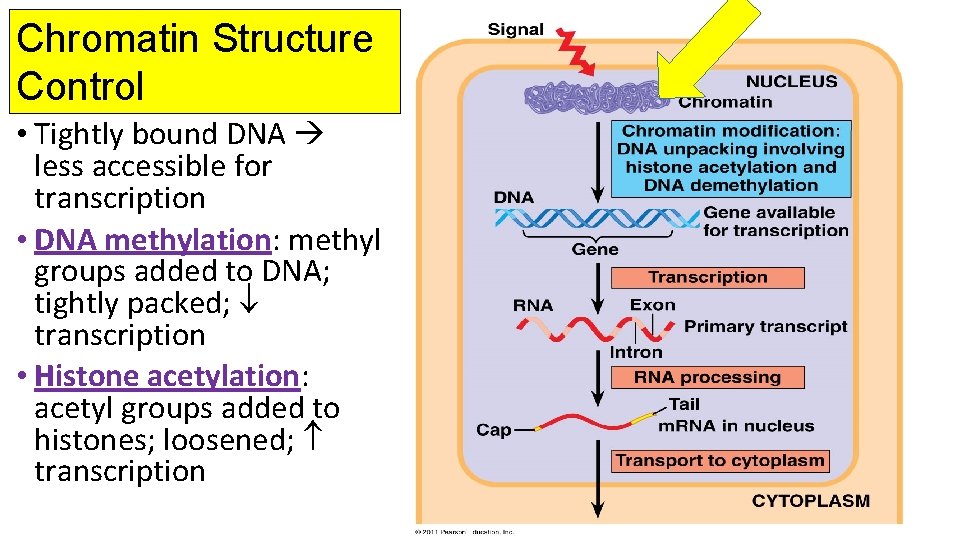
Chromatin Structure Control • Tightly bound DNA less accessible for transcription • DNA methylation: methyl groups added to DNA; tightly packed; transcription • Histone acetylation: acetyl groups added to histones; loosened; transcription

Histone Acetylation Loosens chromatin to allow transcription to take place
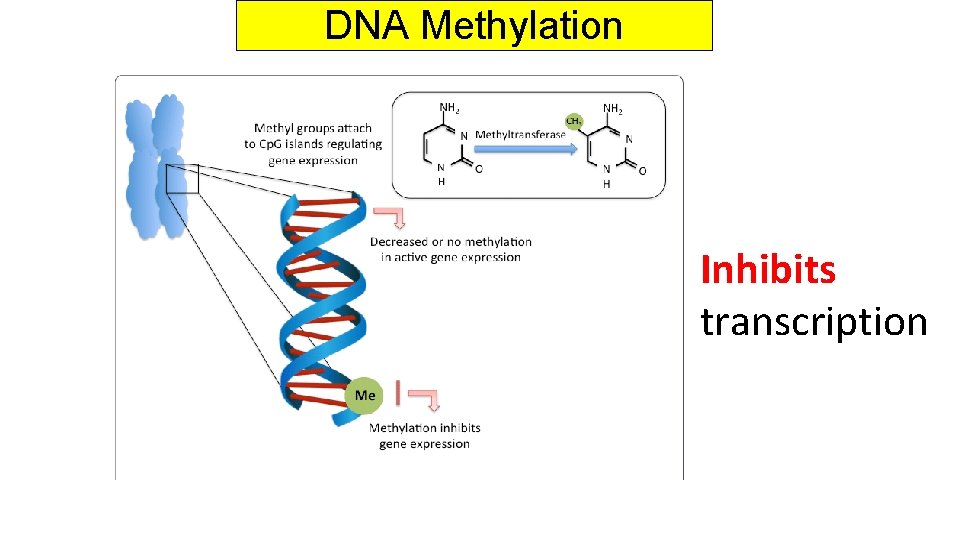
DNA Methylation Inhibits transcription
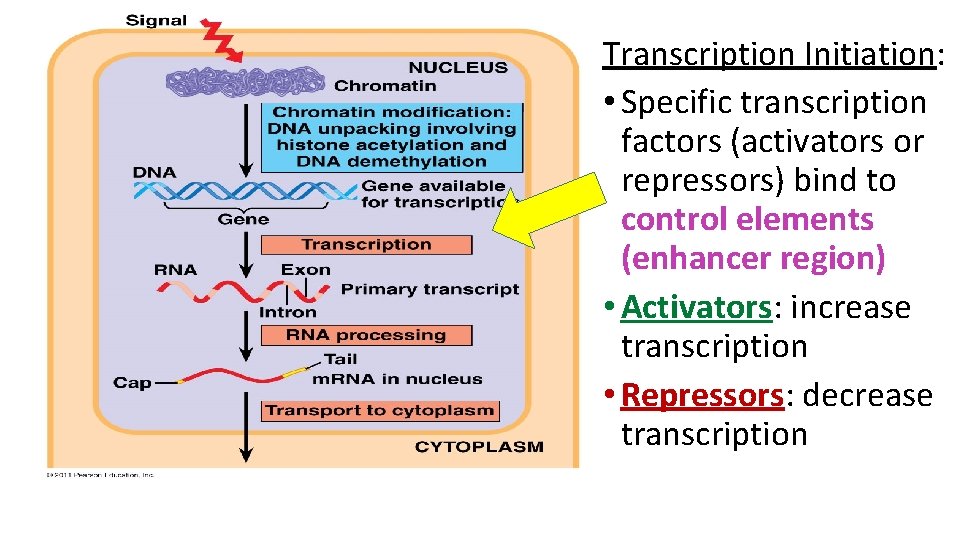
Transcription Initiation: • Specific transcription factors (activators or repressors) bind to control elements (enhancer region) • Activators: increase transcription • Repressors: decrease transcription
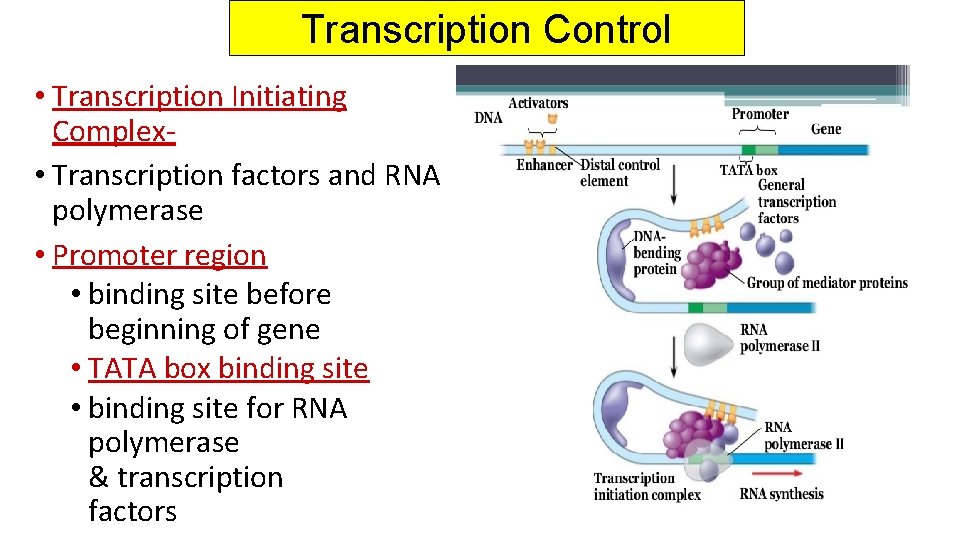
Transcription Control • Transcription Initiating Complex • Transcription factors and RNA polymerase • Promoter region • binding site before beginning of gene • TATA box binding site • binding site for RNA polymerase & transcription factors
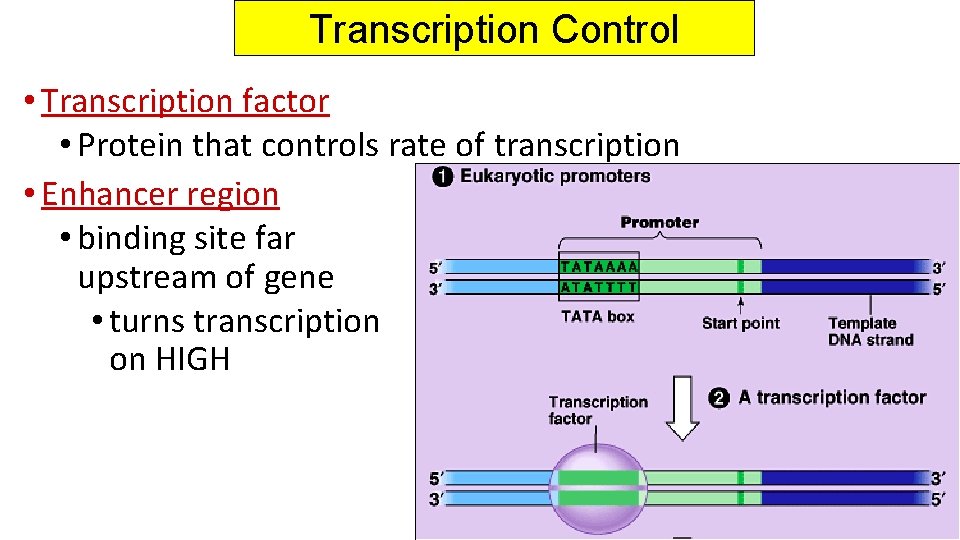
Transcription Control • Transcription factor • Protein that controls rate of transcription • Enhancer region • binding site far upstream of gene • turns transcription on HIGH
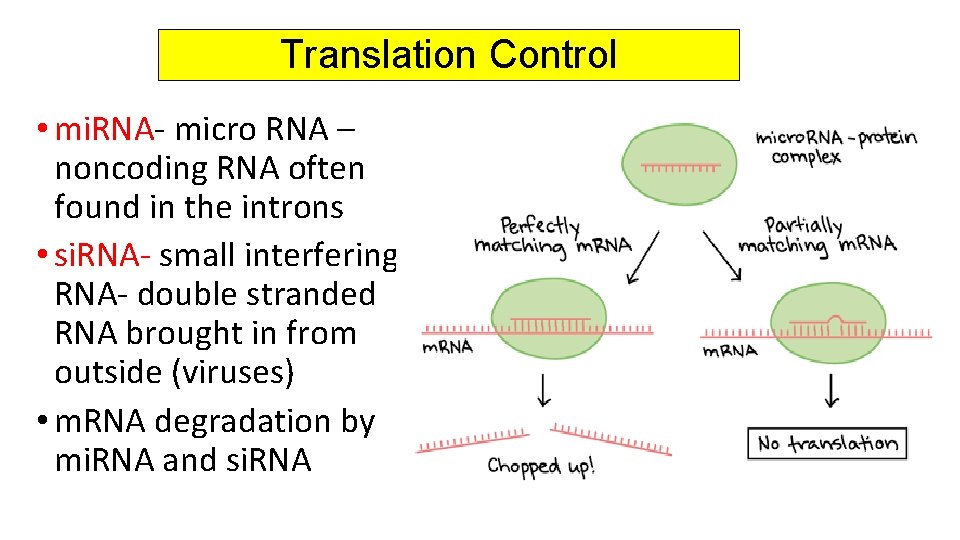
Translation Control • mi. RNA- micro RNA – noncoding RNA often found in the introns • si. RNA- small interfering RNA- double stranded RNA brought in from outside (viruses) • m. RNA degradation by mi. RNA and si. RNA

Epigenetic Inheritance • Modifications on chromatin can be passed on to future generations • Unlike DNA mutations, these changes to chromatin can be reversed (de-methylation of DNA) • Explains differences between identical twins • Eg. DNA methylation (gene silencing), histone acetylation, X chromosome inactivation, heterochromatin (silent chromatin) https: //www. eurekalert. org/pub_releases/2016 -01/cpedw 012016. php
Mutations

Point Mutations- affecting only one or a few bases • Base pair substitution- a single base is substituted for another • Silent – because of wobble, has no effect • Missense- altered codon still codes for an amino acid but not the right one • Nonsense- substitution changes the codon to a stop codon (worst mutation) • Insertion or deletions cause frameshift mutations when the reading of the codons changes in base sequences which are no longer in groups of three
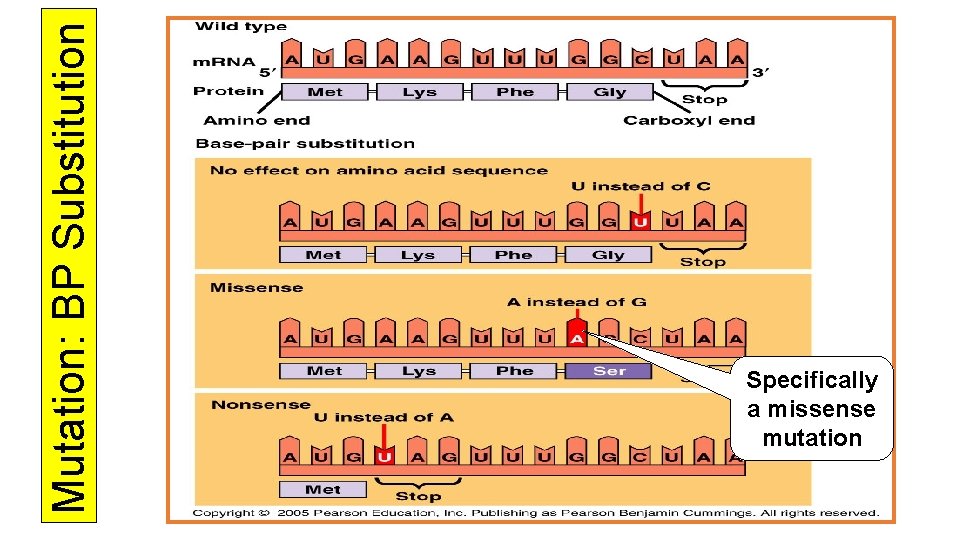
Mutation: BP Substitution Specifically a missense mutation
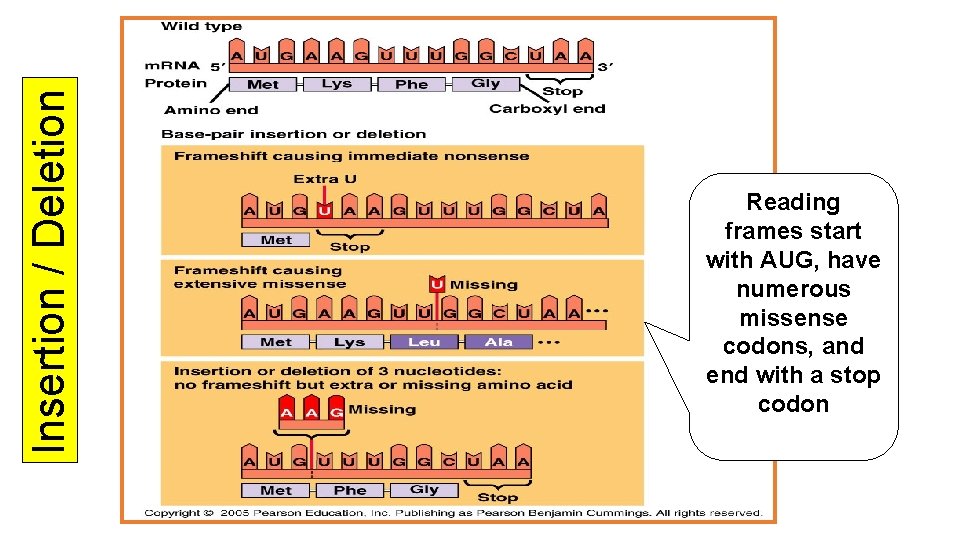
Insertion / Deletion Reading frames start with AUG, have numerous missense codons, and end with a stop codon
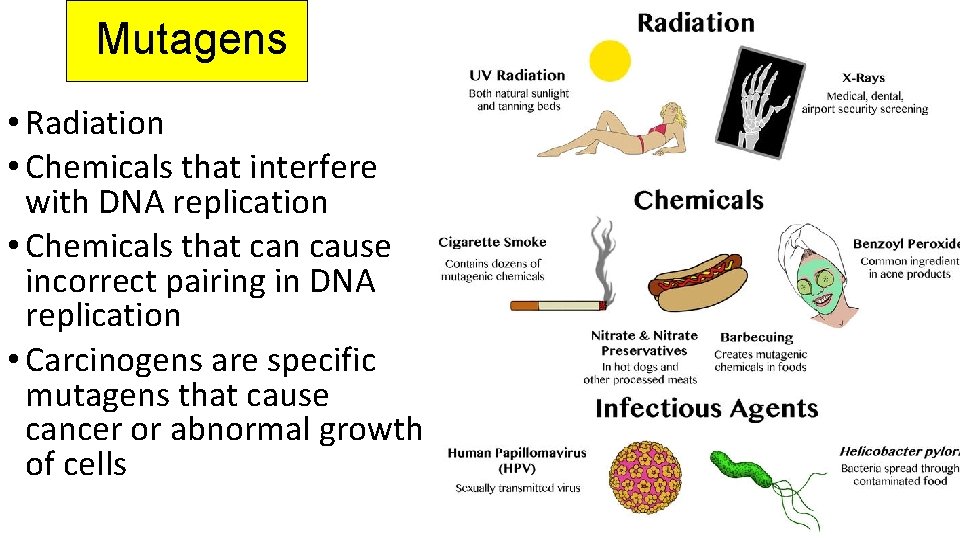
Mutagens • Radiation • Chemicals that interfere with DNA replication • Chemicals that can cause incorrect pairing in DNA replication • Carcinogens are specific mutagens that cause cancer or abnormal growth of cells
- Slides: 26