Functional Genomics Functional genomic datasets Biological networks Integrating

Functional Genomics • Functional genomic datasets • Biological networks • Integrating genomic datasets BIO 520 Bioinformatics Jim Lund

Functional genomics • Genome scale experiments to understand the function of all the proteins--what they do and how they interact. • Many different experimental designs – Different kinds of information generated. • Each has experimental limitations – Coverage: full genome, limited? – False positives. – False negatives.
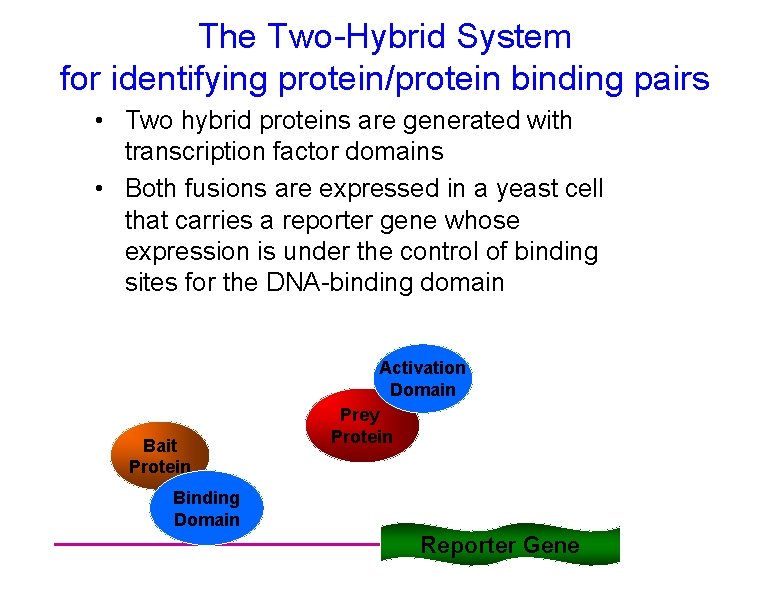
The Two-Hybrid System for identifying protein/protein binding pairs • Two hybrid proteins are generated with transcription factor domains • Both fusions are expressed in a yeast cell that carries a reporter gene whose expression is under the control of binding sites for the DNA-binding domain Bait Protein Activation Domain Prey Protein Binding Domain Reporter Gene
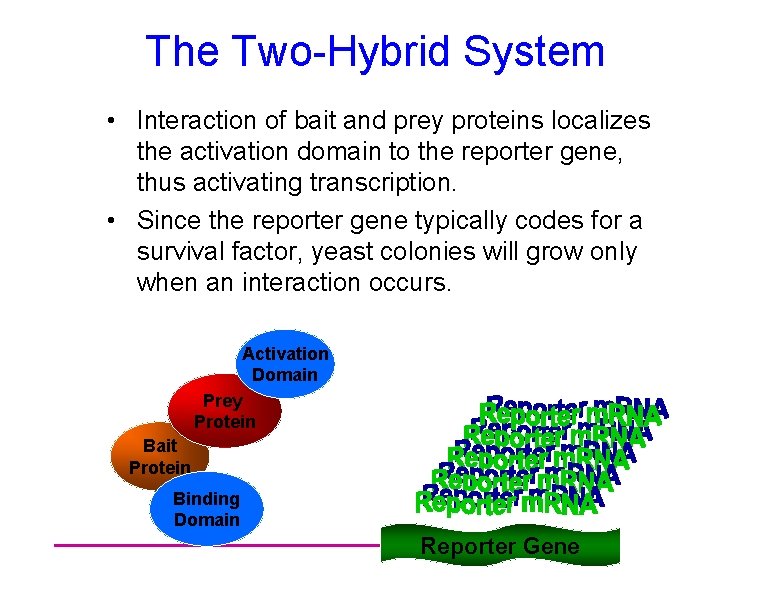
The Two-Hybrid System • Interaction of bait and prey proteins localizes the activation domain to the reporter gene, thus activating transcription. • Since the reporter gene typically codes for a survival factor, yeast colonies will grow only when an interaction occurs. Activation Domain Prey Protein Bait Protein Binding Domain Reporter Gene
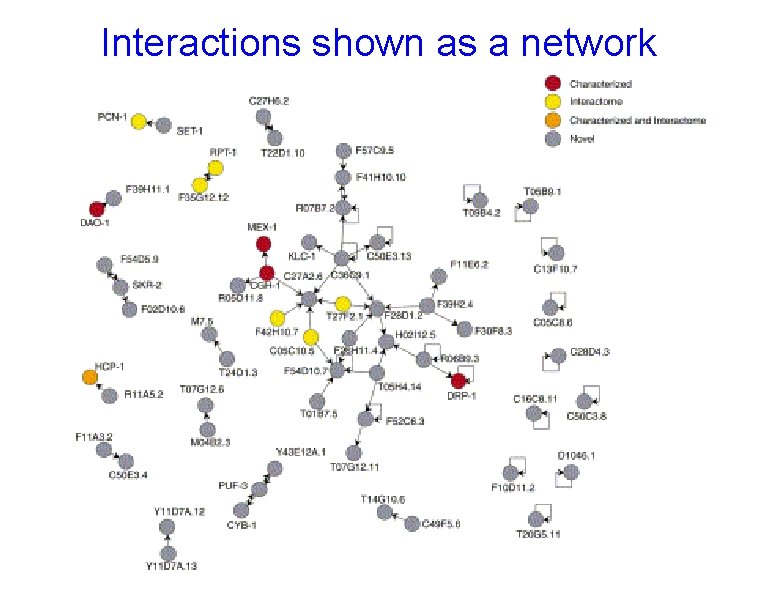
Interactions shown as a network

Networks • When methods of detecting functional linkages are applied to all the proteins of an organism, network of interacting, functionally linked proteins can be traced. • As methods improve for detecting protein linkages, it seems likely that most of the proteins will be included in the network.

What do you miss? • Tertiary interactions • Regulated interactions – Subcellular localization dependent – Cofactor dependent (eg. Hormoneregulated) • Low-affinity (Kd>10 -6)
Cellular Location • Immunolocalization – FUSION PROTEINS YFG GFP • Prediction – Membrane vs non-membrane • improved by homology • WHICH MEMBRANE – Nuclear vs cytoplasmic
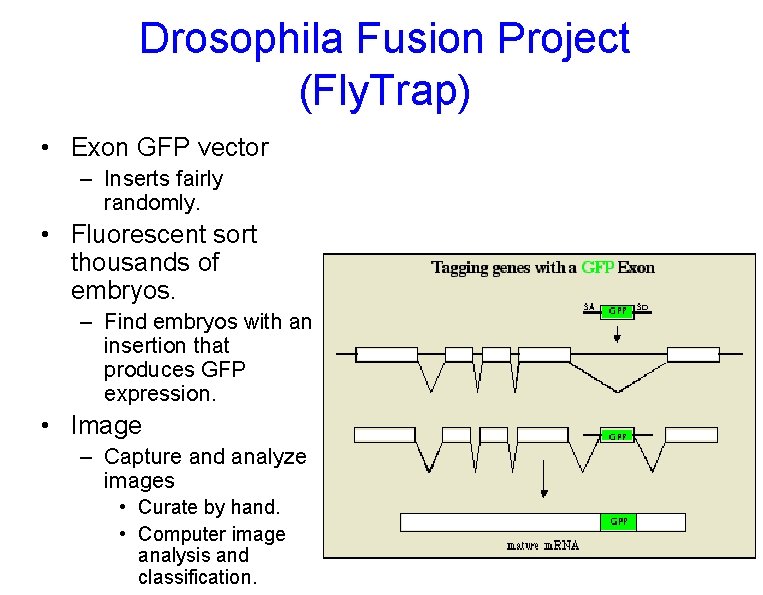
Drosophila Fusion Project (Fly. Trap) • Exon GFP vector – Inserts fairly randomly. • Fluorescent sort thousands of embryos. – Find embryos with an insertion that produces GFP expression. • Image – Capture and analyze images • Curate by hand. • Computer image analysis and classification.
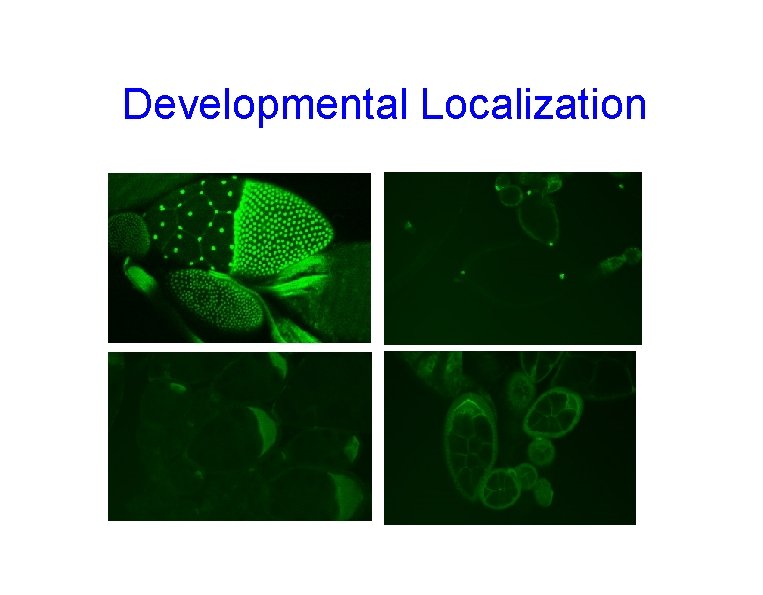
Developmental Localization
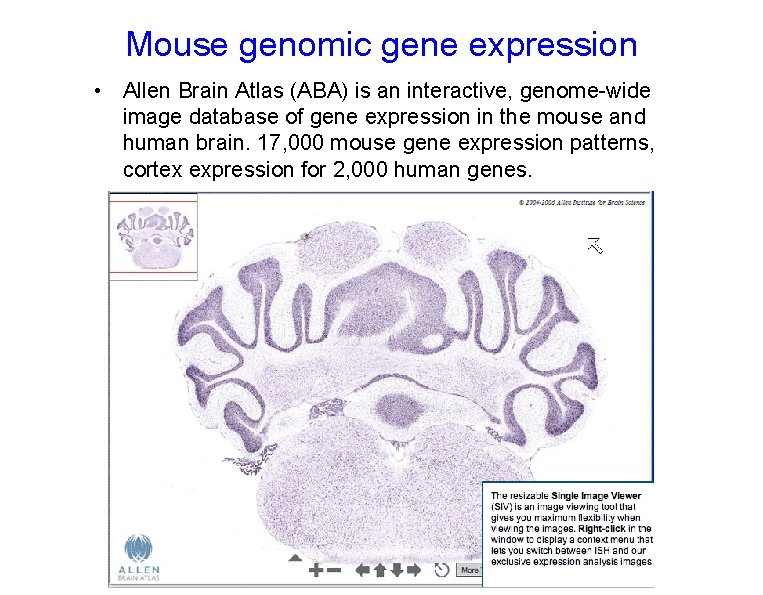
Mouse genomic gene expression • Allen Brain Atlas (ABA) is an interactive, genome-wide image database of gene expression in the mouse and human brain. 17, 000 mouse gene expression patterns, cortex expression for 2, 000 human genes.
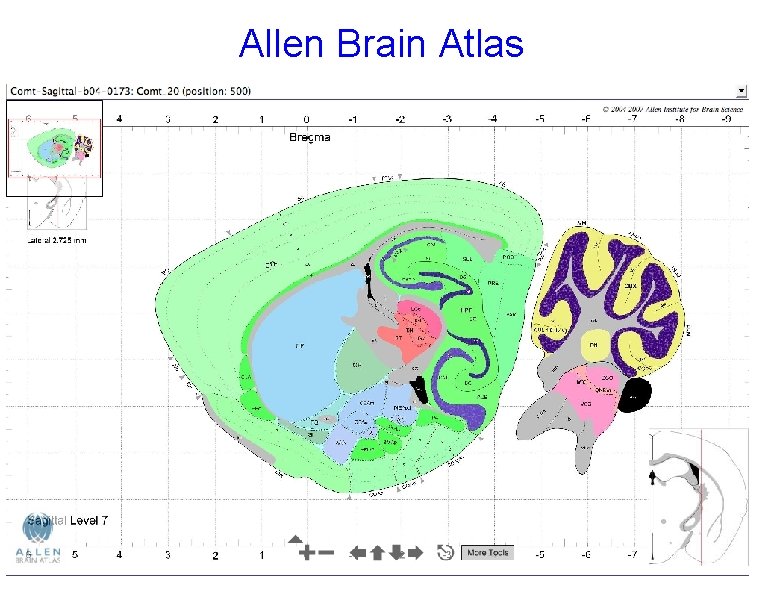
Allen Brain Atlas

3 D mouse gene expression project Single gene expression database for the mouse research community. Integrated in the Mouse Genome Database (MGD) at the Jackson Laboratory. 10, 302 expression entries WT 1 expression (red) on a section of the E 9 (Theiler Stage 14) embryo from the Edinburgh Mouse Atlas. The gut epithelium is shown in yellow and the neural tube in a blue overlay. WT 1 is expressed in the presumptive mesothelium of the coelom and in the intermediate mesoderm (ventral to the somites).

Methods for discovering protein function • Automated Binding Assays • High Throughput Enzyme Assays
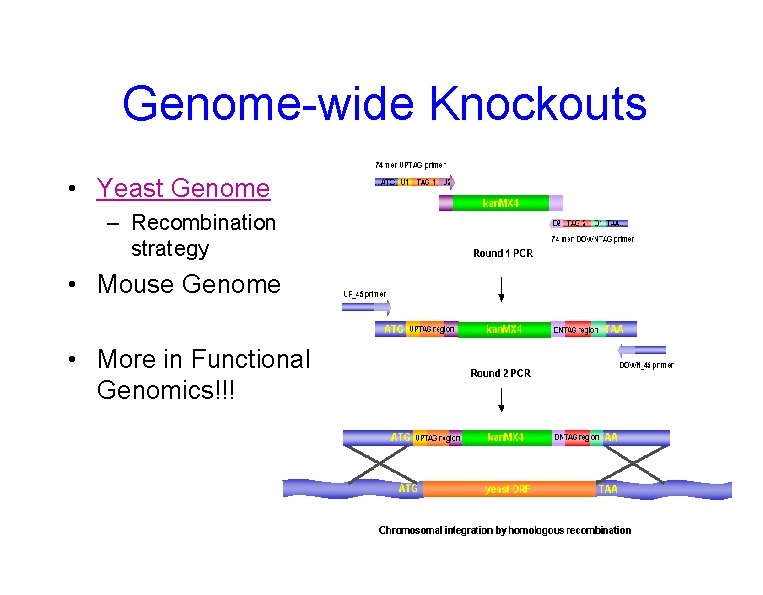
Genome-wide Knockouts • Yeast Genome – Recombination strategy • Mouse Genome • More in Functional Genomics!!!

Essential vs Non-essential • Transcription similar – >99% essential genes transcribed • Transcript level 70% higher – >90% non-essential transcribed • Genome locations similar – Not clustered – Essential genes rarely near telomeres

Why only 20% essential? • Redundant – 8. 5% of non-essential had CLOSE homolog in genome (P<10 -150) • Essential in another condition • Marginal Benefit

Resources YEAST MOUSE • Saccharomyces Genome Deletion Project • Mouse Phenome Database – http: //wwwsequence. stanford. edu/ group/yeast_deletion_p roject/deletions 3. html – http: //phenome. jax. org/pubcgi/phenome/mpdcgi? rtn=docs/h ome • Knockout Mouse Project – http: //www. knockoutmouse. org/
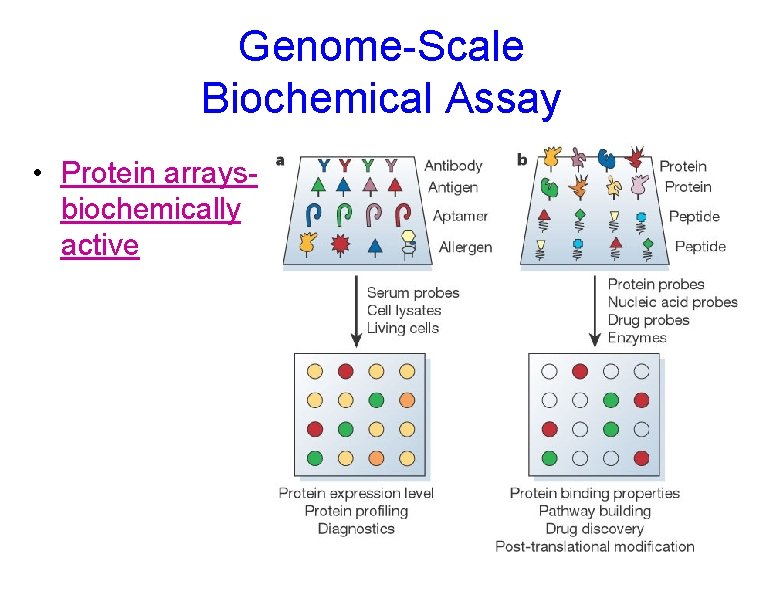
Genome-Scale Biochemical Assay • Protein arraysbiochemically active

Databases • Relationships between genes/proteins. • How are different types of experimental data integrated? – Schema • Data quality – Who curates? – Who revises?

Proteome Projects • Swiss. Prot (Ex. Pasy) – http: //expasy. org/ch 2 d/ • Saccharomyces Genome Database (SGD) Gene Function Information – 2 -hybrid, functional assignments, pathways. – http: //www. yeastgenome. org/Search. Contents. shtml • Yale TRIPLES – Database of TRansposon-Insertion Phenotypes, Localization, and Expression in Saccharomyces. • 2 -hybrid databases – http: //proteome. wayne. edu/YTHwebsites. html

Pathway and interaction databases • KEGG (http: //www. genome. jp/kegg/) – Metabolic and signaling pathways • PUMA (http: //compbio. mcs. anl. gov/puma 2/cgi-bin/index. cgi) – Metabolic and signaling pathways • DIP (http: //dip. doe-mbi. ucla. edu/) – Protein-protein interactions • BIND (http: //bind. ca/) – Molecular and genetic interactions
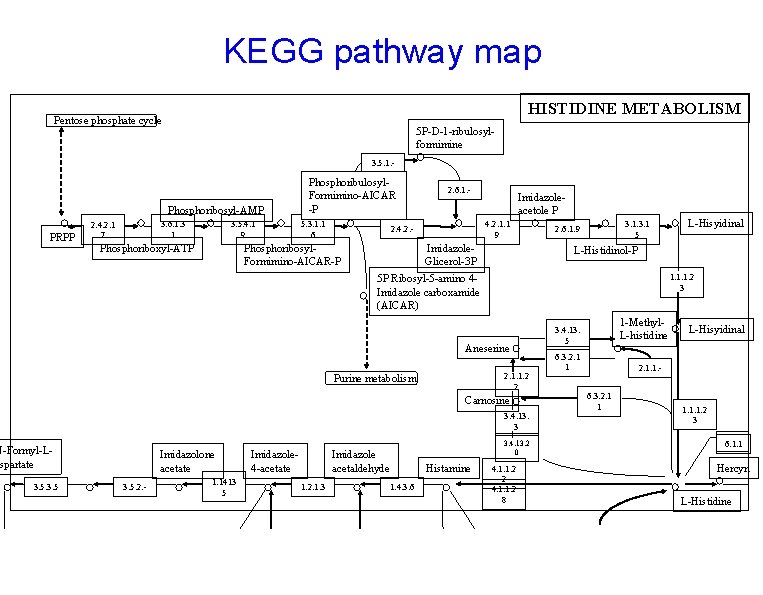
KEGG pathway map HISTIDINE METABOLISM Pentose phosphate cycle 5 P-D-1 -ribulosylformimine 3. 5. 1. - Phosphoribosyl-AMP PRPP 3. 6. 1. 3 1 2. 4. 2. 1 7 3. 5. 4. 1 9 Phosphoribulosyl. Formimino-AICAR -P 5. 3. 1. 1 6 Imidazoleacetole P 4. 2. 1. 1 9 2. 4. 2. - Phosphoribosyl. Formimino-AICAR-P Phosphoriboxyl-ATP 2. 6. 1. - Imidazole. Glicerol-3 P 3. 1 5 2. 6. 1. 9 L-Histidinol-P 1. 1. 1. 2 3 5 P Ribosyl-5 -amino 4 Imidazole carboxamide (AICAR) Aneserine 2. 1. 1. 2 2 Purine metabolism Carnosine 3. 4. 13. 3 N-Formyl-Lspartate 3. 5 Imidazolone acetate 3. 5. 2. - 1. 1413 5 Imidazole 4 -acetate 3. 4. 13. 2 0 Imidazole acetaldehyde 1. 2. 1. 3 Histamine 1. 4. 3. 6 L-Hisyidinal 4. 1. 1. 2 2 4. 1. 1. 2 8 1 -Methyl. L-histidine 3. 4. 13. 5 6. 3. 2. 1 1 L-Hisyidinal 2. 1. 1. 6. 3. 2. 1 1 1. 1. 1. 2 3 6. 1. 1 Hercyn L-Histidine
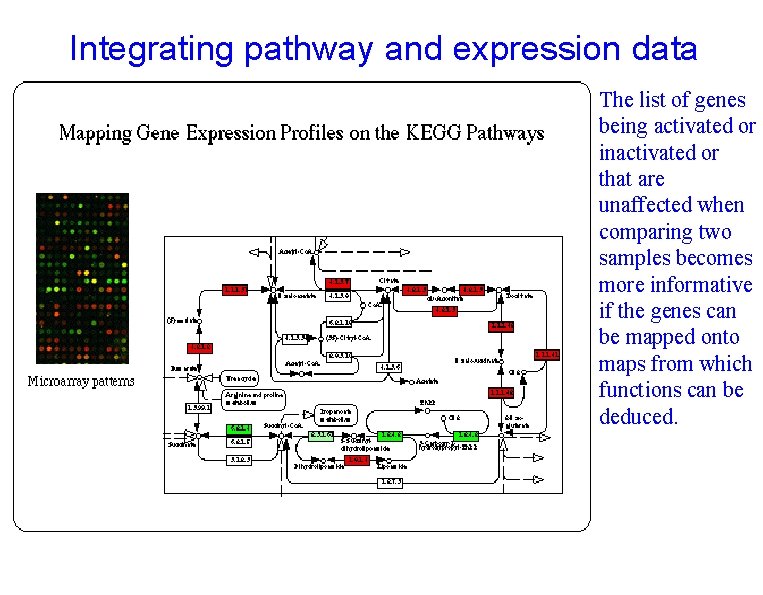
Integrating pathway and expression data The list of genes being activated or inactivated or that are unaffected when comparing two samples becomes more informative if the genes can be mapped onto maps from which functions can be deduced.
- Slides: 24