GAlib A C Library of Genetic Algorithm Components

GAlib A C++ Library of Genetic Algorithm Components Vanessa Herves Gómez vherves@fi. upm. es Department of Computer Architecture and Technology, Technical University of Madrid, Spain

Types of genetic algorithms (GAs) n The library contains four types of genetic algorithms: Simple genetic algorithm n Steady state genetic algorithm n Incremental genetic algorithm n Deme genetic algorithm. n
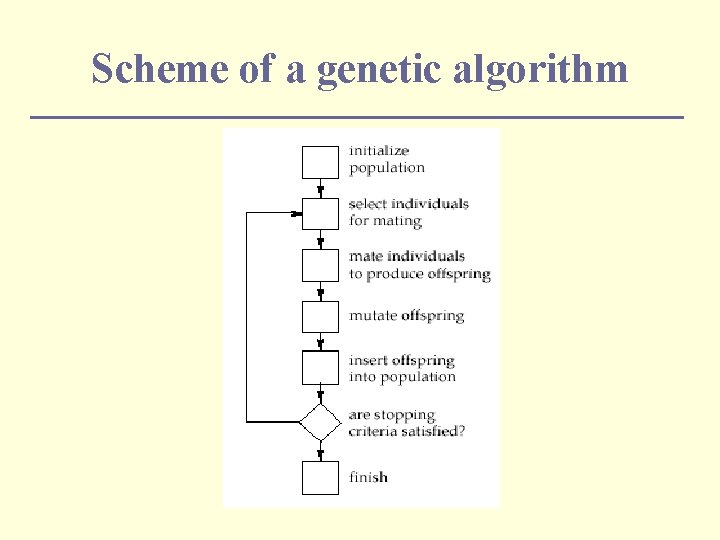
Scheme of a genetic algorithm

Resolve a problem with GAs n In order to solve a problem using a genetic algorithm you must do three things: Define a representation n Define the genetic operators n Define the objective function → Defined by you n n To generate new individuals the genetic algorithm uses: Genetic operators n Selection/replacement strategies n

Representation n A single solution to the problem is represented in a single data structure called genome. n The library contains four types of genomes: GAList. Genome n GATree. Genome n GAArray. Genome n GABinary. String. Genome n

The Genetic Operators n Built into the genome. n Each genome have three primary operators: n Initialization n Mutation n Crossover n GAlib comes whith these operators pre-defined for each genome type. n You can customize any of them. n Each genome must also contain an objective function and may also contain a comparator.

Selection strategies n Is defined in the population. n It is use by the genetic algorithms to choose which individuals should mate. n The library contains the following selection strategies: n n n GARank. Selector GARoulette. Wheel. Selector GATournament. Selector GADSSelector GASRSSelector GAUniform. Selector

Objective Functions and Fitness Scaling (I) n Given a single solution to a problem, the objective function provides a measure of how good is it. n The objective score is the value returned by your objective function. n The fitness score is a possibly-transformed rating used by the GA to determinate the fitness of individuals for mating. n The genetic algoritm uses the fitness score to do selection.

Objective Functions and Fitness Scaling (II) n GAlib contains the following scaling scheme: n GANo. Scaling n GALinear. Scaling n GASigma. Truncation. Scaling n Ga. Power. Law. Scaling n GASharing

Stopping criterion n You must tell the algorithm when to stop. n The library contains the following stopping criterion: Terminate. Upon. Generation n Terminate. Upon. Convergence n Terminate. Upon. Pop. Convergence n
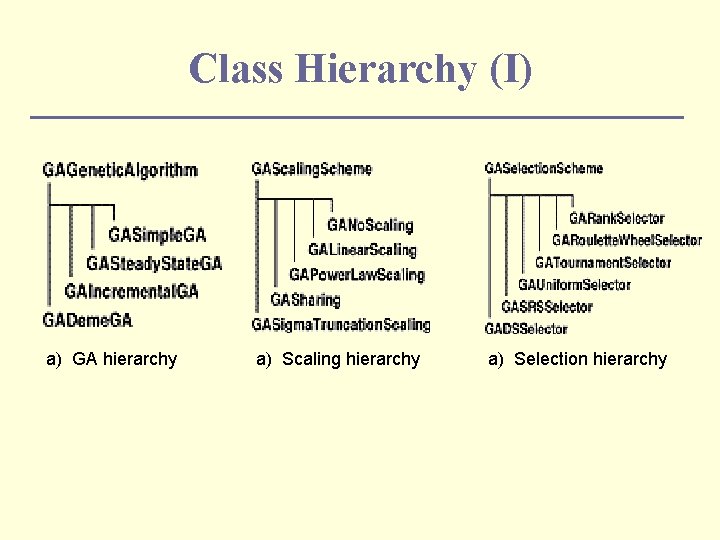
Class Hierarchy (I) a) GA hierarchy a) Scaling hierarchy a) Selection hierarchy
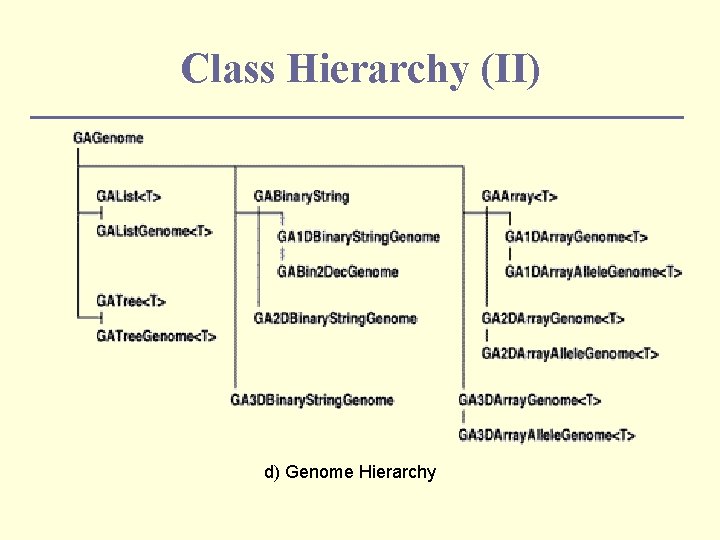
Class Hierarchy (II) d) Genome Hierarchy

The form of a typical optimization problem #include <ga/GASimple. GA. h> #include <ga/GA 1 DBin. Str. Genome. h> float Objective(GAGenome &); main(int argc, char **argv){ int length = 10; GA 1 DBinary. String. Genome genome(length, Objective); //create a genome GASimple. GA ga(genome); //create the genetic algorithm ga. evolve(); //do the evolution cout << ga. statistics() << endl; //print out the results } float Objective(GAGenome & g) { GA 1 DBinary. String. Genome & genome = (GA 1 DBinary. String. Genome &)g; //put here your objective function }
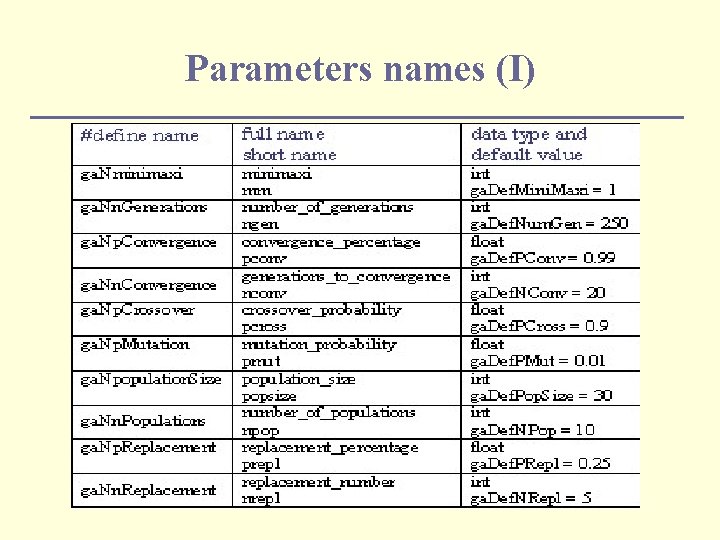
Parameters names (I)
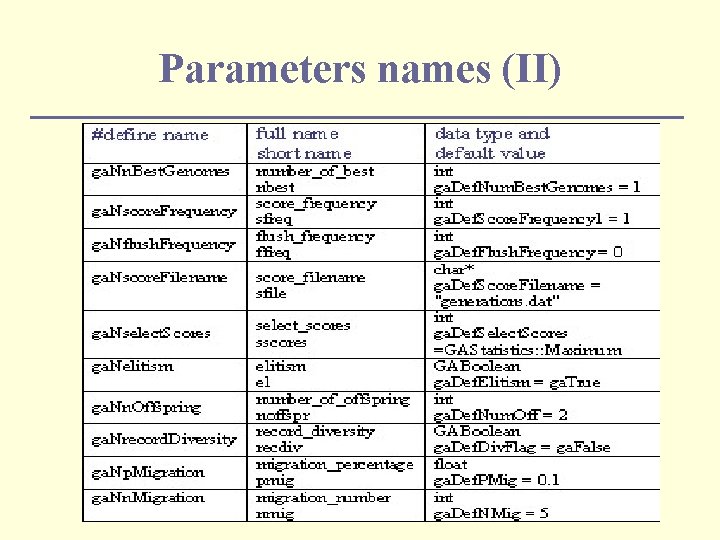
Parameters names (II)

Change the behaviour of the GA (I) n By setting various parameters: ga. population. Size(popsize); ga. n. Generations(ngen); ga. p. Mutation(pmut); ga. p. Crossover(pcross); n Parameters in comand line: GAParameter. List params; GASimple. GA: : register. Default. Parameters(params); params. set(ga. Nn. Generations, 2000); params. parse(argc, argv, ga. False); GA 1 DBinary. String. Genome genome(length, Objective); GASimple. GA ga(genome); ga. parameters(params);

Change the behaviour of the GA (II) n GASigma. Truncation. Scaling scale; n n n Ga. scaling(scale); GARank. Selector scheme; ga. selector(scheme); ga. terminator(GAGenetic. Algorithm: : Terminate. Upon. Co nvergence); genome. crossover(GA 2 DBinary. String. Genome: : One. Poi nt. Crossover); genome. mutator(GA 2 DBinary. String. Genome: : Flip. Mutat or); ga. replacement(GAIncremental. GA: : RANDOM);

Define our own operators int My. Mutator(GAGenome&, float); void My. Initializer(GAGenome&); float My. Comparator(const GAGenome&, const GAGenome&); int My. Crossover(const GAGenome&, GAGenome*, GAGenome*); GA 1 DBinary. String. Genome genome(20); genome. initializer(My. Initializer); genome. mutator(My. Mutator); genome. comparator(My. Comparator); genome. crossover(My. Crossover);

Traveling Salesman Problem (TSP) n Given a finite set of cities C={c 1, c 2, …, cn} and a distance d(ci, cj), i≠j, between each pair, the TSP asks for finding a tour through all of the cities, visiting each exactly once, and returning to the originating city such that the total distance traveled is minimized.

Format of the file that contains cities City X Y 1 1 1 2 4 2 3 4 1 4 5 12 6 … … … N -1 29 3 N 17 20

Genome and Initializer n To resolve this problem: n Use GAList. Genome. The cities are listed in the order in which they are visited. 10 n 3 5 9 2 7 4 1 6 8 Define your own Initializer n n Use GARandom. Int(int, int); Use a vector in which are stored the visited cities.

Crossover n Define your own crossover: n This crossover selects the first city of one parent, compares the cities leaving that city in both parents, and chooses the closer one to extend the tour. If one city has already appeared in the tour, we choose the other city. If both cities have already appeared, we randomly select a nonselected city. "

Mutator n Define your own mutator n This mutator randomly select two cities from one chromosome and swap them if the new (swapped) tour length is shorter than the old one. n Example: Before mutation 10 3 5 9 2 7 4 1 6 8 9 1 6 8 ≥ After mutation 10 3 5 4 2 7
- Slides: 23