function from sequence Peer Bork EMBL MDC Heidelberg

function from sequence Peer Bork EMBL & MDC Heidelberg & Berlin bork@embl-heidelberg. de http: //www. bork. embl-heidelberg. de/

Bioinformatics Generation of information (biophysics) n Storage and retrieval of information (informatics for biodatabases) n Translation of information into knowledge (computational biology) n www. bork. embl-heidelberg. de
www. bork. embl-heidelberg. de
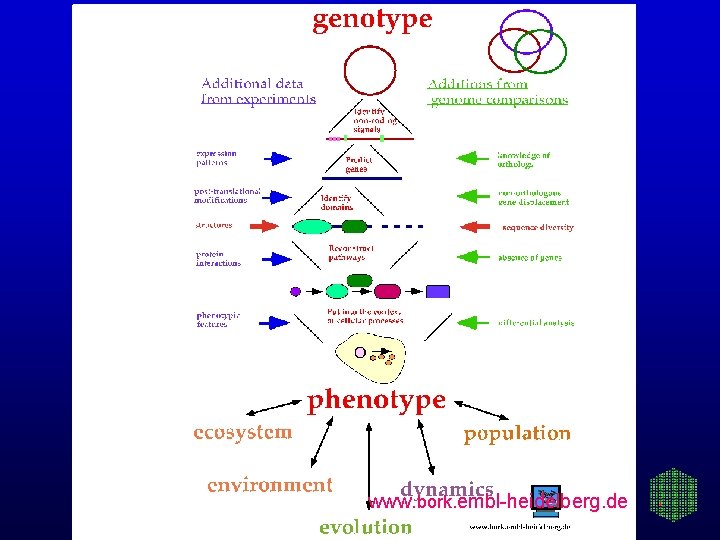
www. bork. embl-heidelberg. de

Chance of deducing structural and functional featur featu by homology Many homologues, an increasing number o predictable folds, bu tough times for automatic function prediction www. bork. embl-heidelberg. de

Function prediction from sequence Quality and heterogeneity of da Prediction accuracy: 70% hurd Function and domain prediction Function prediction by gene co www. bork. embl-heidelberg. de
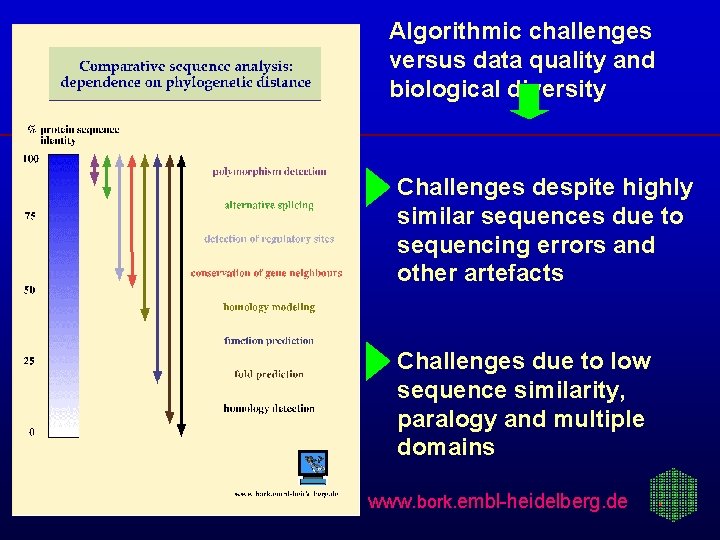
Algorithmic challenges versus data quality and biological diversity Challenges despite highly similar sequences due to sequencing errors and other artefacts Challenges due to low sequence similarity, paralogy and multiple domains www. bork. embl-heidelberg. de

No human genes in thousands Number of human genes in time 120 100 80 60 40 20 HGS, Incyte and co Textbooks, public opinion 52 Celera HGP 38 39 32 HGS others 27 24 21 0 Mar 00 Aug 00 Oct 00 Dec 00 Feb 01 Apr 01 www. bork. embl-heidelberg. de

Nature 304, 16. November 2000

Heterogenous data from large scale approaches Gene expression (correlation to proteins poor) Yeast two hybrid (8% overlap with each other) Many others…. www. bork. embl-heidelberg. de
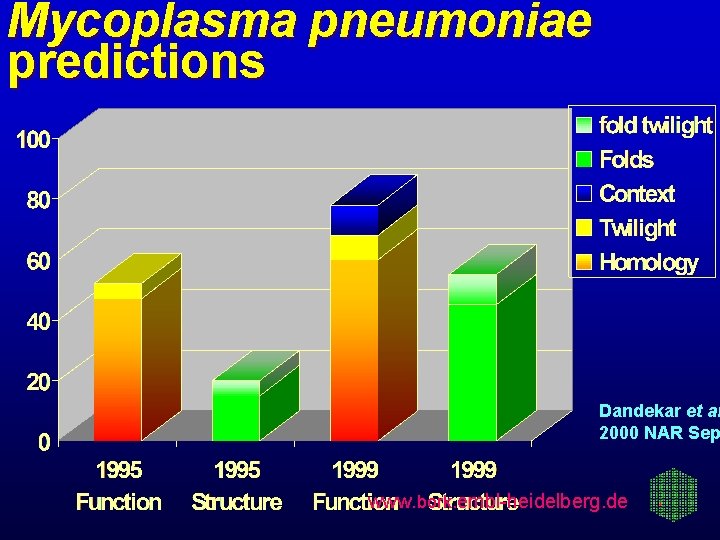
Mycoplasma pneumoniae predictions Dandekar et al 2000 NAR Sep www. bork. embl-heidelberg. de

Mycoplasma pneumoniae reannotation 1995 vs 1999 RNAs: +9 = 42 ORFs: +12 -1 = 688 with functions: +105 = 458 ORFs changed: 16 extended, 8 shorte Function changed: 30 more, 18 less s 57% of all entrieswww. were re-annotated bork. embl-heidelberg. de

Function prediction from sequence Quality and heterogeneity of da Prediction accuracy: 70% hurd Function and domain prediction Function prediction by gene co www. bork. embl-heidelberg. de
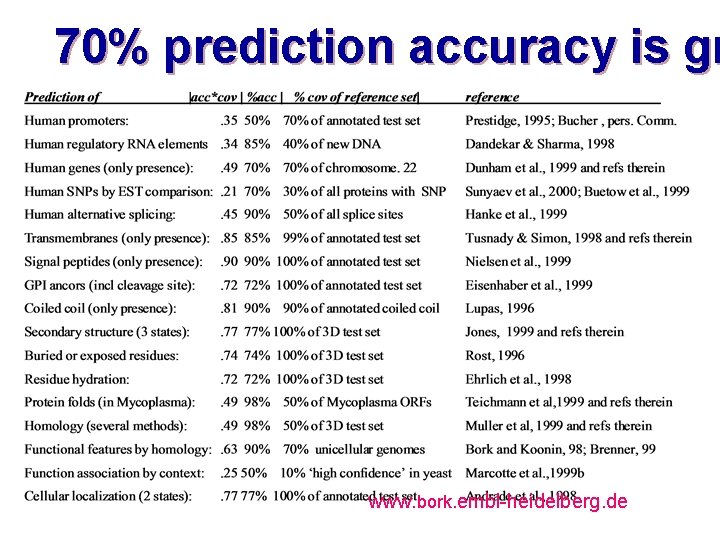
70% prediction accuracy is gr www. bork. embl-heidelberg. de

Clear homology via Blast; yet, misleading otation hampers automatic function predic www. bork. embl-heidelberg. de

Phylogenetic tree of Blast hits reveals that no function prediction is possible www. bork. embl-heidelberg. de

Molecular Functions have to be defined on a domain basis i. e. separately for each structurally independent unit within Henikoff etaal. 1997 www. bork. embl-heidelberg. de Science 278, 609 sequence

Function prediction from sequence Quality and heterogeneity of da Prediction accuracy: 70% hurd Function and domain prediction Function prediction by gene co www. bork. embl-heidelberg. de
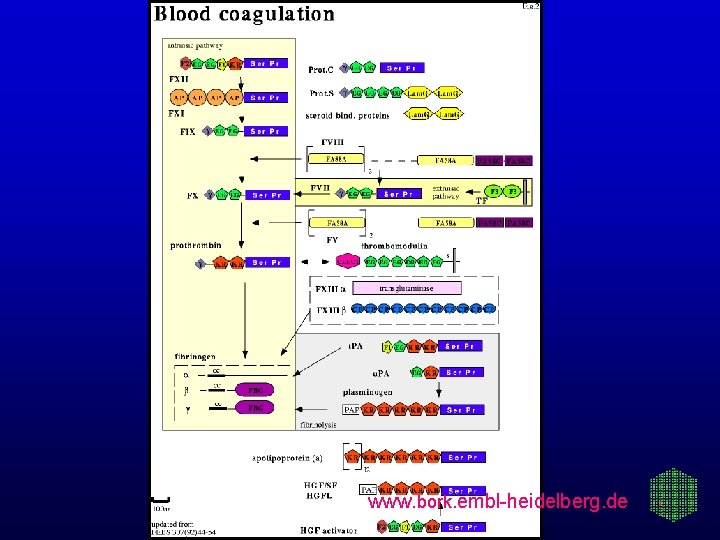
www. bork. embl-heidelberg. de
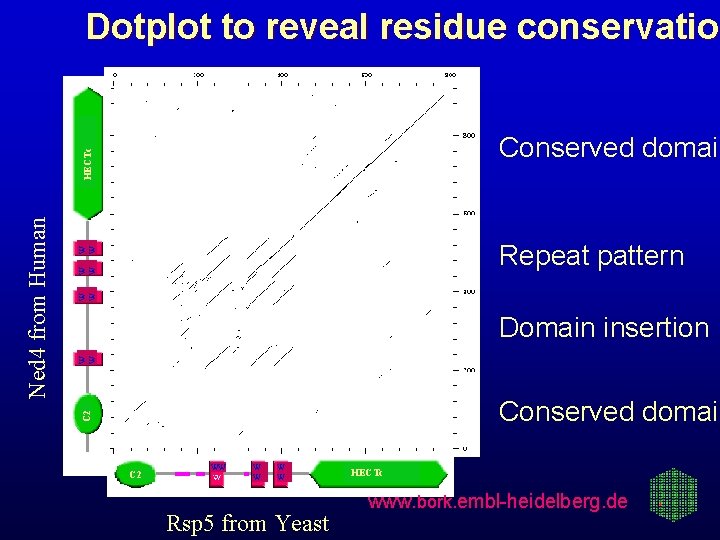
Dotplot to reveal residue conservation conservatio W W W Repeat pattern W W Domain insertion Conserved domain C 2 Ned 4 from Human HECTc Conserved domain C 2 WW W W Rsp 5 from Yeast HECTc www. bork. embl-heidelberg. de

Function prediction for disease genes Breast cancer gene BRCA 1 Positionally cloned 1994 (Miki et al. Science 266, Features originally deduced from the 1857 aa sequen Contains a RING finger (30 aa, usually bind diverse p Function unknown, even localization unc www. bork. embl-heidelberg. de

Localization experiments on BRCA 1 Title/Journal Conclusion www. bork. embl-heidelberg. de
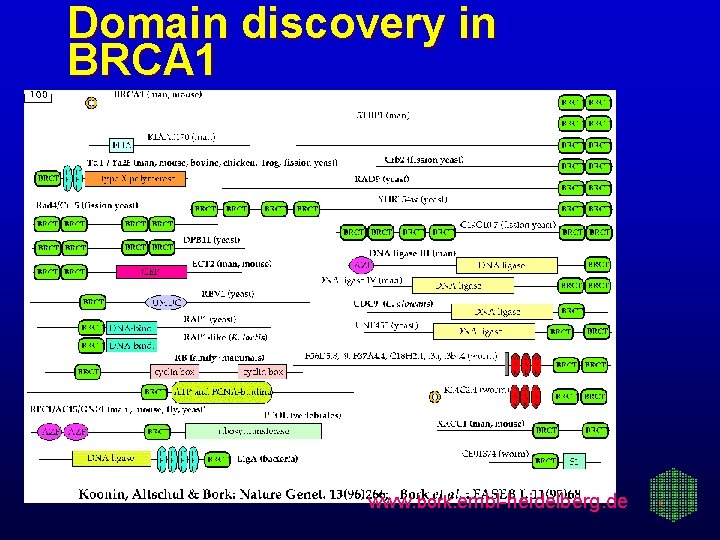
Domain discovery in BRCA 1 www. bork. embl-heidelberg. de
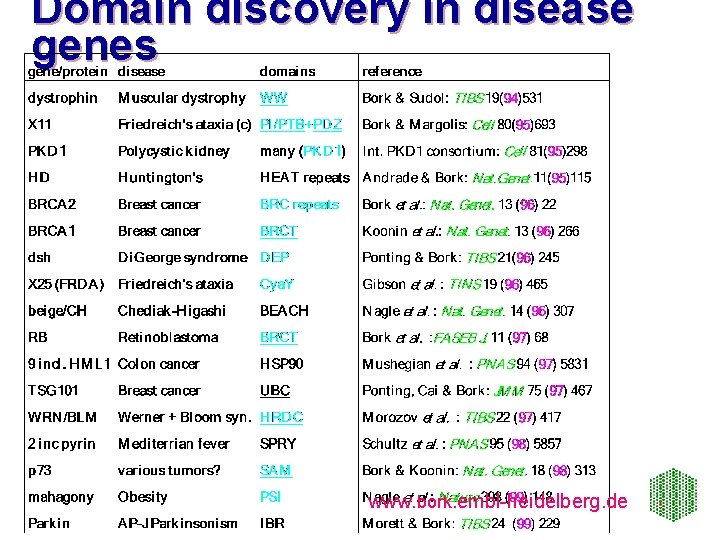
Domain discovery in disease genes www. bork. embl-heidelberg. de
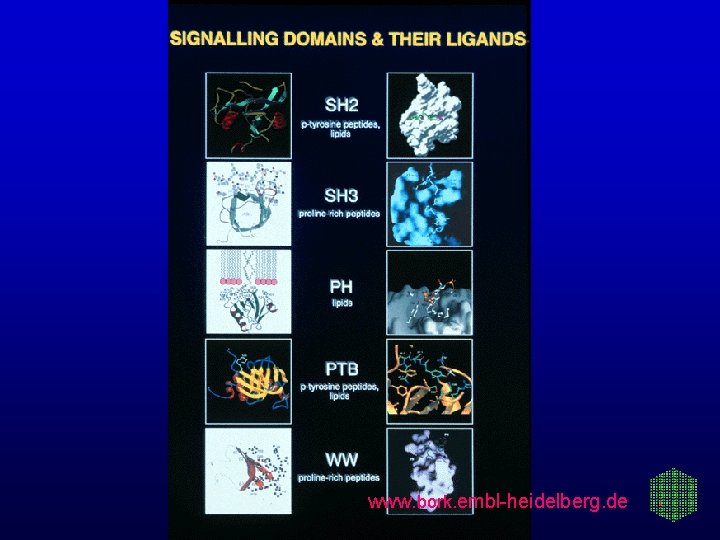
www. bork. embl-heidelberg. de
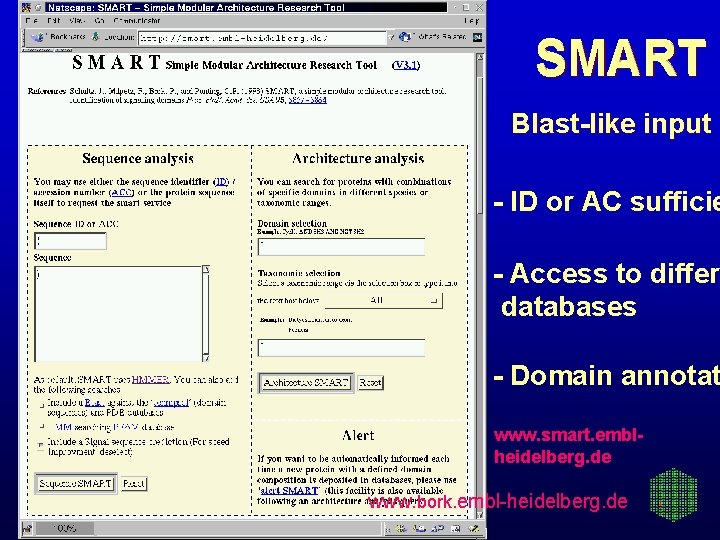
SMART Blast-like input - ID or AC sufficie - Access to differ databases - Domain annotat www. smart. emblheidelberg. de www. bork. embl-heidelberg. de

SMART Digested output -signal sequence -transmembrane regions -comparison of domain context www. smart. emblheidelberg. de www. bork. embl-heidelberg. de

Non-globuar functional features in protein sequences n n n Transmembrane regions signal sequences GPI anchors coiled-coiled other compositionally biased regions (short internal repeats) www. bork. embl-heidelberg. de

SMART Blast with “in between” regions -automatically cuts respective region -cut and paste for other programs -some specific outpu features www. smart. emblwww. bork. embl-heidelberg. de

SMART Digested output -signal sequence -transmembrane regions -comparison of domain context www. smart. emblheidelberg. de www. bork. embl-heidelberg. de

SMART Domain annotation -description -multiple alignmen -consensus featur -residue annotatio -search options www. smart. emblheidelberg. de www. bork. embl-heidelberg. de

SMART Species distributio -total occurrence -protein and domain statistics -taxonomic break do -model organisms www. smart. emblheidelberg. de www. bork. embl-heidelberg. de

Annotation improvement using domain correlation n Query: VAV H. sapiens Find closest hit: selective SMART n Domain architecture of C 35 B 8. 2 C. elegans Evaluate correlation; scan genome region n Reconstructed structure of C 35 B 8. 2 SH 3 www. bork. embl-heidelberg. de
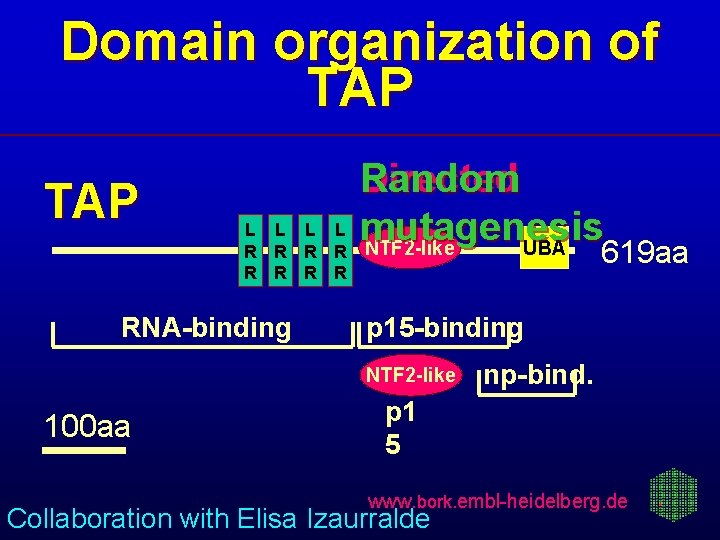
Domain organization of TAP L L R R R R RNA-binding Random Directed mutagenesis NTF 2 -like UBA 619 aa p 15 -binding NTF 2 -like 100 aa np-bind. p 1 5 www. bork. embl-heidelberg. de Collaboration with Elisa Izaurralde
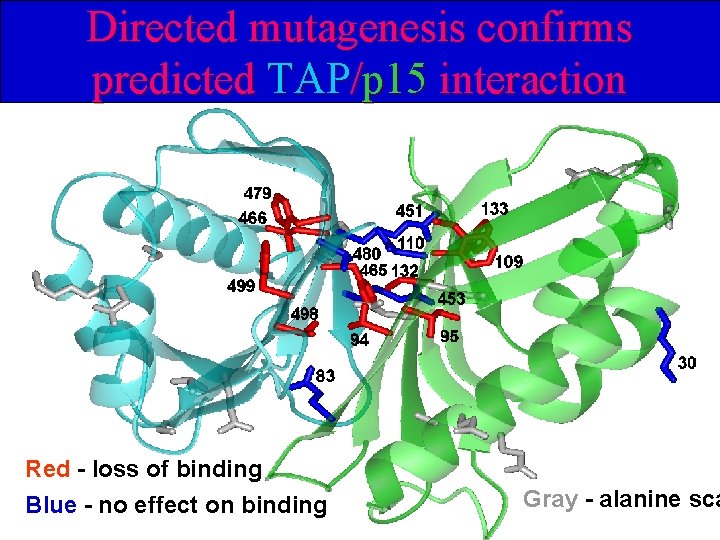
Directed mutagenesis confirms predicted TAP/p 15 interaction Red - loss of binding Blue - no effect on binding Gray - alanine sca
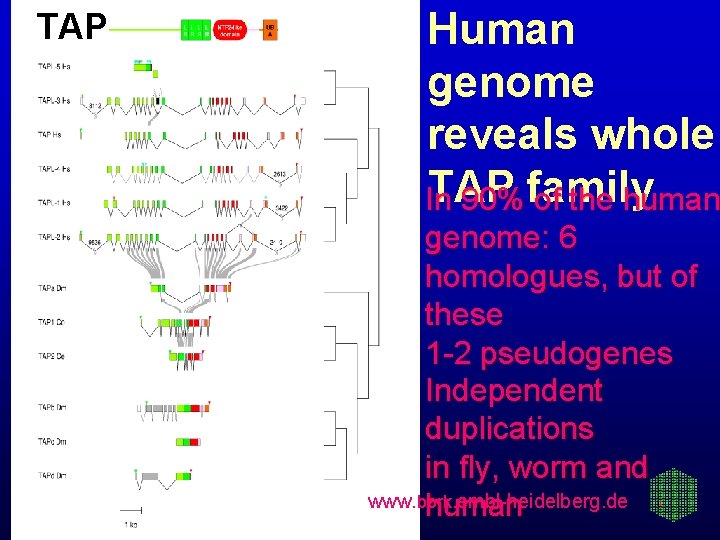
TAP Human genome reveals whole TAP In 90% family of the human genome: 6 homologues, but of these 1 -2 pseudogenes Independent duplications in fly, worm and www. bork. embl-heidelberg. de human
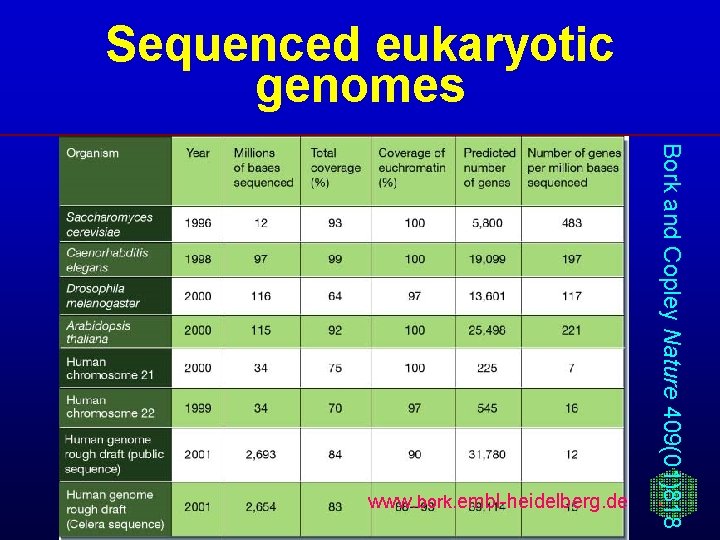
Sequenced eukaryotic genomes Bork and Copley Nature 409(01)818 www. bork. embl-heidelberg. de
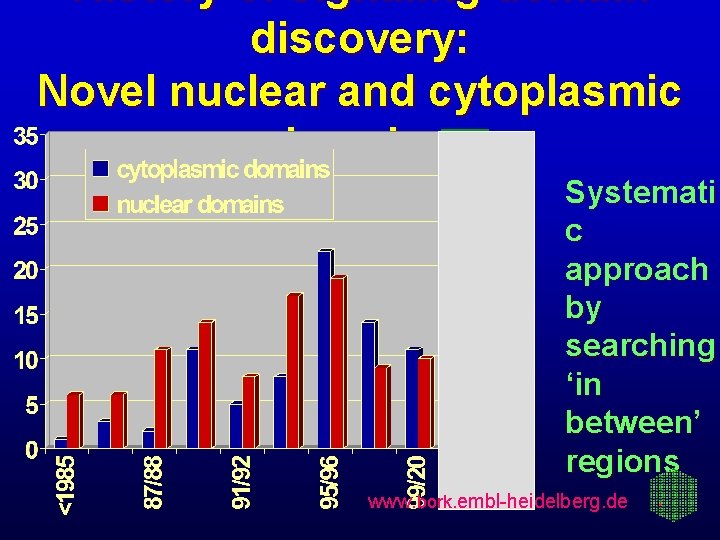
History of signaling domain discovery: Novel nuclear and cytoplasmic domains Systemati c approach by searching ‘in between’ regions www. bork. embl-heidelberg. de

Top 10 domains* in human Species man fly wormyeast cress Total no genes 13300 18200 6100 25700 26500(26500) 1 64 0 Immunoglobulin 765(381) 140 115 48 C 2 H 2 zinc finger 706(607) 357 151 Protein kinase 575(501) 319 437 121 1049 16 0 Rhod. -like GPCR 569(616) 97 358 331 198 183 97 P-loop NTPase 433 80 10 50 6 Rev. transcriptase 350 255 96 54 300(224) 157 RRM (RNA-binding) 210 91 WD 40 (G-protein)277(136) 162 102 19 120 Ankyrin repeat 276(145) 105 107 109 9 118 267(160) 148 Homeobox www. bork. embl-heidelberg. de *Only no of genes given, no of domains higher; note that only around Nature 409 (01)860; Science 291(01)1304 90% is sequenced
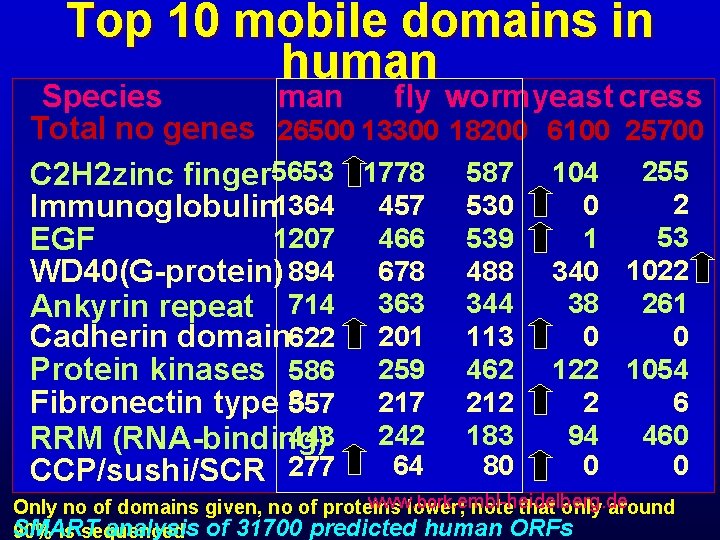
Top 10 mobile domains in human Species man fly wormyeast cress Total no genes 26500 13300 18200 6100 25700 C 2 H 2 zinc finger 5653 1778 587 104 255 2 0 Immunoglobulin 1364 457 530 53 466 539 1 1207 EGF WD 40(G-protein) 894 678 488 340 1022 261 38 Ankyrin repeat 714 363 344 0 0 Cadherin domain 622 201 113 Protein kinases 586 259 462 122 1054 6 217 212 2 Fibronectin type 3 557 242 183 94 460 443 RRM (RNA-binding) 64 80 0 0 CCP/sushi/SCR 277 www. lower; bork. embl-heidelberg. de Only no of domains given, no of proteins note that only around SMART analysis of 31700 predicted human ORFs 90% is sequenced

Correlation between domains other Marker PX extra nuclear intra www. bork. embl-heidelberg. de

Function prediction from sequence Quality and heterogeneity of da Prediction accuracy: 70% hurd Function and domain prediction Function prediction by gene co www. bork. embl-heidelberg. de

Phenotypic features do not coincide with species evolution. . . yeast . . . but gene content does www. bork. embl-heidelberg. de
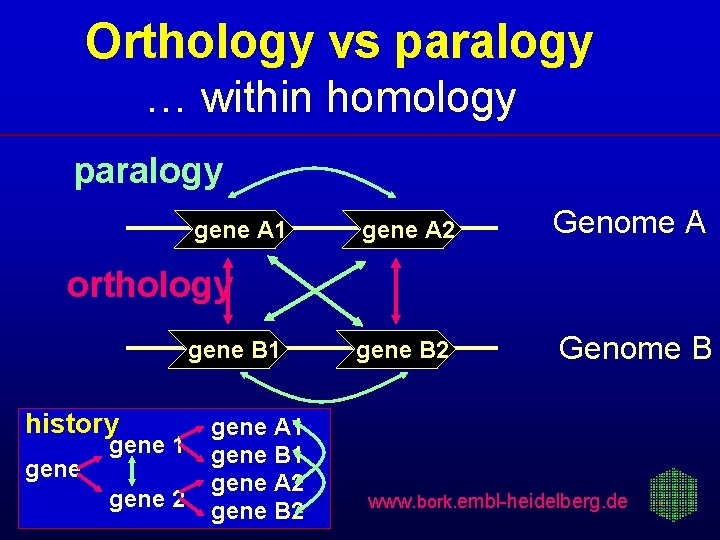
Orthology vs paralogy … within homology paralogy gene A 1 gene A 2 Genome A gene B 2 Genome B orthology gene B 1 history gene 1 gene 2 gene A 1 gene B 1 gene A 2 gene B 2 www. bork. embl-heidelberg. de
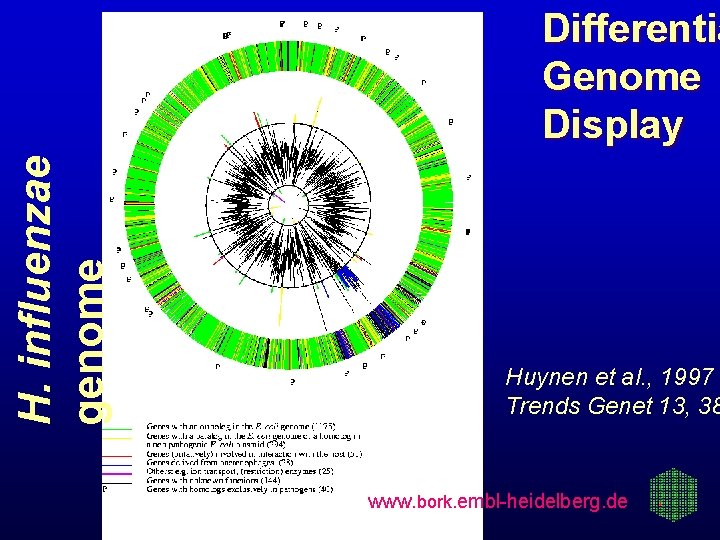
H. influenzae genome Differentia Genome Display Huynen et al. , 1997 Trends Genet 13, 38 www. bork. embl-heidelberg. de

Exploiting the absence of genes Huynen et al. , 1998, FEBS Lett 426, 1 -5 www. bork. embl-heidelberg. de

Predicting functional interactions between proteins by the co-occurrence of their genes in genomes Distribution of four M. genitalium genes among 25 genomes MG 299 (pta) 0 0 0 1 1 0 1 0 1 1 0 0 0 1 1 1 1 MG 357(ack. A) 0 0 0 1 1 0 1 0 1 1 0 0 0 1 1 1 1 MG 019(dna. J) 0 0 1 1 1 0 1 1 1 0 0 1 1 1 MG 305(dna. K)0 0 1 1 1 0 1 1 1 0 0 1 1 1 Using the mutual information between genes as a scoring heuristic for their co-occurrence. M(pta, ack. A)=0. 69 (phospotransacetylase, acet M(dna. J, dna. K)=0. 55 (heat shock proteins) M(dna. J, ack. A)=0. 19 www. bork. embl-heidelberg. de
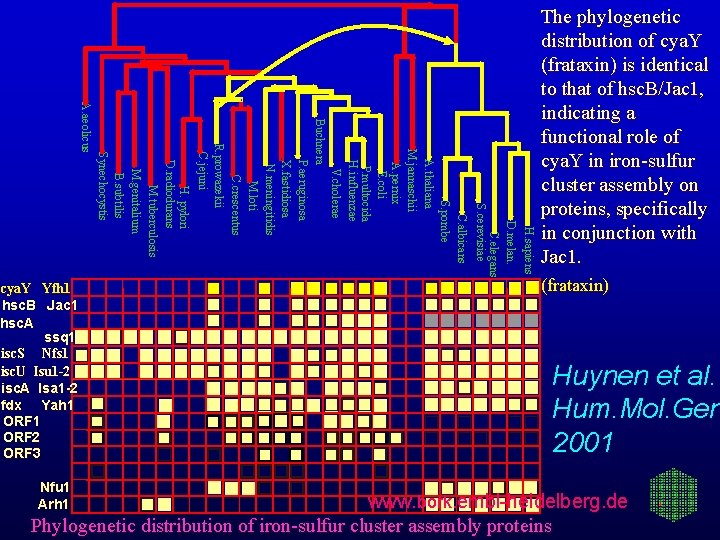
H. sapiens D. melan. C. elegans S. cerevisiae Nfu 1 Arh 1 C. albicans S. pombe A. thaliana M. jannaschii A. pernix E. coli P. multocida H. influenzae V. cholerae Buchnera P. aeruginosa X. fastidiosa N. meningitidis M. loti C. crescentus R. prowazekii C. jejuni H. pylori D. radiodurans M. tuberculosis M. genitalium B. subtilis Synechocystis A. aeolicus cya. Y Yfh 1 hsc. B Jac 1 hsc. A ssq 1 isc. S Nfs 1 isc. U Isu 1 -2 isc. A Isa 1 -2 fdx Yah 1 ORF 2 ORF 3 The phylogenetic distribution of cya. Y (frataxin) is identical to that of hsc. B/Jac 1, indicating a functional role of cya. Y in iron-sulfur cluster assembly on proteins, specifically in conjunction with Jac 1. (frataxin) Huynen et al. Hum. Mol. Gen 2001 www. bork. embl-heidelberg. de Phylogenetic distribution of iron-sulfur cluster assembly proteins

Function prediction via gene context information Genomic context - Conserved gene neighborhood in genomes information: - Gene fusion as distinct neighborhood subset - Conserved co-occurrence of genes in species (‘phylogentic profile’, ‘COG pattern’) - Surrounding and shared regulatory elements Knowledge-based context information - Pathway data (can overrule homology!) - Gene expression data (co-expression etc. ) - Protein interaction /localisation www. bork. embl-heidelberg. de - Scientific literature
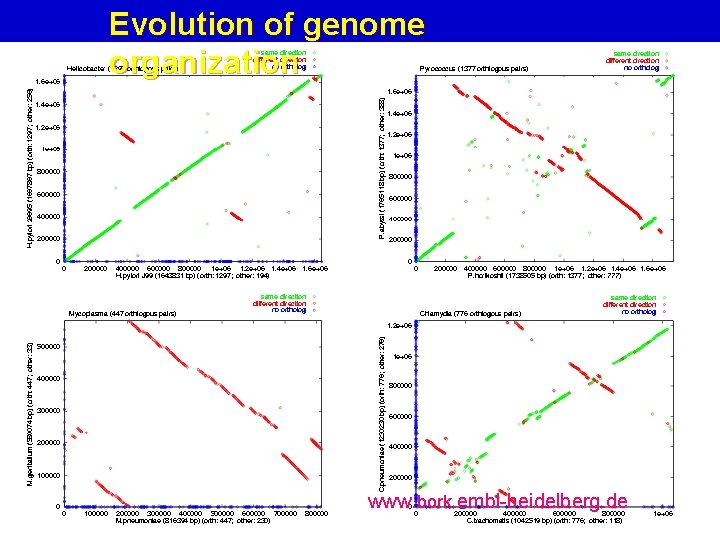
Evolution of genome organization www. bork. embl-heidelberg. de

Dotplot to reveal gene order conservatio www. bork. embl-heidelberg. de
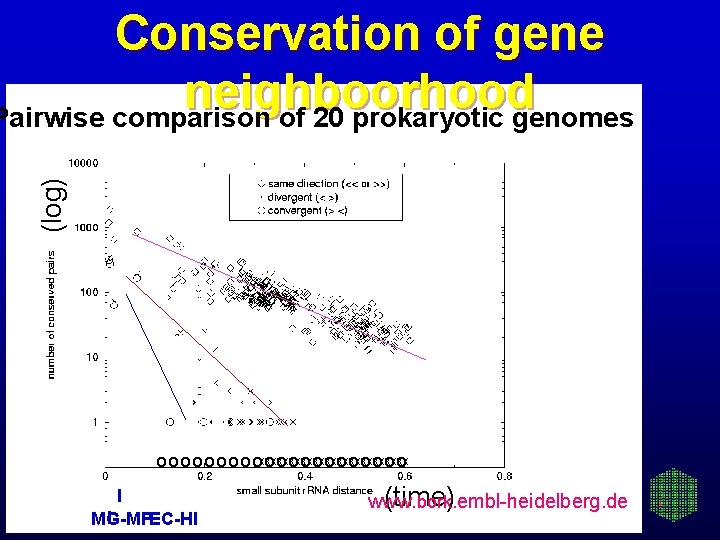
(log) Conservation of gene neighboorhood Pairwise comparison of 20 prokaryotic genomes o o o o oxxxxxxxxxxxxx oooooo I I MG-MP EC-HI (time) www. bork. embl-heidelberg. de
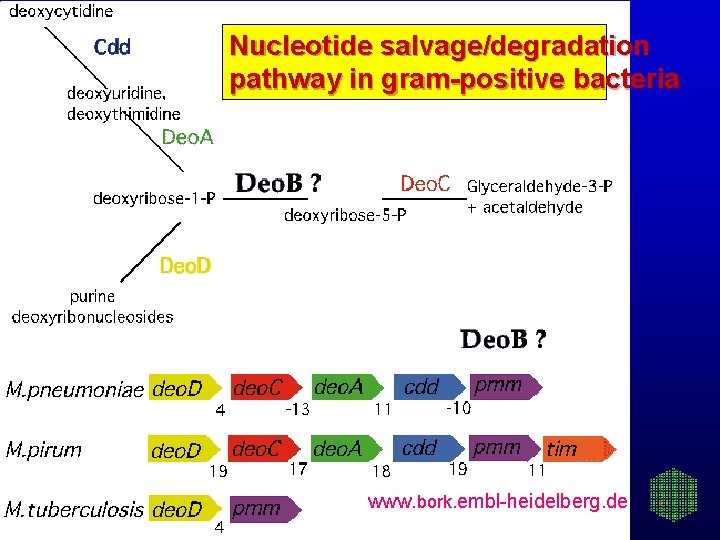
Nucleotide salvage/degradation pathway in gram-positive bacteria www. bork. embl-heidelberg. de
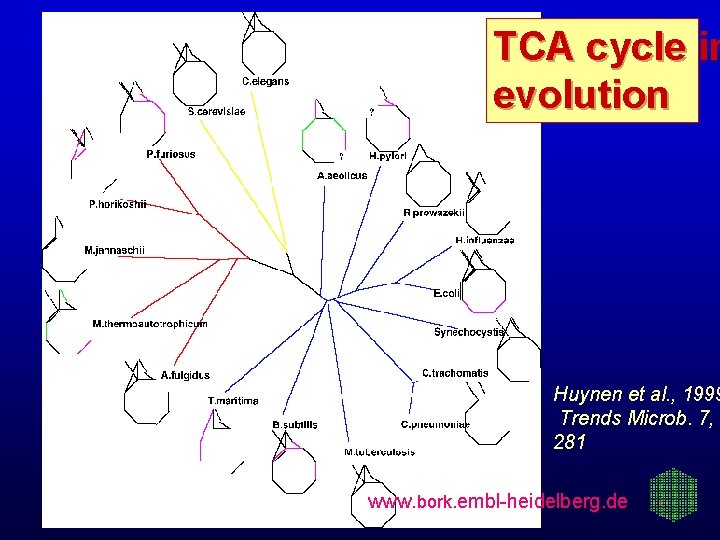
TCA cycle in evolution Huynen et al. , 1999 Trends Microb. 7, 281 www. bork. embl-heidelberg. de
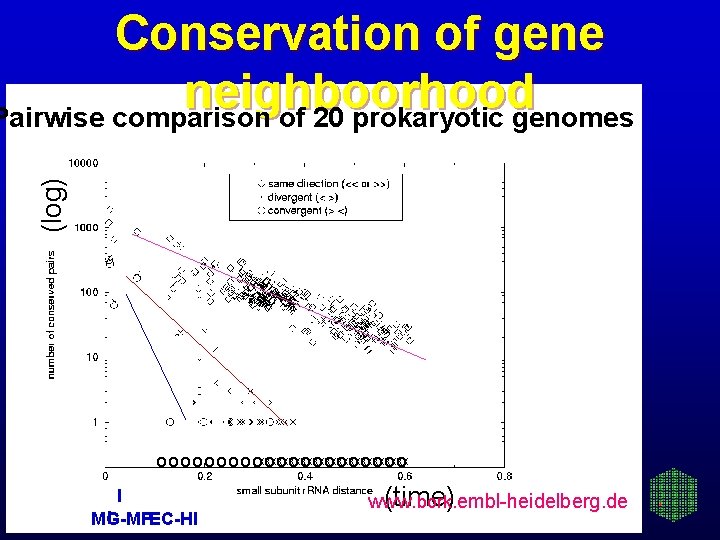
(log) Conservation of gene neighboorhood Pairwise comparison of 20 prokaryotic genomes o o o o oxxxxxxxxxxxxx oooooo I I MG-MP EC-HI (time) www. bork. embl-heidelberg. de
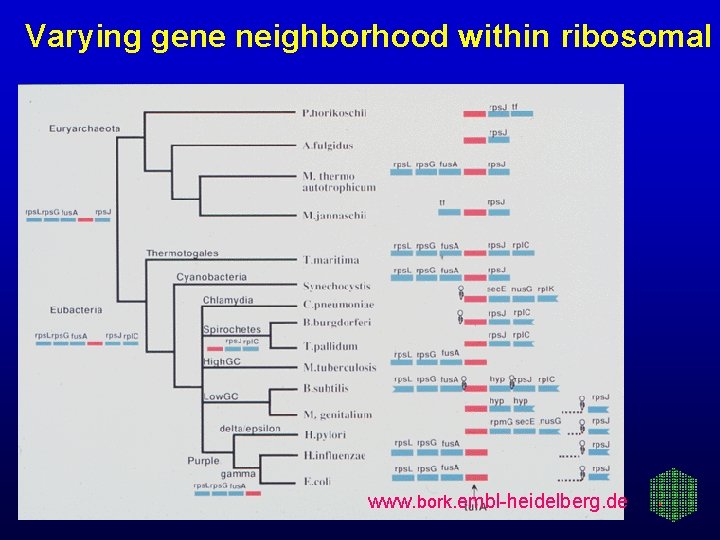
Varying gene neighborhood within ribosomal www. bork. embl-heidelberg. de
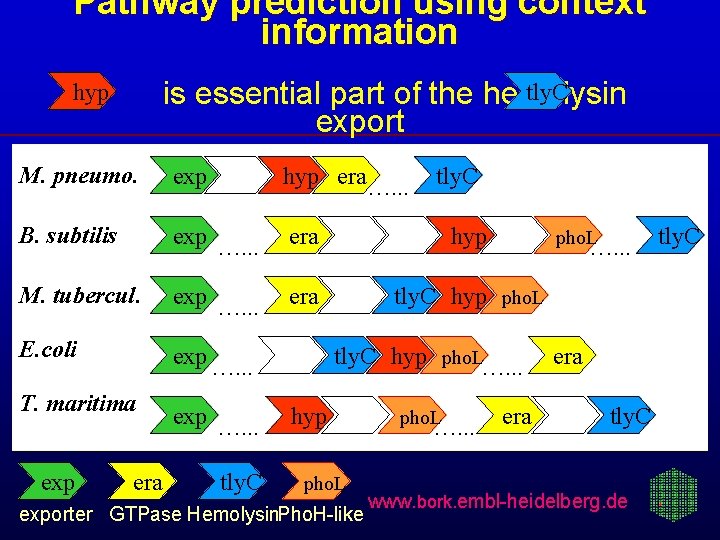
Pathway prediction using context information tly. C is essential part of the hemolysin export hyp M. pneumo. exp hyp era…. . . B. subtilis exp …. . . era hyp M. tubercul. exp …. . . era tly. C …. . . hyp pho. L E. coli exp …. . . T. maritima exp era exp …. . . tly. C pho. L …. . . tly. C…. . . hyp pho. L exporter GTPase Hemolysin. Pho. H-like pho. L …. . . era tly. C www. bork. embl-heidelberg. de tly. C

www. bork. embl-heidelberg. de/STR Tryptopha n biosynthe Snel et al. NAR 28(00)3442 sis www. bork. embl-heidelberg. de/STRING server for context retrieval
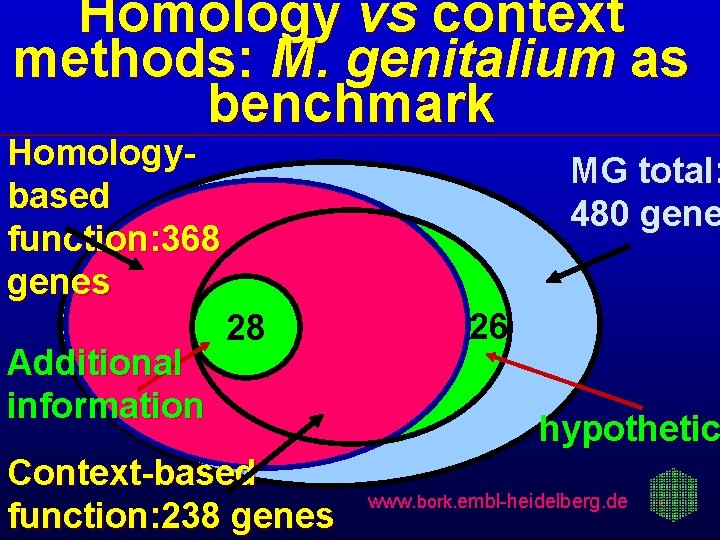
Homology vs context methods: M. genitalium as benchmark Homologybased function: 368 genes Additional information MG total: 480 gene 28 33 Context-based function: 238 genes 26 hypothetic www. bork. embl-heidelberg. de

www. bork. embl-heidelberg. de Frank* (D) Yan (C) Peer (D) Martijn (NL) Gert* (D) Shamil (RU) Vassily (RU), Ina* (D) Birgit* (D) Tobias (D) Richard (UK) Luis* (E) Mikita (J) Miguel (E) Jörg* (D) *left EMB Warren (US) Berend (NL) +Thomas* (D), David (E), Ivica (Hr), Carolina (E), www. bork. embl-heidelberg. de Steffen (D), Francesca (I), Jan (D)
www. bork. embl-heidelberg. de
- Slides: 61