From DNA to Protein By RNA Transcription Similarity

분자생물학 From DNA to Protein By 배형섭
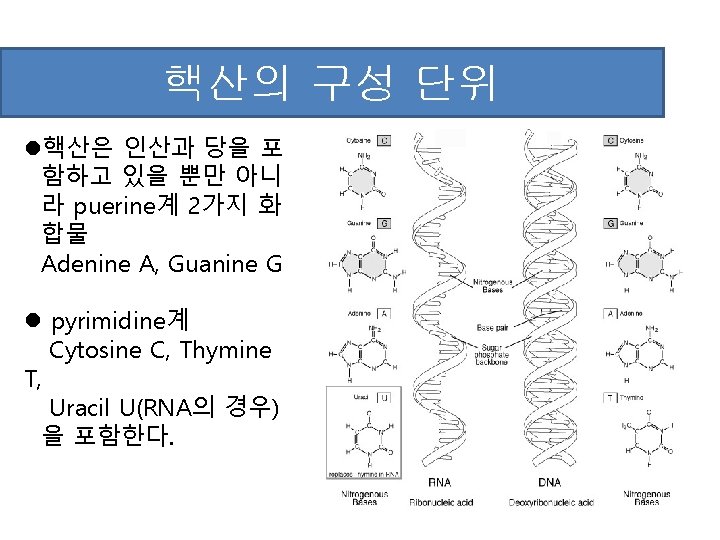
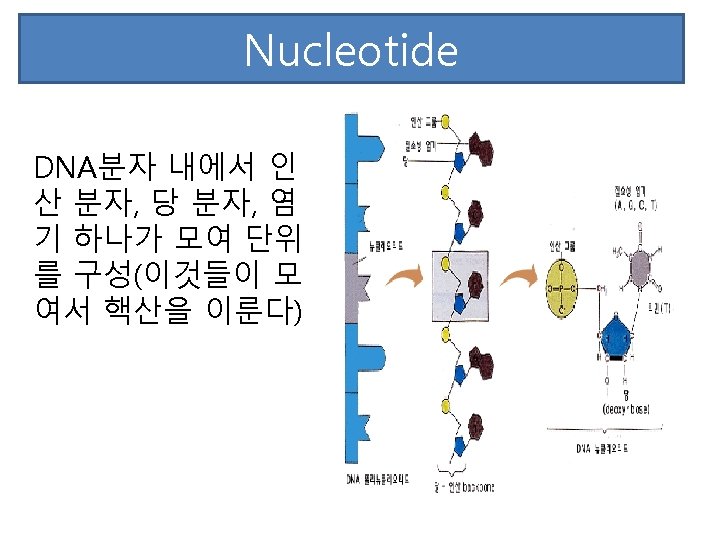
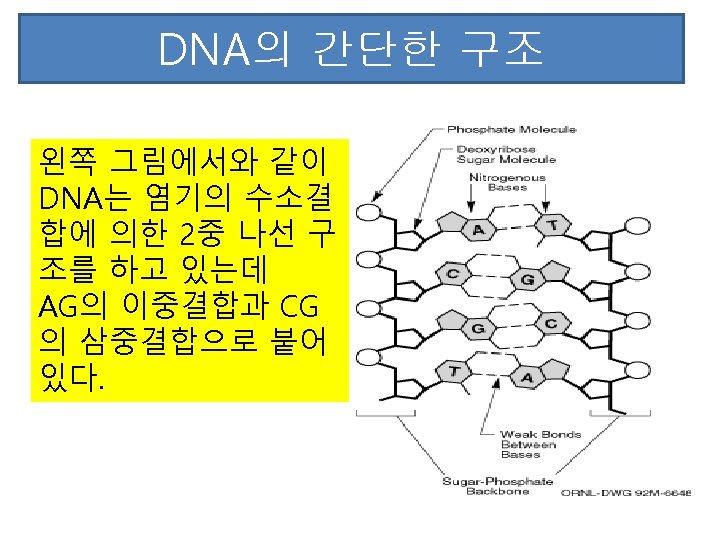

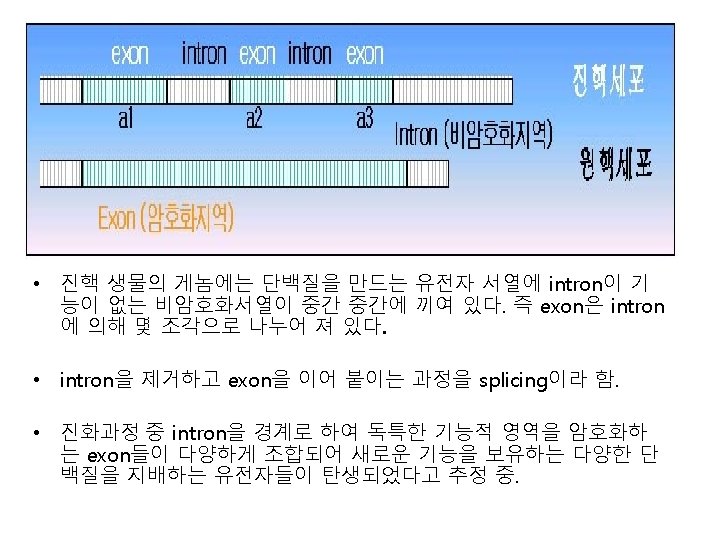

RNA
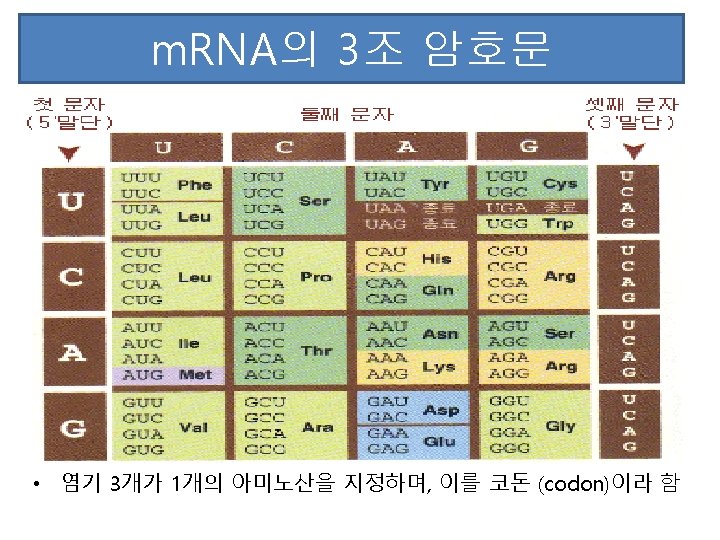
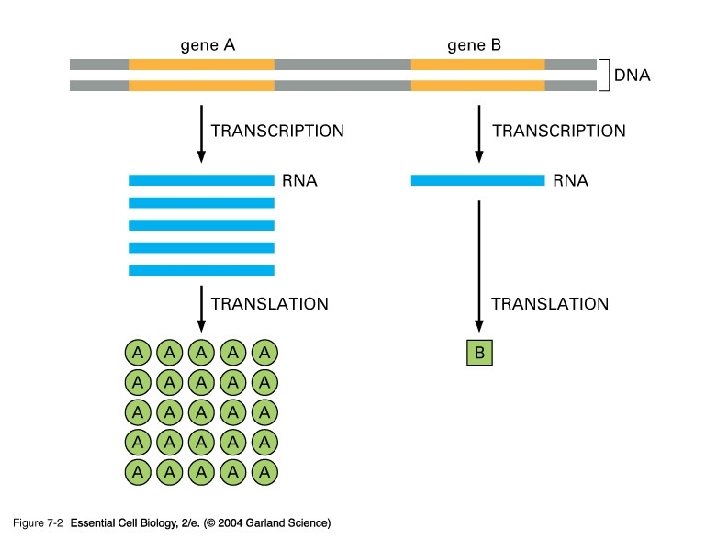
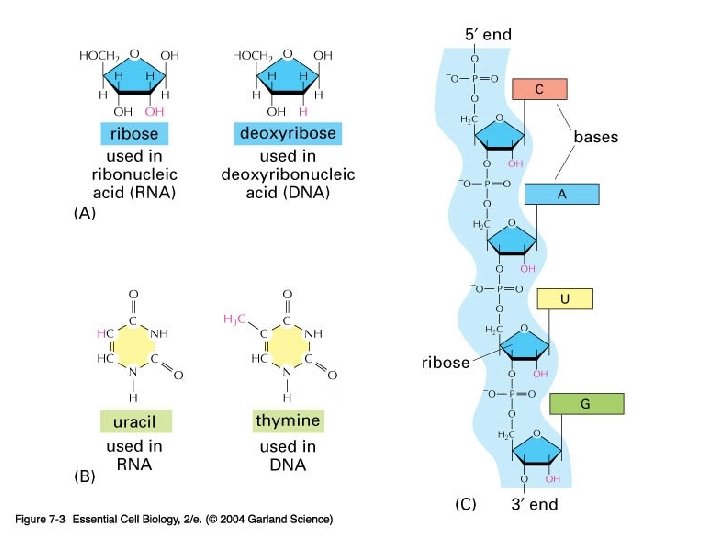
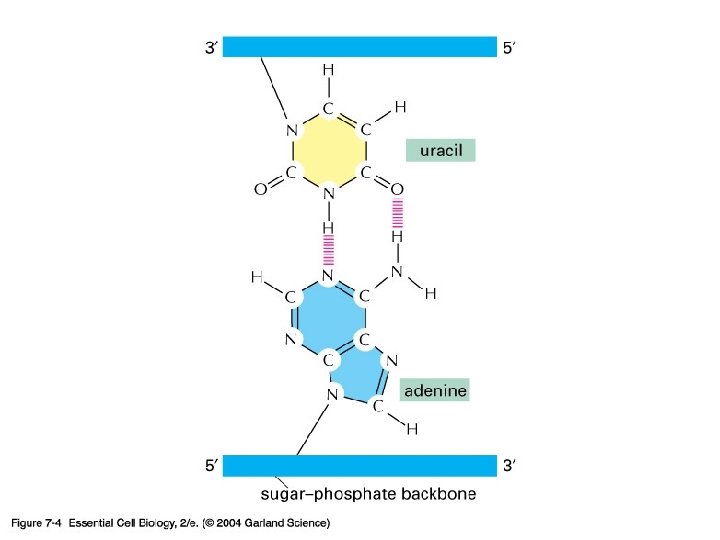


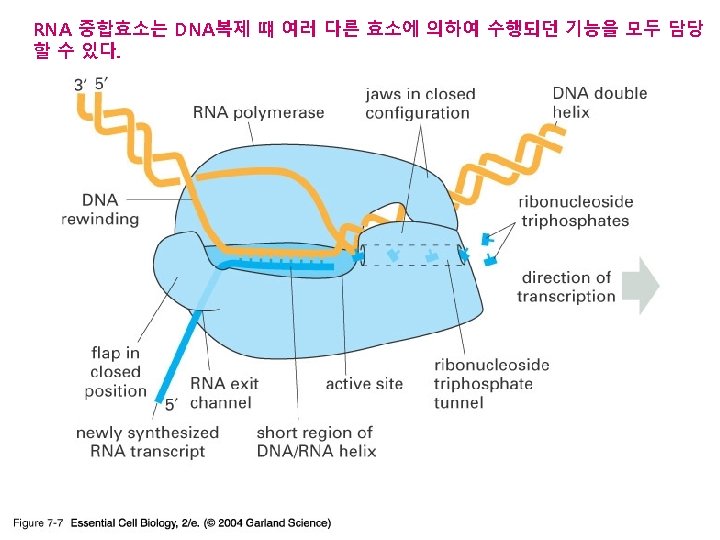



Transcription

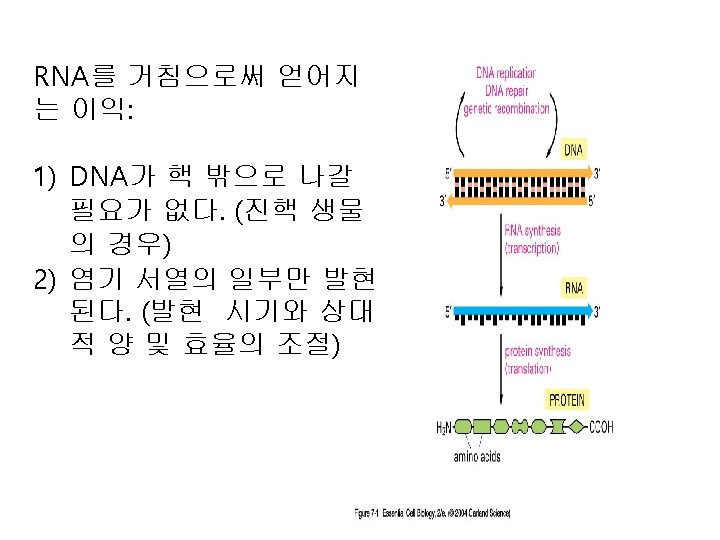
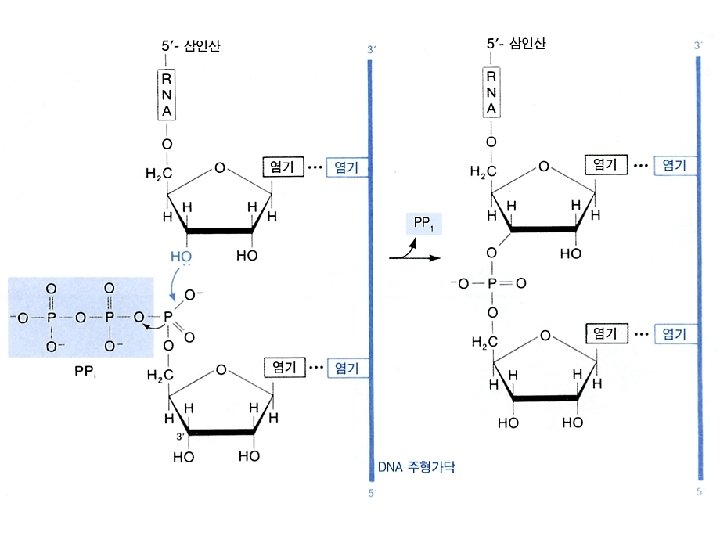
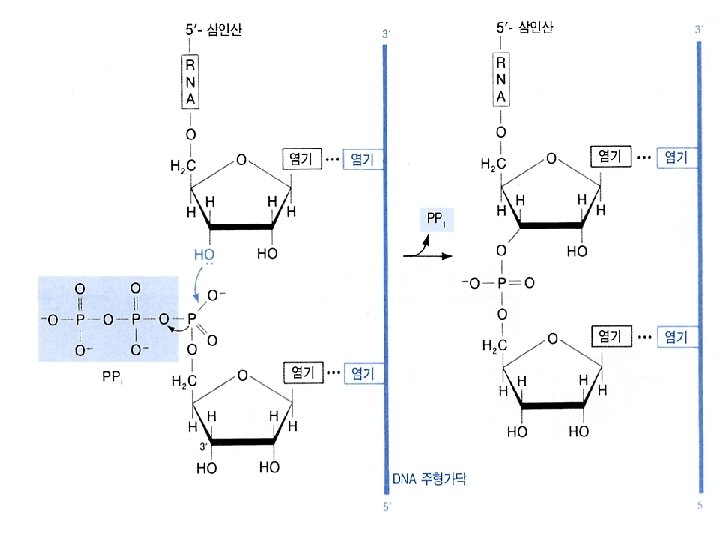
Similarity between replication and transcription l. Both processes use DNA as the template l Phophodiester bonds are formed in both cases. l Both synthesis directions are from 5 to 3

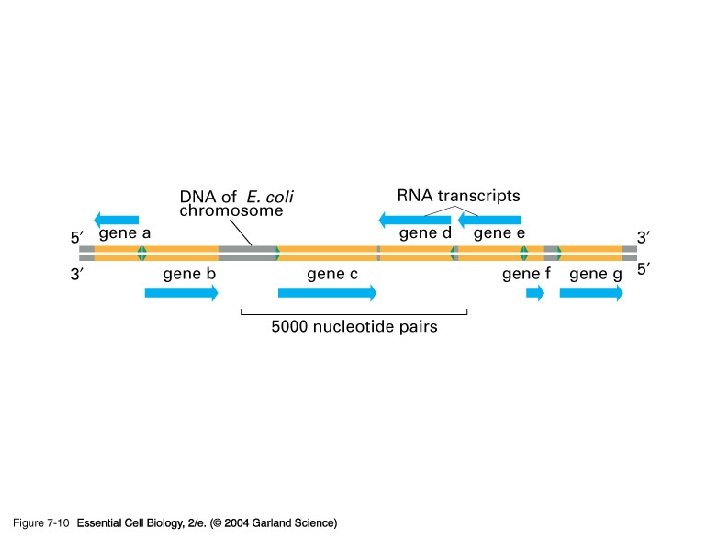
Template and Enzyme The whole genome of DNA needs to be replicated, but only small portion of genome is transcribed in response to the development requirement, physiological need and environmental changes. DNA regions that can be transcribed into RNA are called structural genes

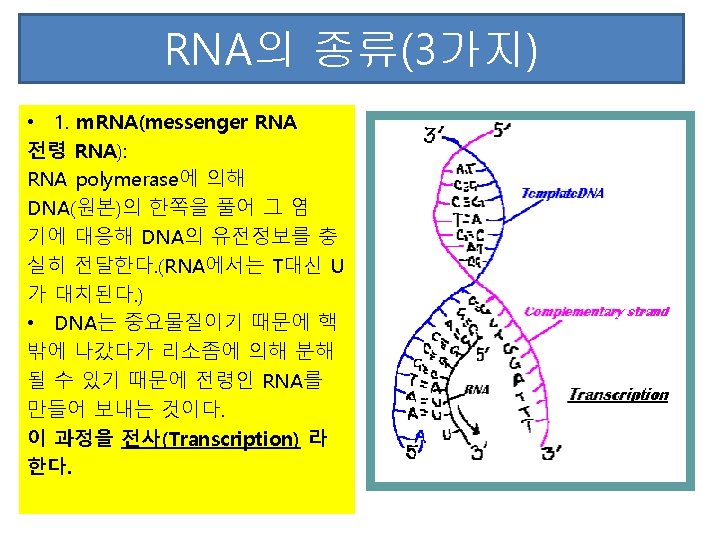
Template l. The template strand is the strand from which the RNA is actually transcribed. It is also termed as antisense strand. l The coding strand is the strand whose base sequence specifies the amino acid sequence of the encoded protein. Therefore, it is also called as sense strand

Asymmetric transcription l Only the tempate strand is used for the transcription, but the coding strand is not. l Both strands can be used as the templates. l The transcription direction on different strands is opposite. l This feature is referred to as the asymmetric transcription.

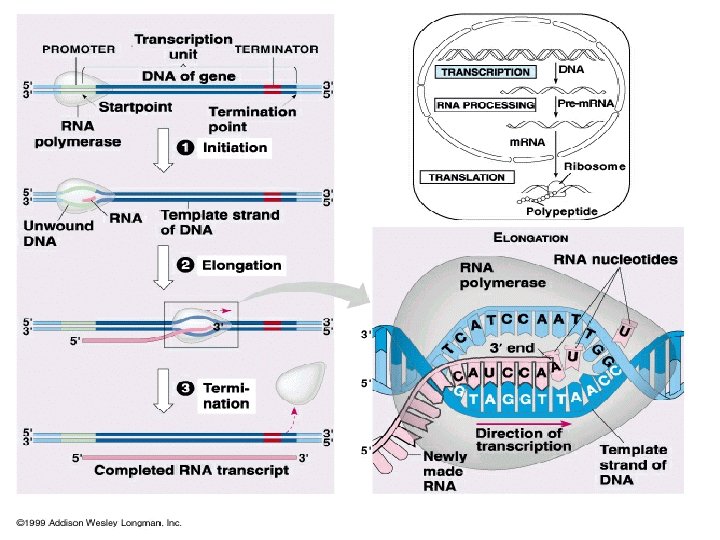
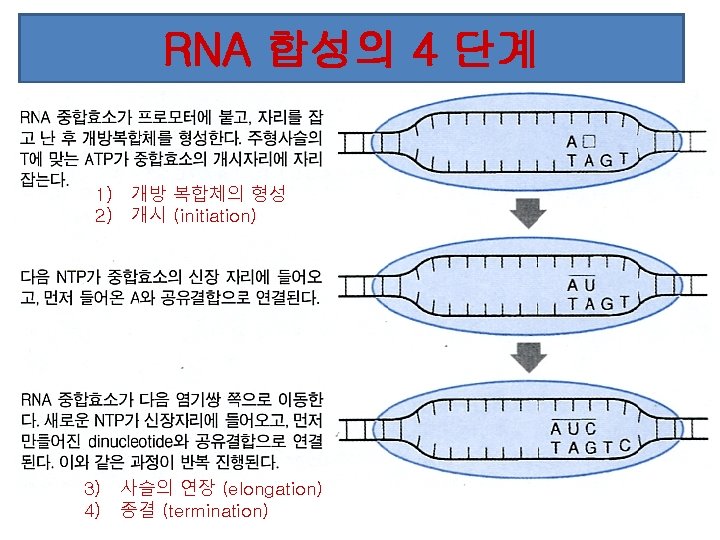
Transcription process l. RNA polymerase (an enzyme) attaches to DNA at a special sequence that serves as a “start signal”. l. The DNA strands are separated and one strand serves as a template. l The RNA bases attach to the complementary DNA template, thus synthesizing m. RNA. l. The RNA polymerase recognizes a termination site on the DNA molecule and releases the new m. RNA molecule.
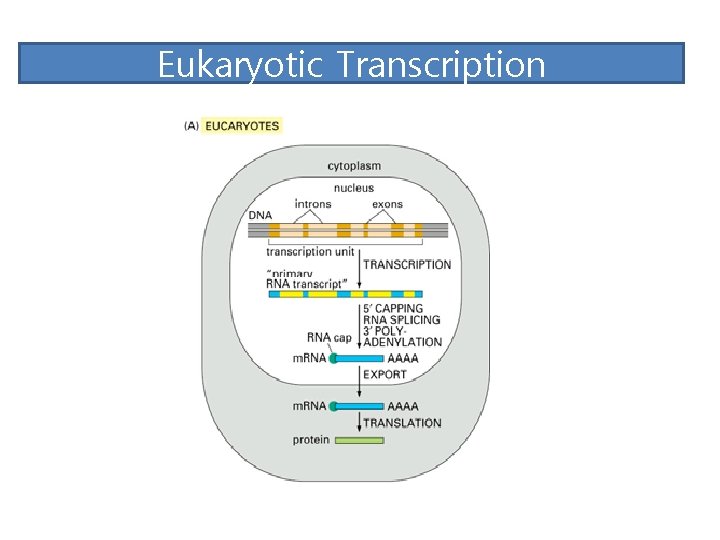
Eukaryotic Transcription
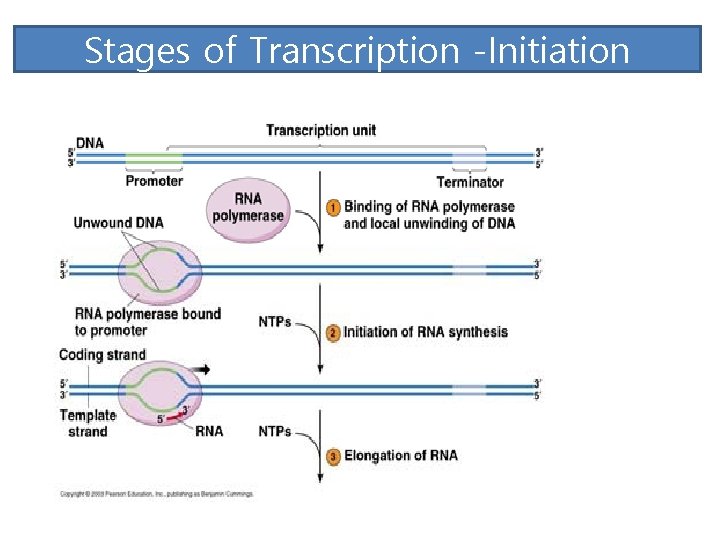
Stages of Transcription -Initiation
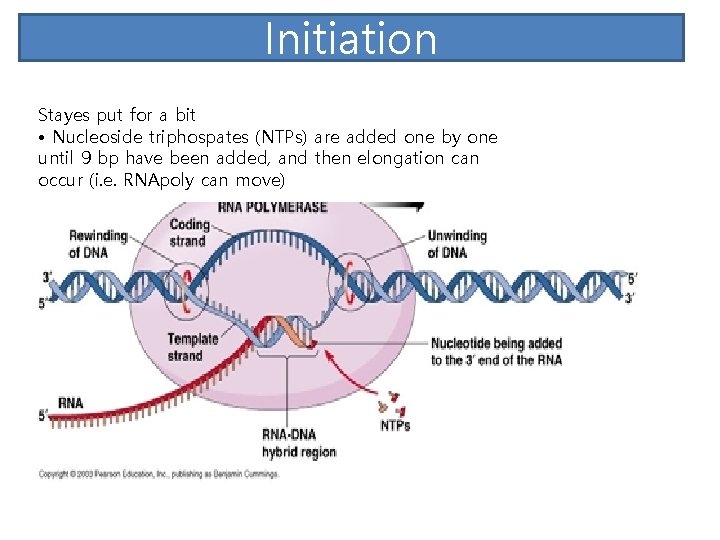
Initiation Stayes put for a bit • Nucleoside triphospates (NTPs) are added one by one until 9 bp have been added, and then elongation can occur (i. e. RNApoly can move)

RNA Polymerase l The enzyme responsible for the RNA synthesis is DNA-dependent RNA polymerase. l The prokaryotic RNA polymerase is a multiplesubunit protein of ~480 k. D. l Eukaryotic systems have three kinds of RNA polymerases, each of which is a multiple-subunit protein and responsible for transcription of different RNAs.
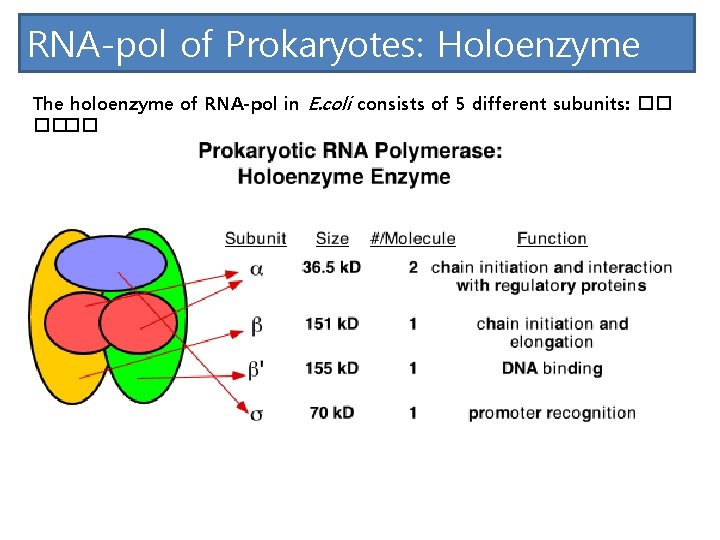
RNA-pol of Prokaryotes: Holoenzyme The holoenzyme of RNA-pol in E. coli consists of 5 different subunits: �� ����

• Rifampicin, a therapeutic drug for tuberculosis treatment, can bind specifically to the subunit of RNApol, and inhibit the RNA synthesis. • RNA-pol of other prokaryotic systems is similar to that of E. coli in structure and functions
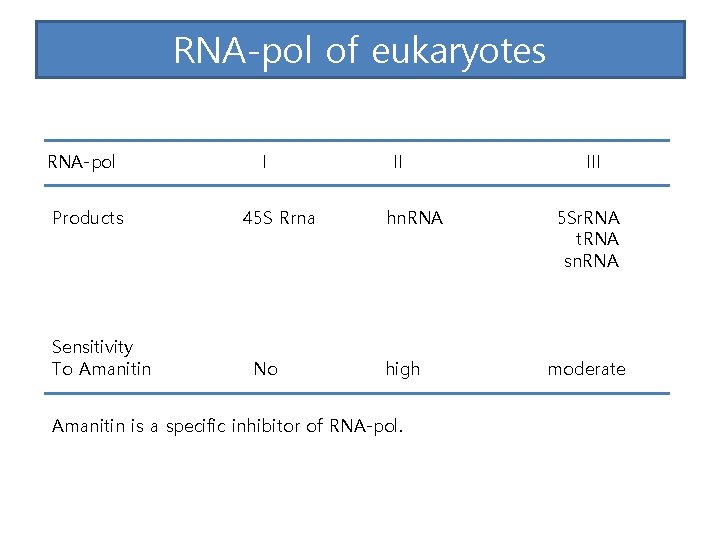
RNA-pol of eukaryotes RNA-pol Products Sensitivity To Amanitin I 45 S Rrna No II hn. RNA high Amanitin is a specific inhibitor of RNA-pol. III 5 Sr. RNA t. RNA sn. RNA moderate

Three RNA polymerases • • One for each major type of RNApoly I makes pre-r. RNApoly II makes pre-m. RNApoly III makes pre-t. RNA Each polymerase has a different promoter structure

RNApoly II promoter • • Initiator sequence surrounding the start point TATA box at about – 25 bp TATA+Initiator = core promoter A transcription factor binds to the TATA box before RNAPoly II

Bacterial Promoters • • A promoter is where RNApoly binds Template strand versus Coding strand Upstream – in the 5’ direction on the coding strand Consensus sequences
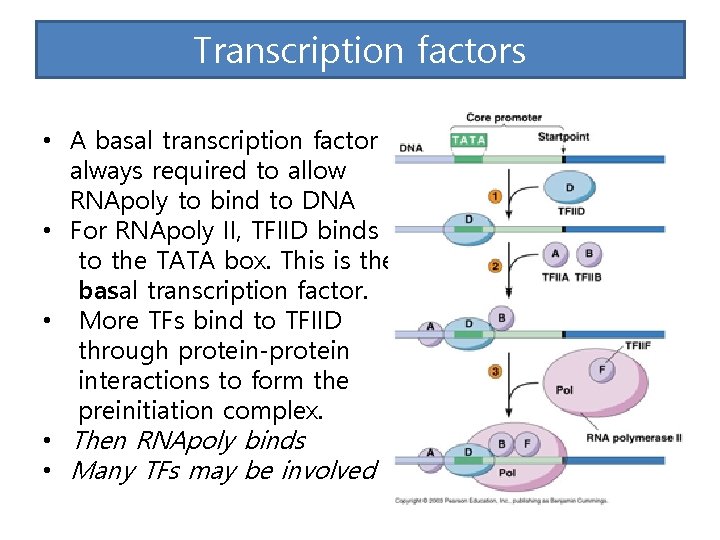
Transcription factors • A basal transcription factor is always required to allow RNApoly to bind to DNA • For RNApoly II, TFIID binds to the TATA box. This is the basal transcription factor. • More TFs bind to TFIID through protein-protein interactions to form the preinitiation complex. • Then RNApoly binds • Many TFs may be involved

Recognition of Origins • Each transcriptable region is called operon. • One operon includes several structural genes and upstream regulatory sequences (or regulatory regions). • The promoter is the DNA sequence that RNA-pol can bind. It is the key point for the transcription control.

Promoter • regulatory • sequences structural gene
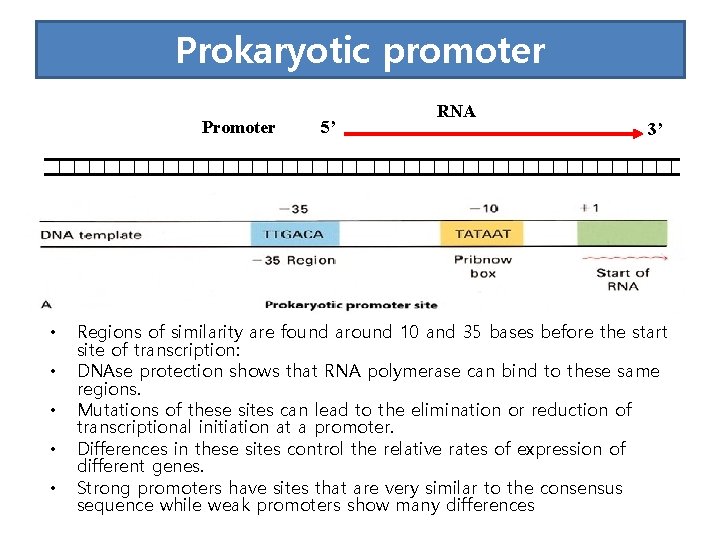
Prokaryotic promoter Promoter • • • 5’ RNA 3’ Regions of similarity are found around 10 and 35 bases before the start site of transcription: DNAse protection shows that RNA polymerase can bind to these same regions. Mutations of these sites can lead to the elimination or reduction of transcriptional initiation at a promoter. Differences in these sites control the relative rates of expression of different genes. Strong promoters have sites that are very similar to the consensus sequence while weak promoters show many differences

Consensus Sequence

• The -35 region of TTGACA sequence is the recognition site and the binding site of RNA-pol. • The -10 region of TATAAT is the region at which a stable complex of DNA and RNApol is formed.

Transcription Process
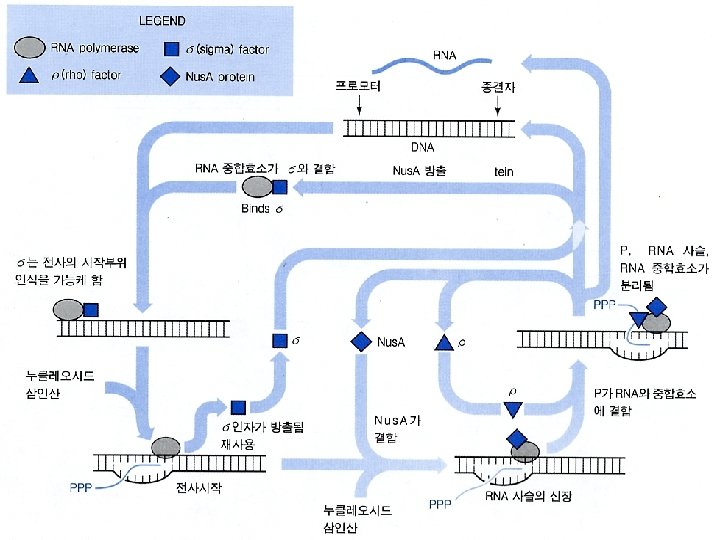
General concepts • Three phases: initiation, elongation, and termination. • The prokaryotic RNA-pol can bind to the • DNA template directly in the transcription process. • The eukaryotic RNA-pol requires cofactors to bind to the DNA template together in the transcription process

Transcription of Prokaryotes • Initiation phase: RNA-pol recognizes the promoter and starts the transcription. • Elongation phase: the RNA strand is continuously growing. • Termination phase: the RNA-pol stops synthesis and the nascent RNA is separated from the DNA template.
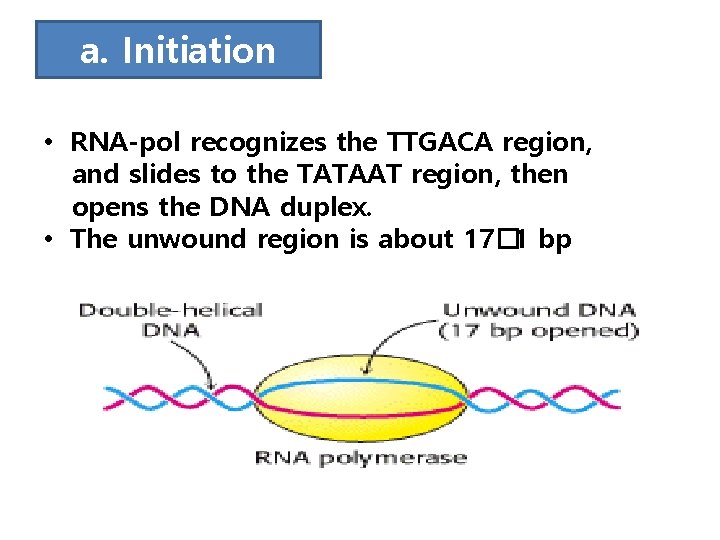
a. Initiation • RNA-pol recognizes the TTGACA region, and slides to the TATAAT region, then opens the DNA duplex. • The unwound region is about 17� 1 bp

• The first nucleotide on RNA transcript is always purine triphosphate. GTP is more often than ATP. • The ppp. Gp. N-OH structure remains on the RNA transcript until the RNA synthesis is completed. • The three molecules form a transcription initiation complex.

• No primer is needed for RNA synthesis. • The subunit falls off from the RNA-pol once the first 3, 5 phosphodiester bond is formed. • The core enzyme moves along the DNA template to enter the elongation phase.

b. Elongation • The release of the subunit causes the conformational change of the core enzyme. The core enzyme slides on the DNA template toward the 3 end. • Free NTPs are added sequentially to the 3 -OH of the nascent RNA strand. (NMP)n + NTP (NMP)n+1 + PPi RNA strand substrate elongated RNA strand

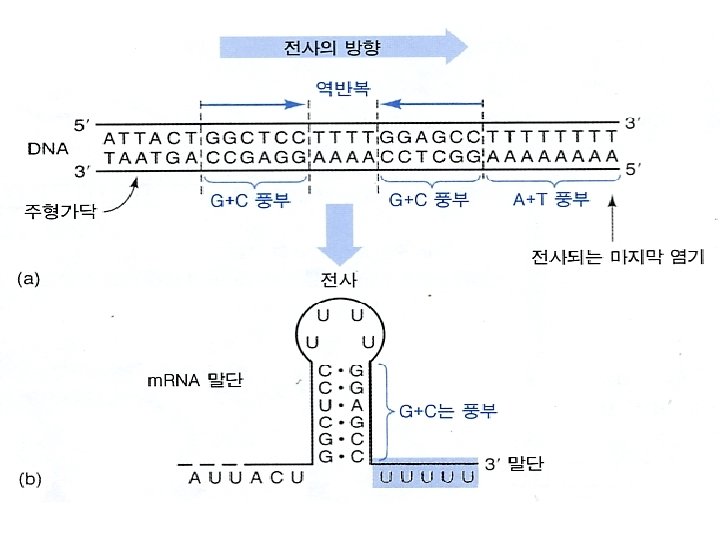
• RNA-pol, DNA segment of ~40 nt and the nascent RNA form a complex called the transcription bubble. • The 3 segment of the nascent RNA hybridizes with the DNA template, and its 5 end extends out the transcription bubble as the synthesis is processing.
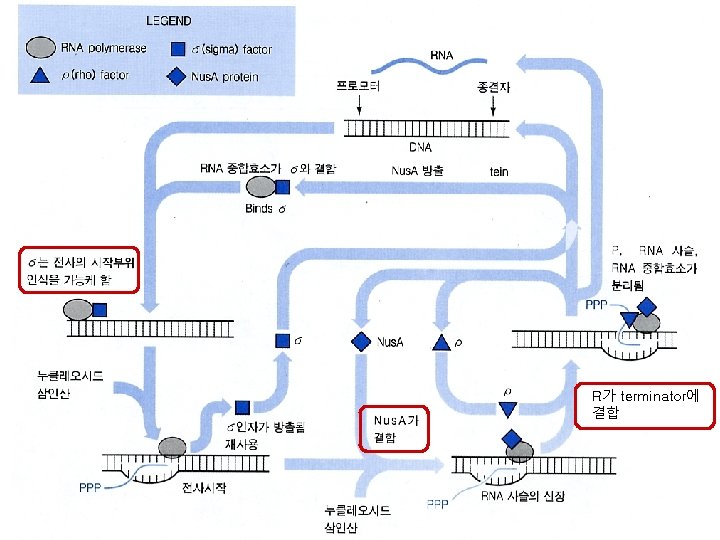
R가 terminator에 결합
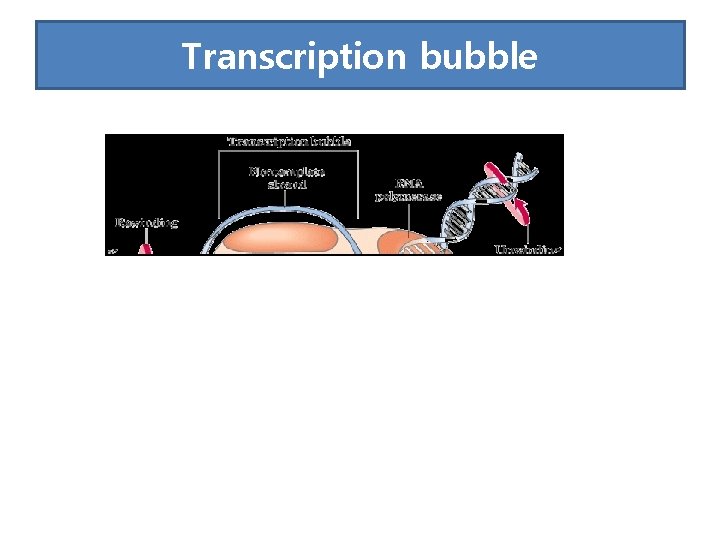
Transcription bubble
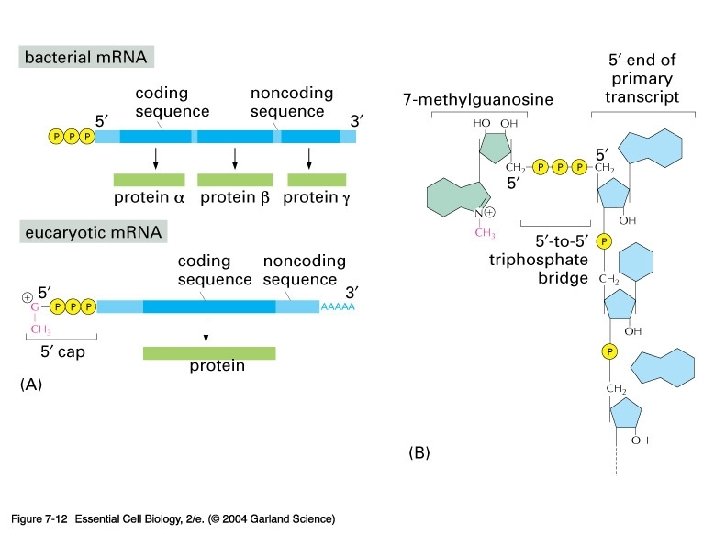

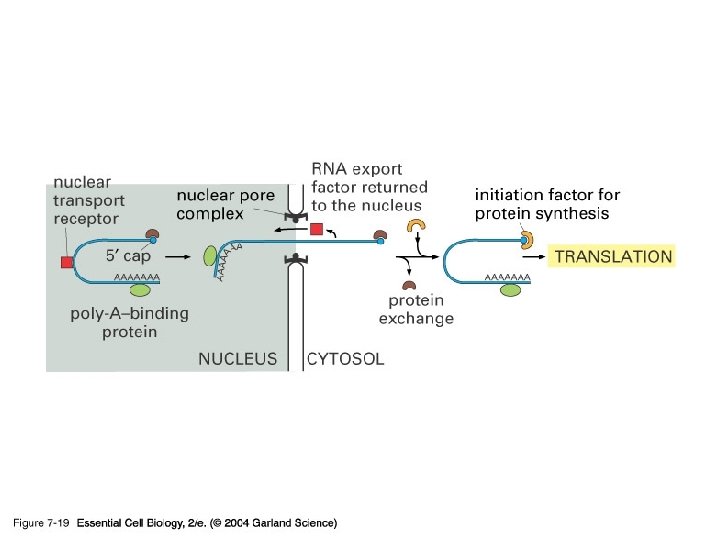
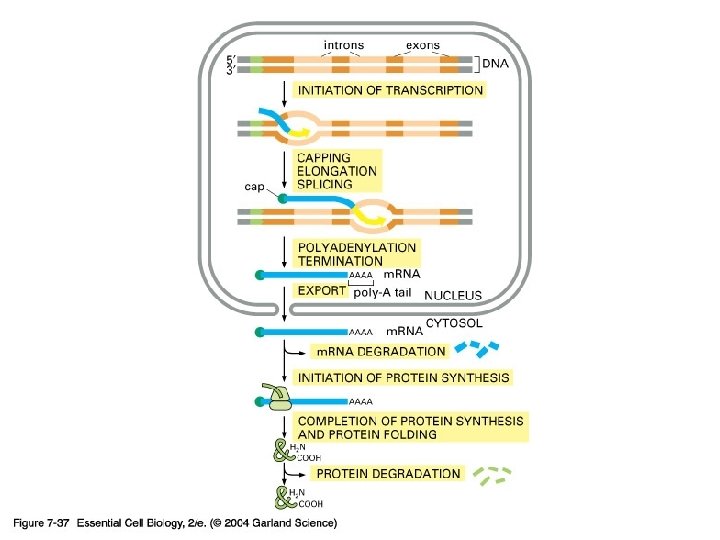
5’-capping과 poly(A) tail 의 기능 -Protection from degradation (due to longer half life---think maternal transcripts) -방향 표지 -Translation 조절에 관여 (고도로 분화된 진핵 생물 유전자 발현 조절의 한 단면)


sn. RNPS (small nuclear ribonucleoprotein particles) , 작은핵 RNA (sn. RNA)와 단백질의 복합체 “Splisosome”을 형성한다. RNA가 효소 작용을 하는 ribozyme의 대표적인 예. Other ribozymes: ribonuclease P ribosomal large subunit (28 S r. RNA) many more to come
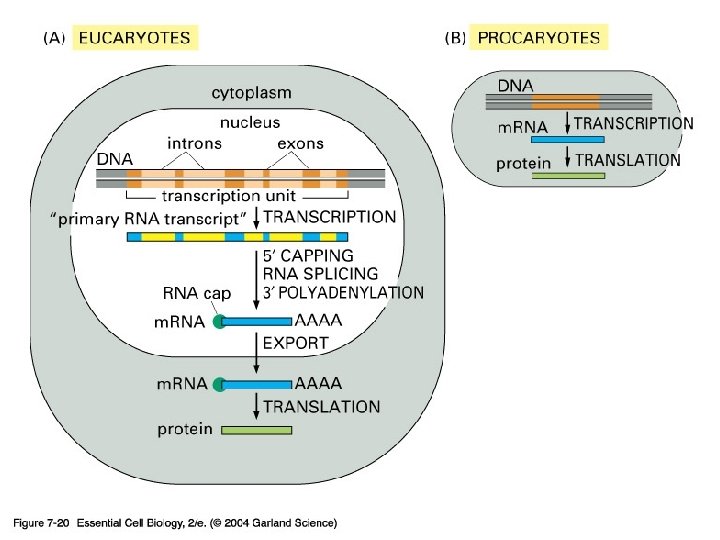
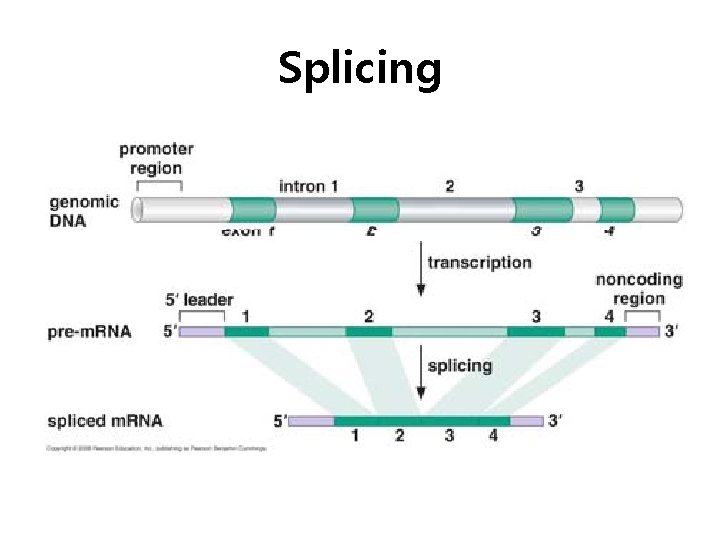
Splicing

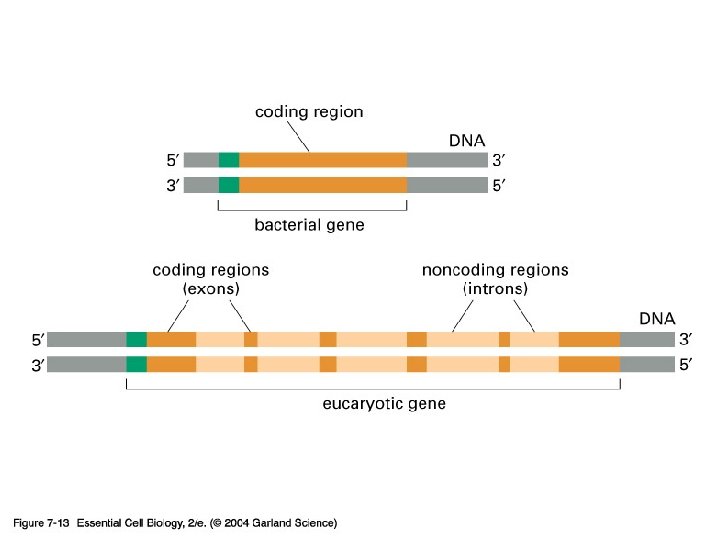
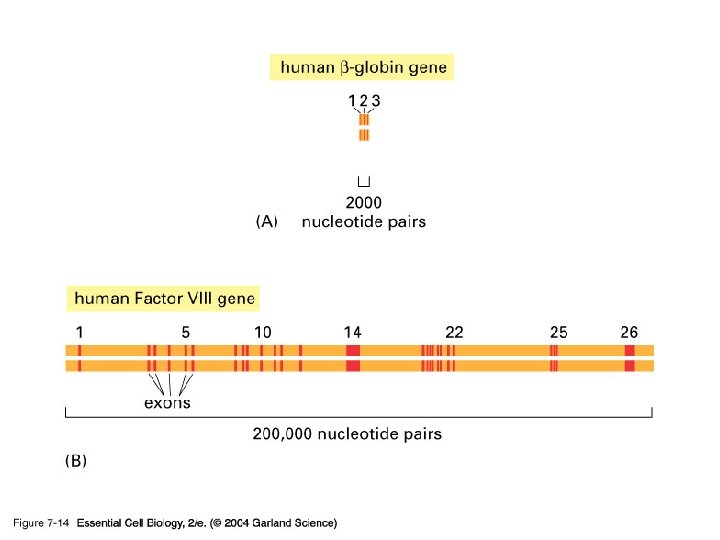
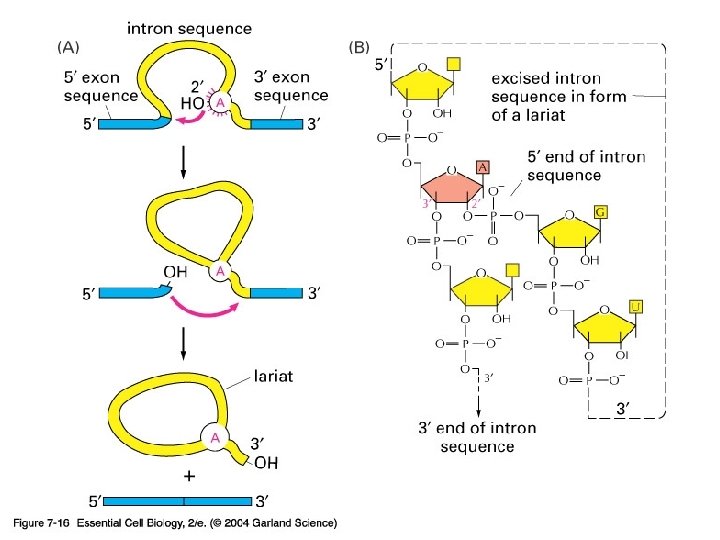
Exon and intron Exons are the coding sequences that appear on split genes and primary transcripts, and will be expressed to matured m. RNA. Introns are the non-coding sequences that are transcripted into primary m. RNAs, and will be cleaved out in the later splicing process.
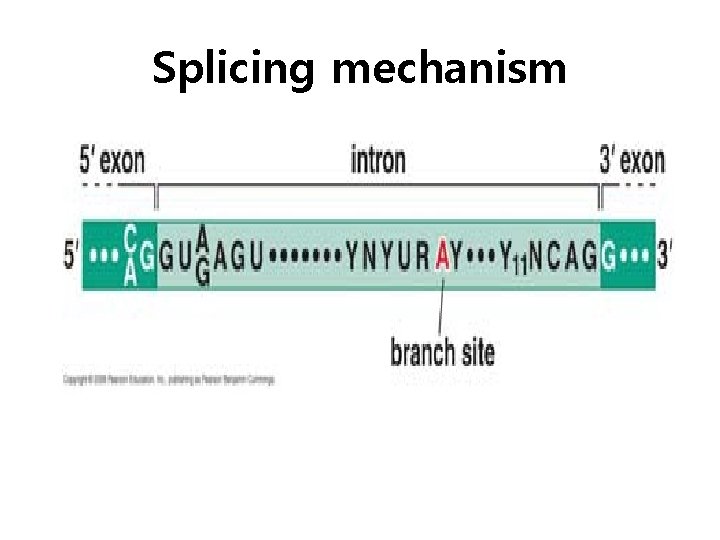
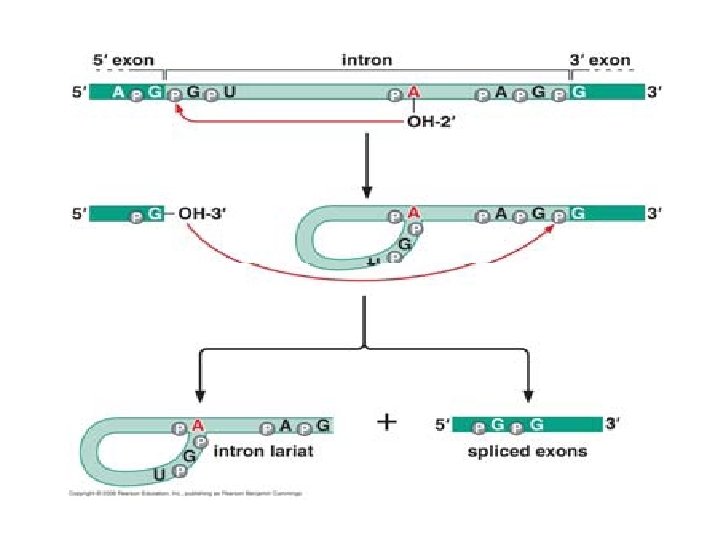
Splicing mechanism
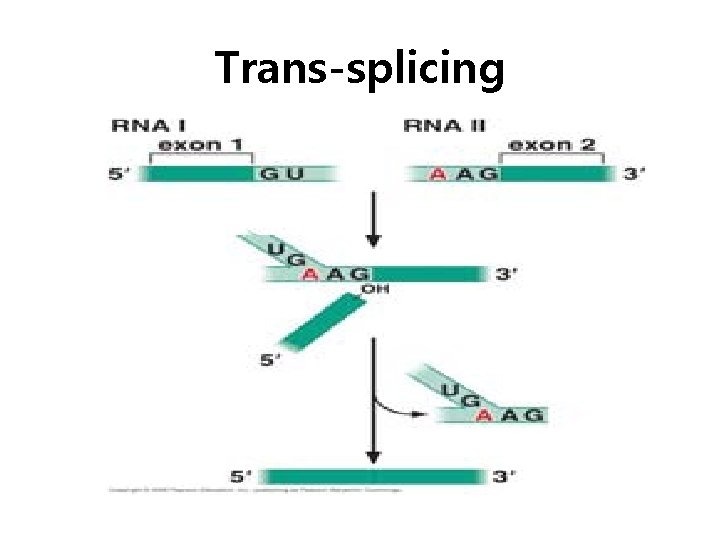
Trans-splicing

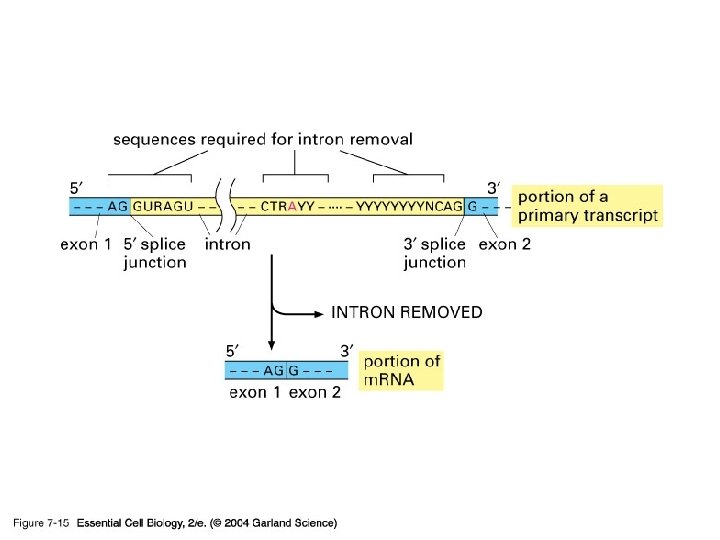
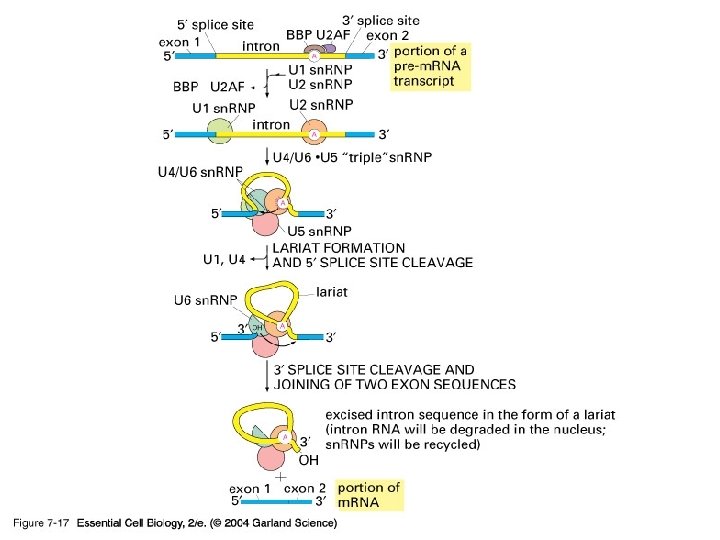
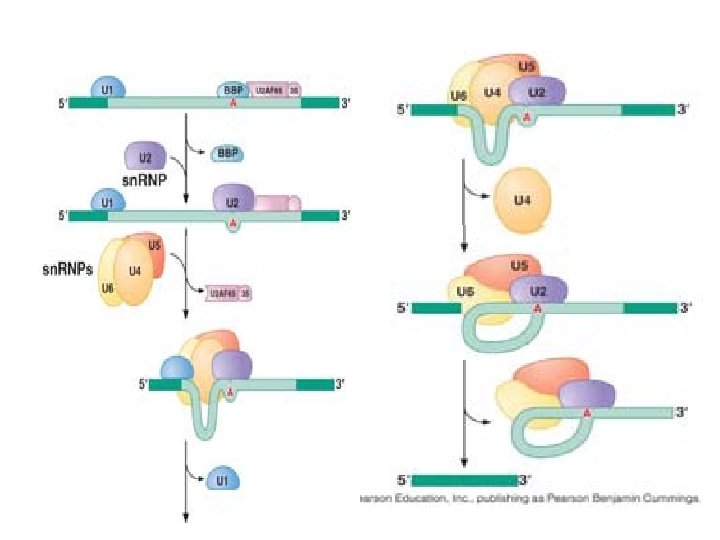
Spliceosome • Splicing introns from m. RNA occurs at short, conserved sequences called splice sites which specify the beginning and ends of introns. • GU on the 5' end and AG on the 3' end are 100% conserved. • Catalyzed by a structure called a spliceosome, composed of protein and RNA (sn. RNA – small nuclear RNA)
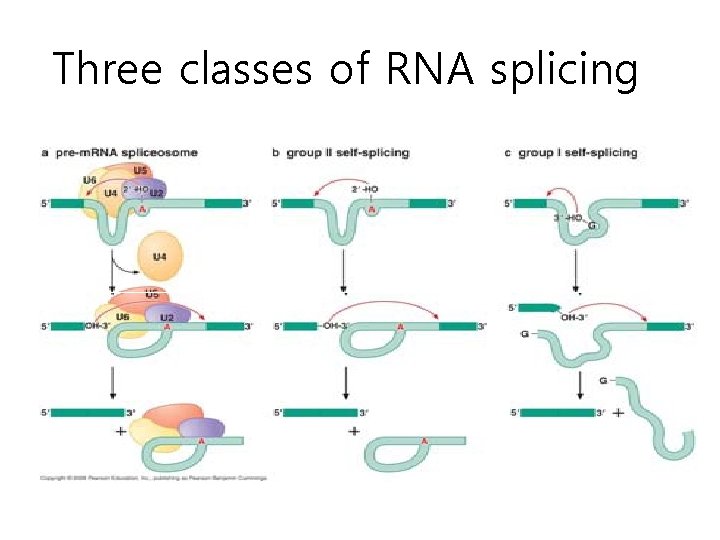
Three classes of RNA splicing
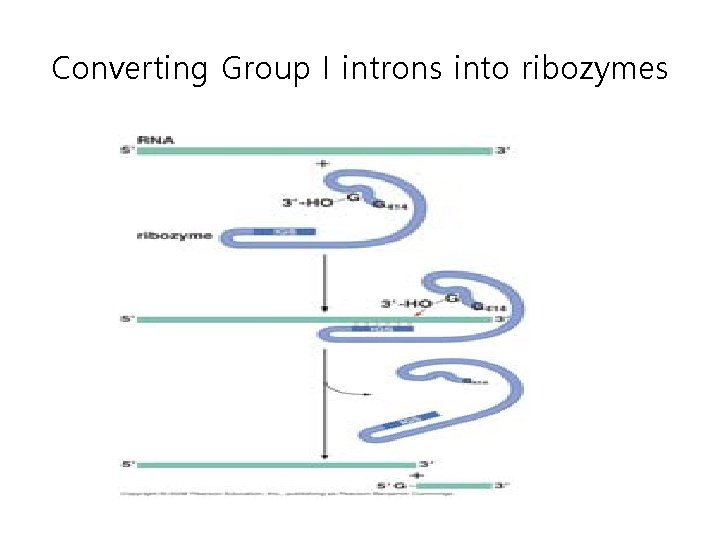
Converting Group I introns into ribozymes
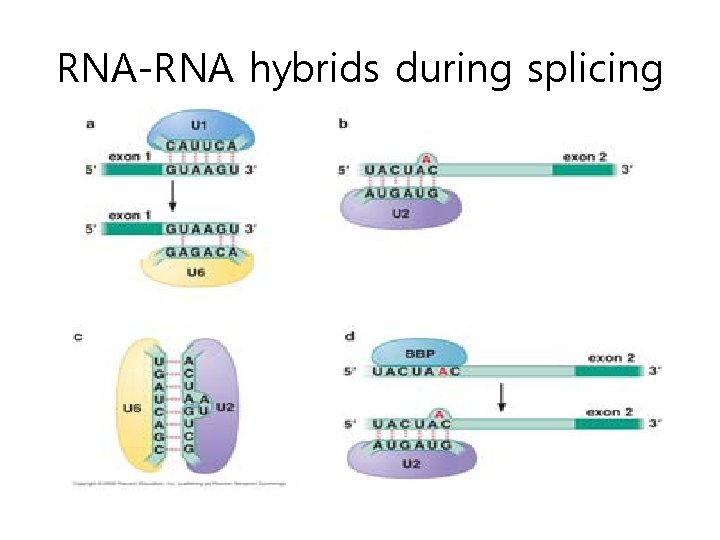
RNA-RNA hybrids during splicing

Errors by mistakes in splice-site selection
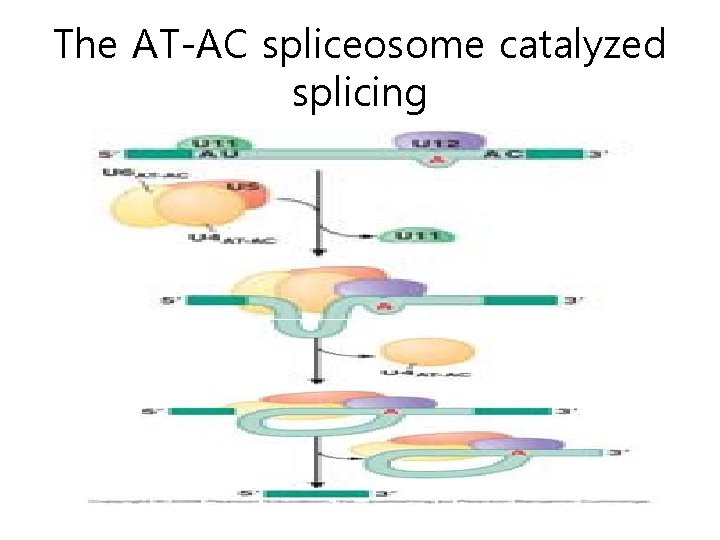
The AT-AC spliceosome catalyzed splicing

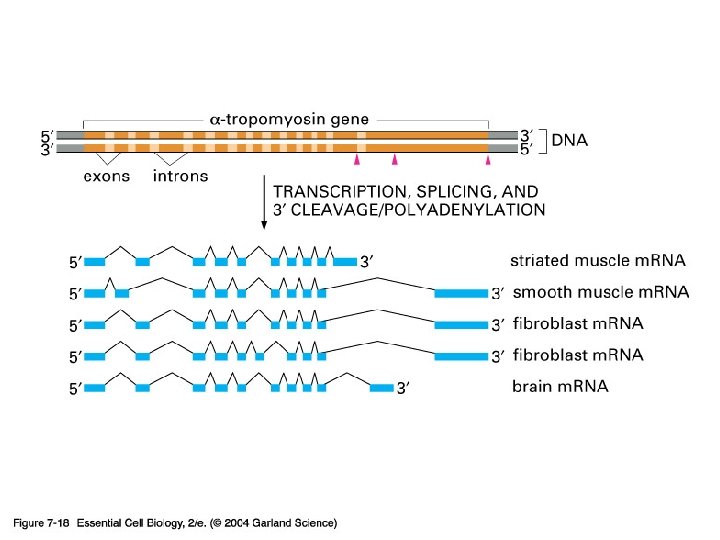
Alternative splicing

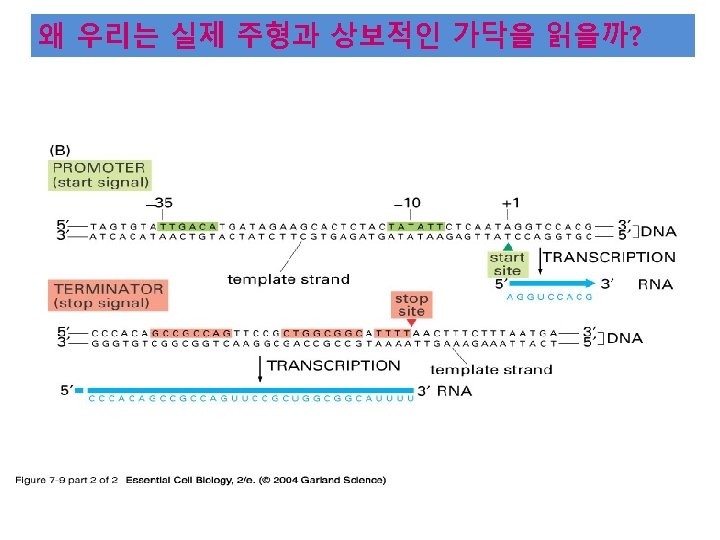

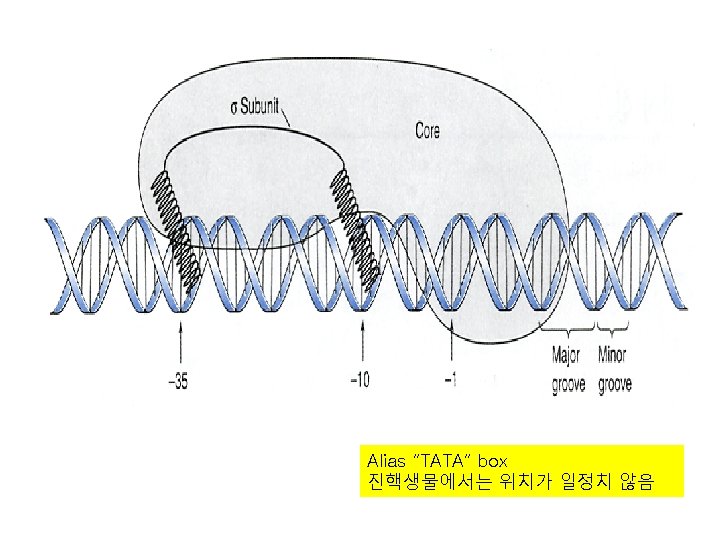

TTGACA TATAAT

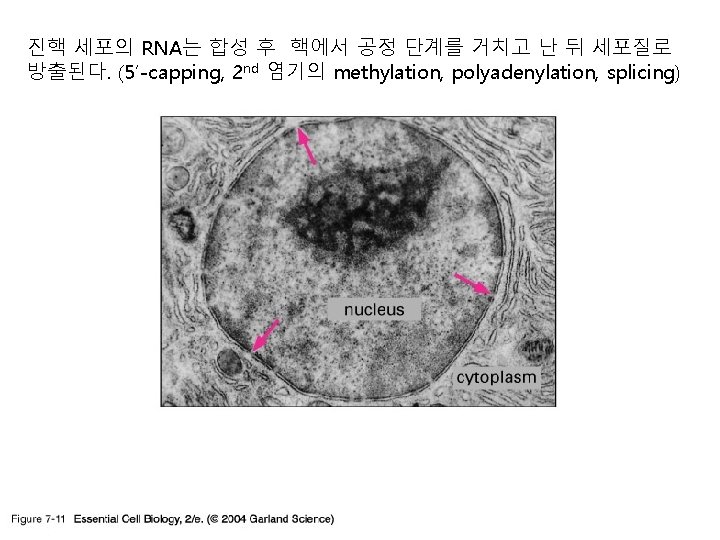

5’-capping과 poly(A) tail 의 기능 -Protection from degradation (due to longer half life---think maternal transcripts) -방향 표지 -Translation 조절에 관여 (고도로 분화된 진핵 생물 유전자 발현 조절의 한 단면)
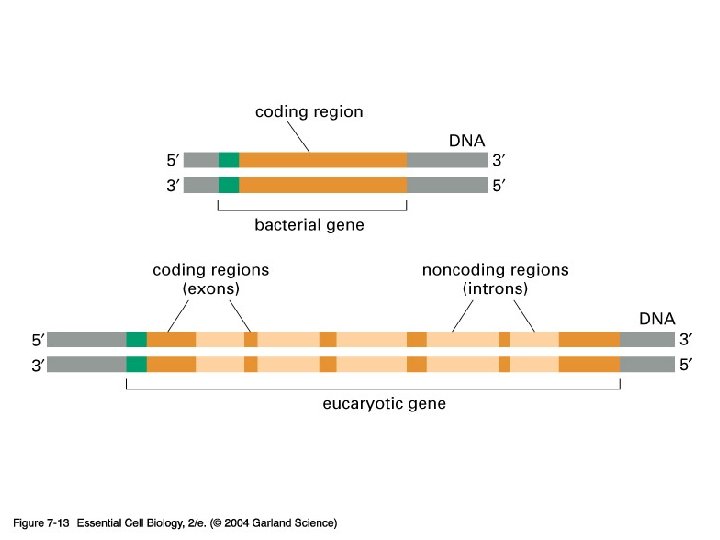

INTRON


sn. RNPS (small nuclear ribonucleoprotein particles) , 작은핵 RNA (sn. RNA)와 단백질의 복합체 “Splisosome”을 형성한다. RNA가 효소 작용을 하는 ribozyme의 대표적인 예. Other ribozymes: ribonuclease P ribosomal large subunit (28 S r. RNA) many more to come

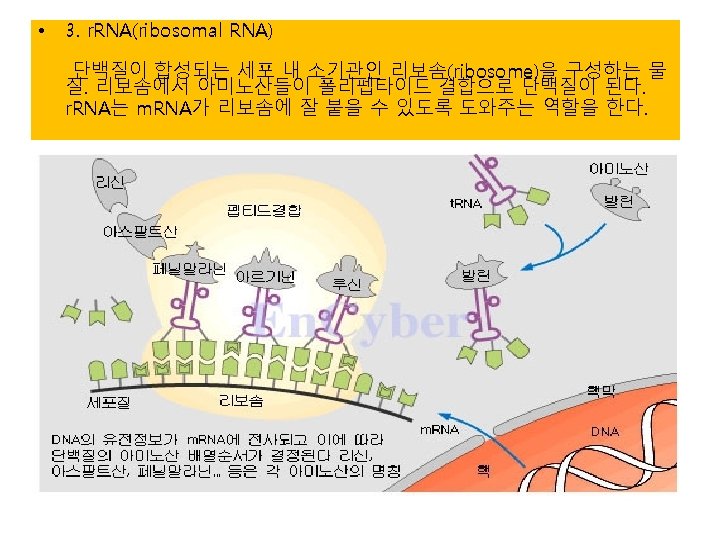
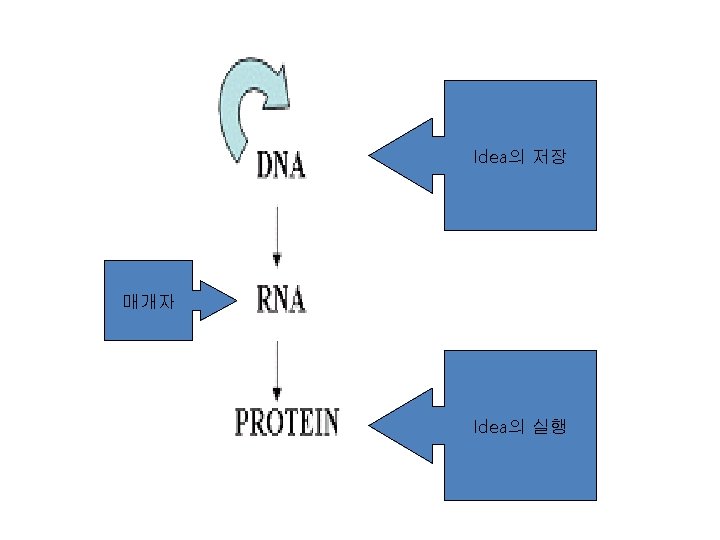
Translation

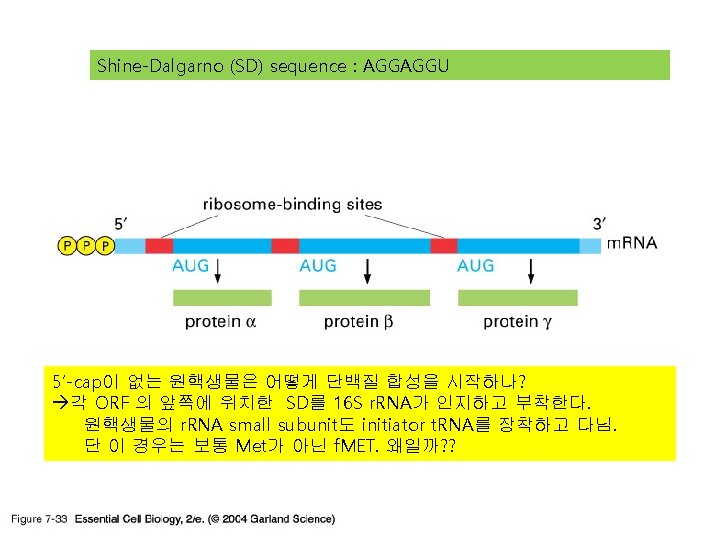
단백질 합성의 세 기구: 1) m. RNA-the template 2) t. RNA-the delieverer 3) r. RNA (ribosome)-the synthesizer

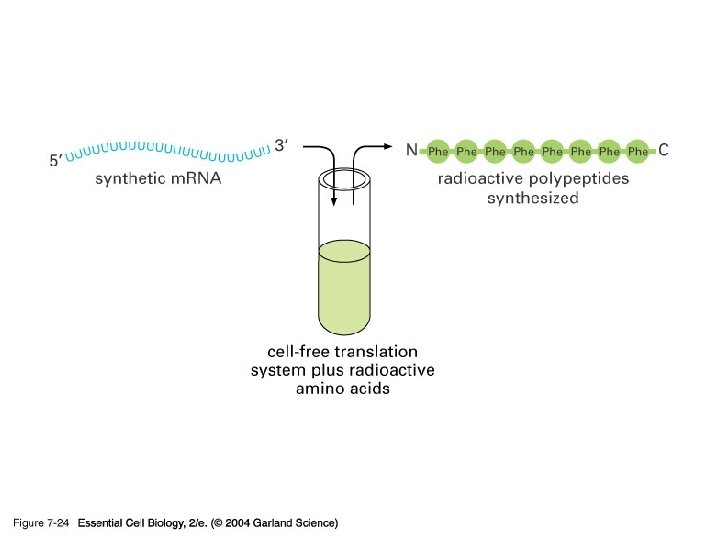
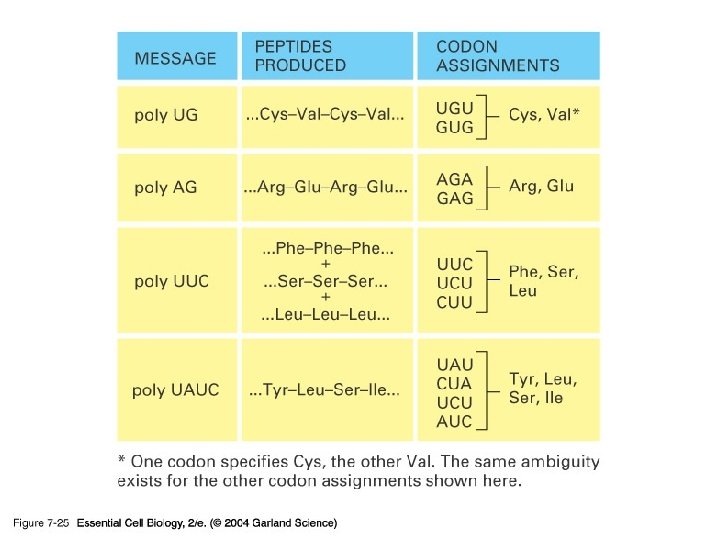
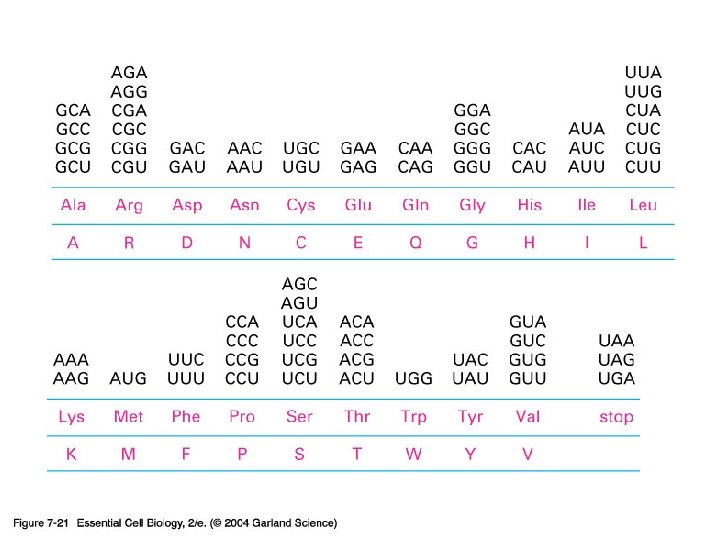
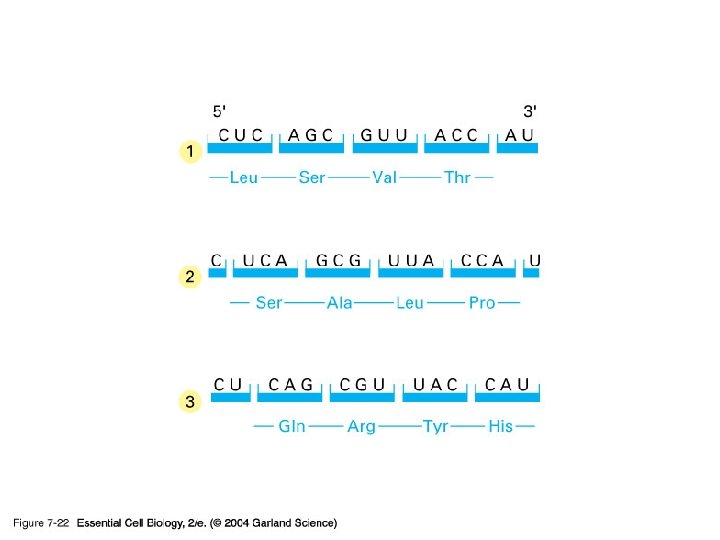
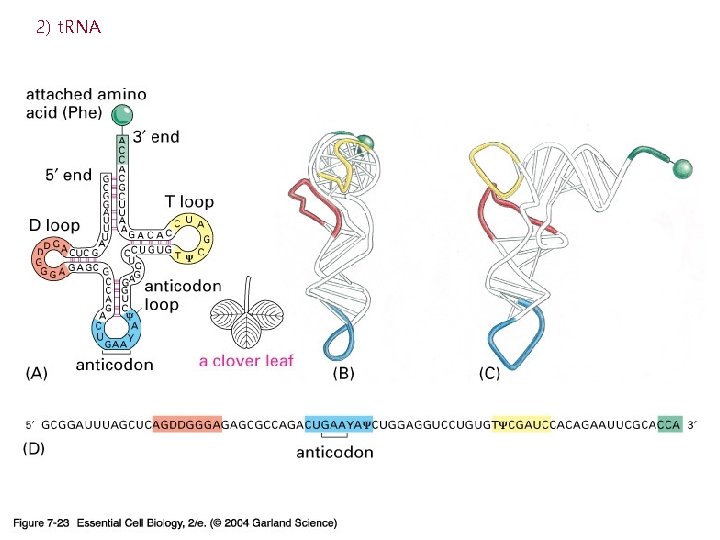
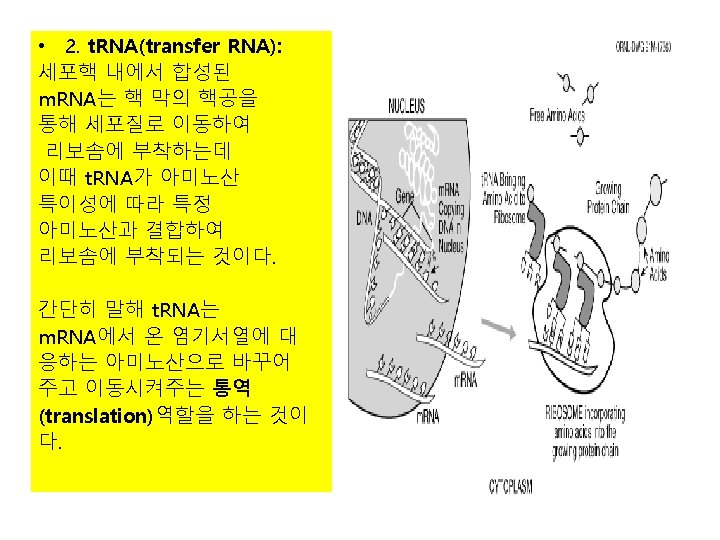
2) t. RNA

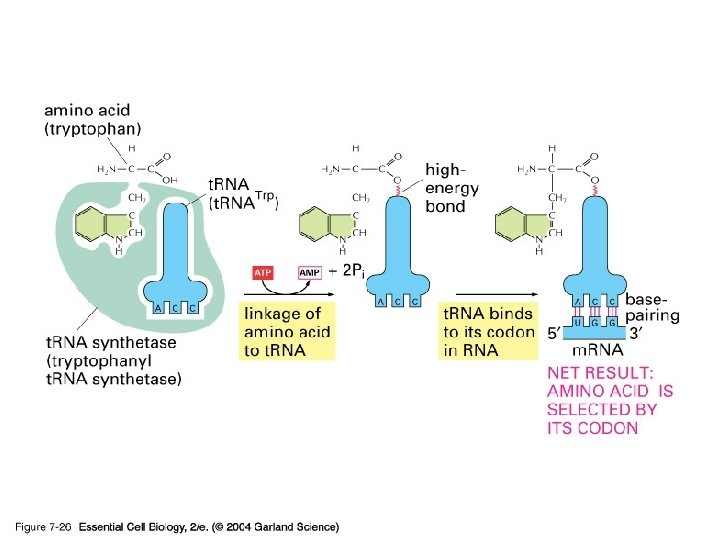

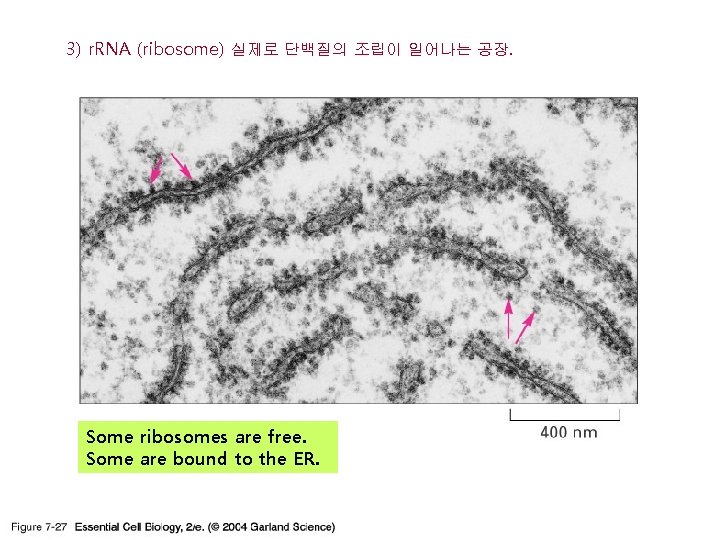
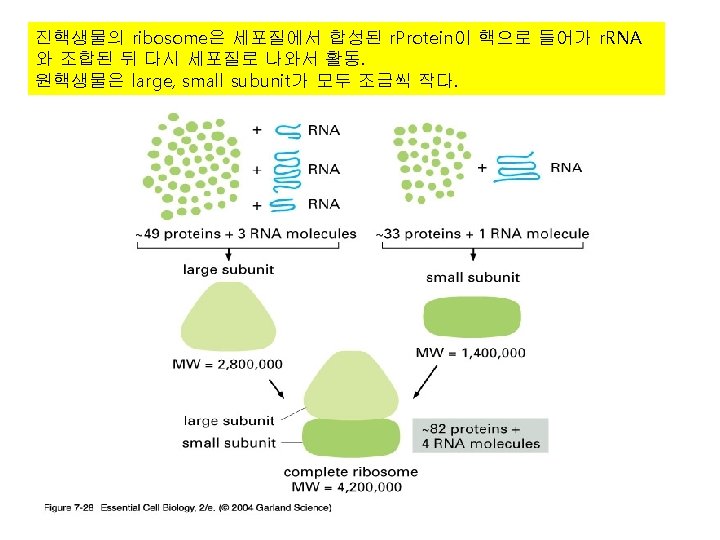
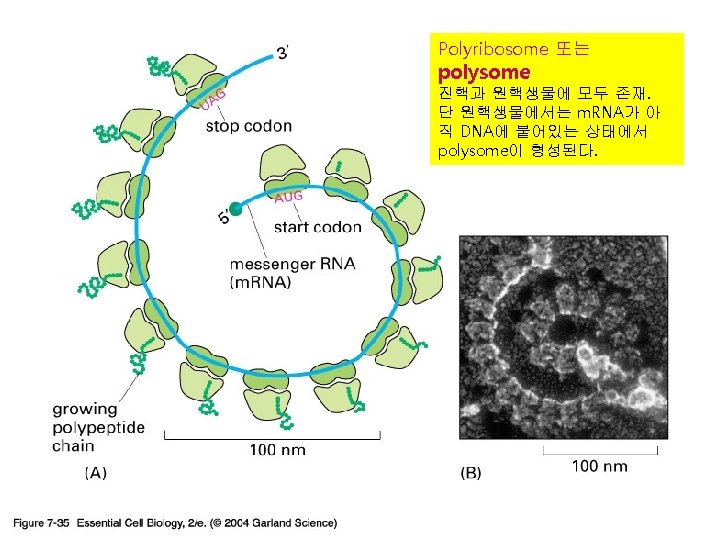
3) r. RNA (ribosome) 실제로 단백질의 조립이 일어나는 공장. Some ribosomes are free. Some are bound to the ER.
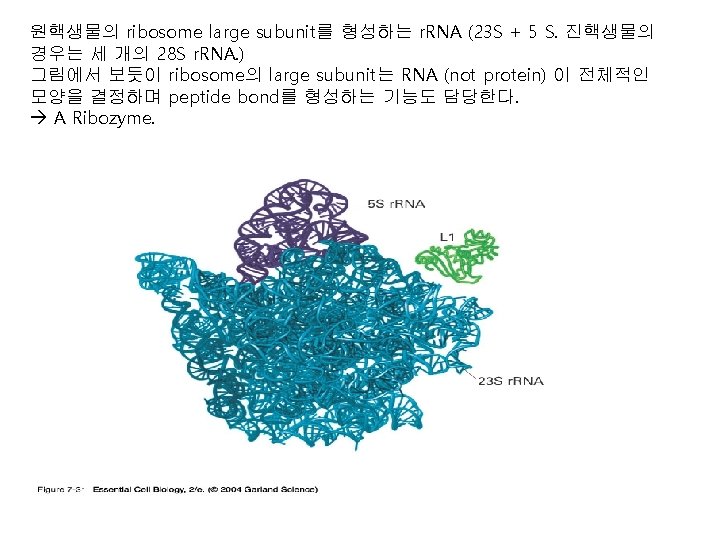
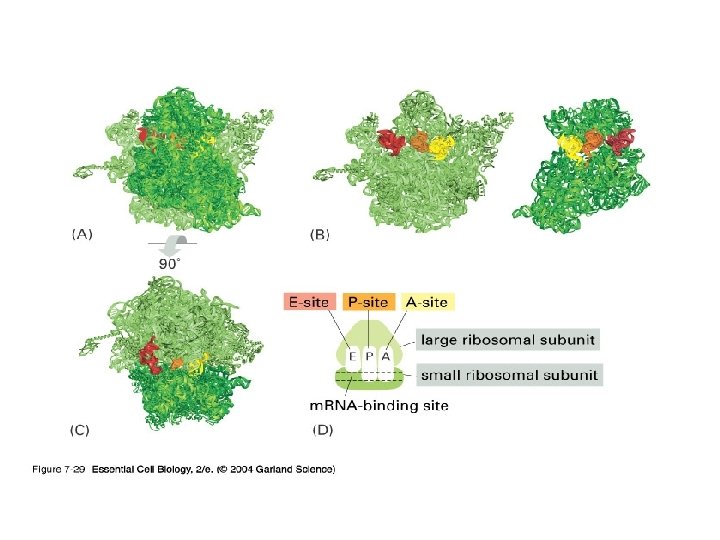
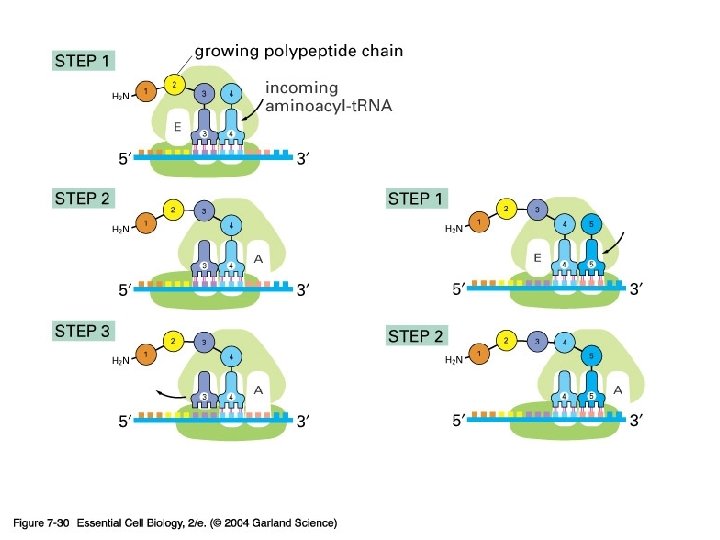
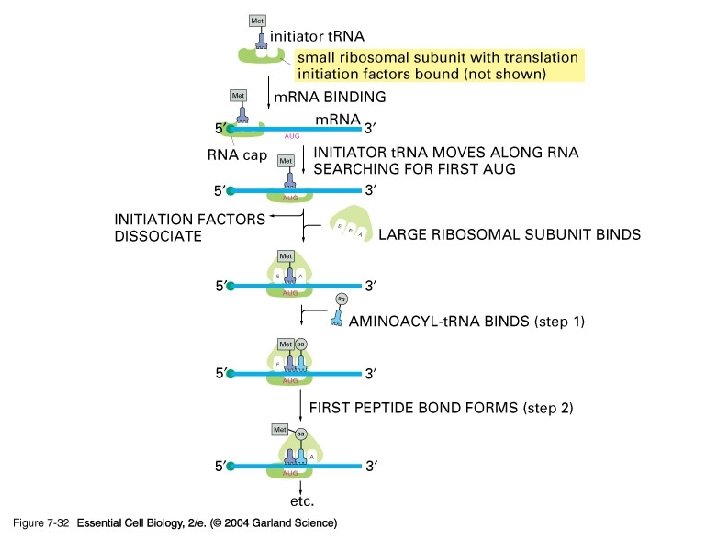
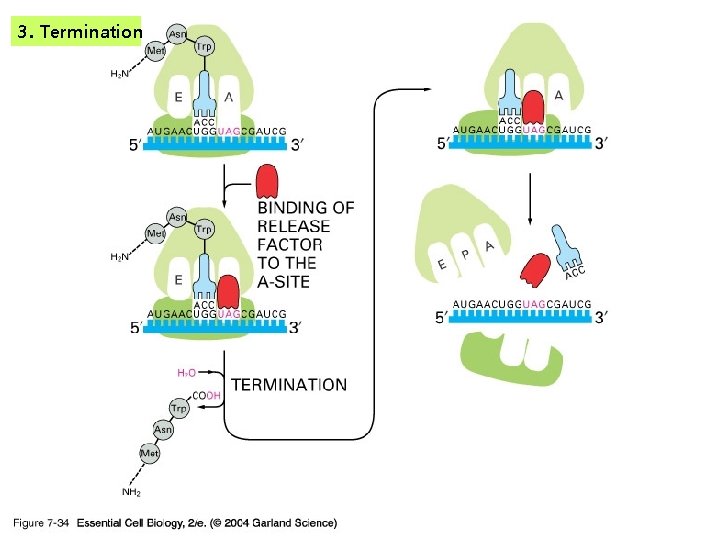
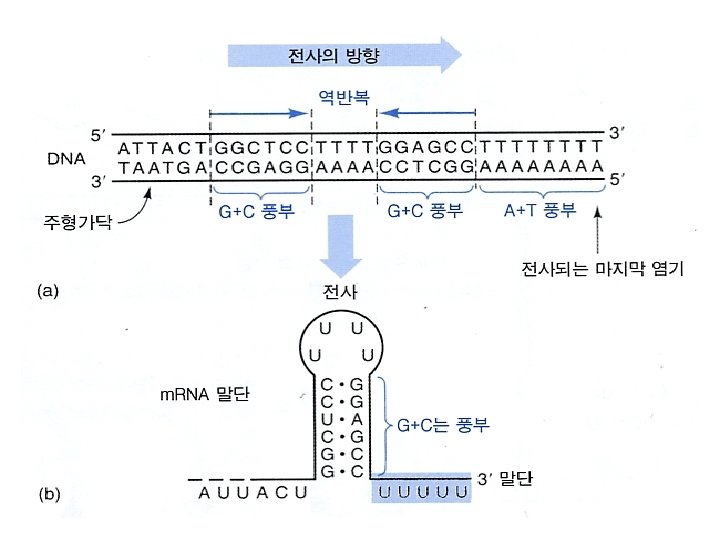
3. Termination


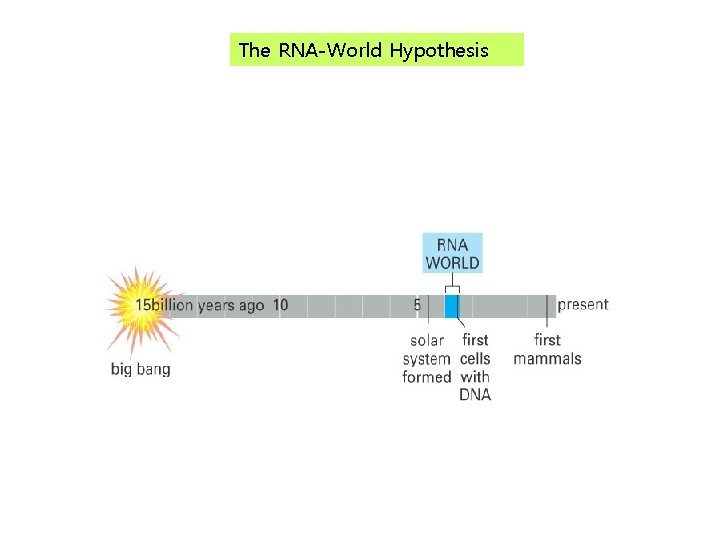
The RNA-World Hypothesis


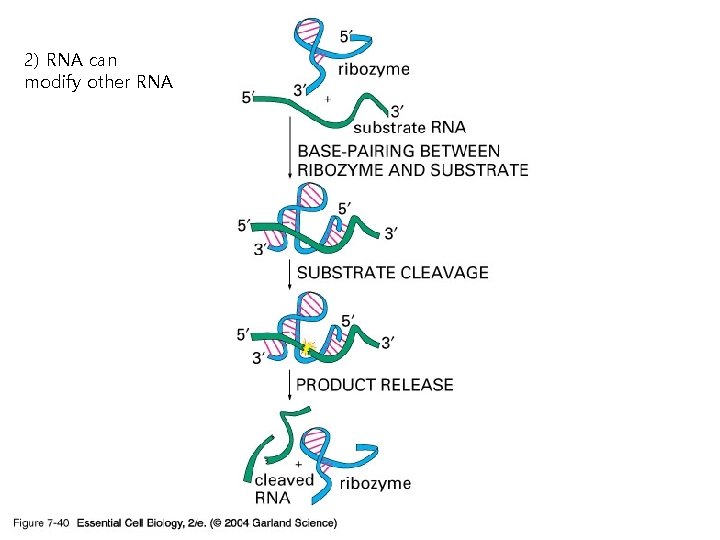
2) RNA can modify other RNA
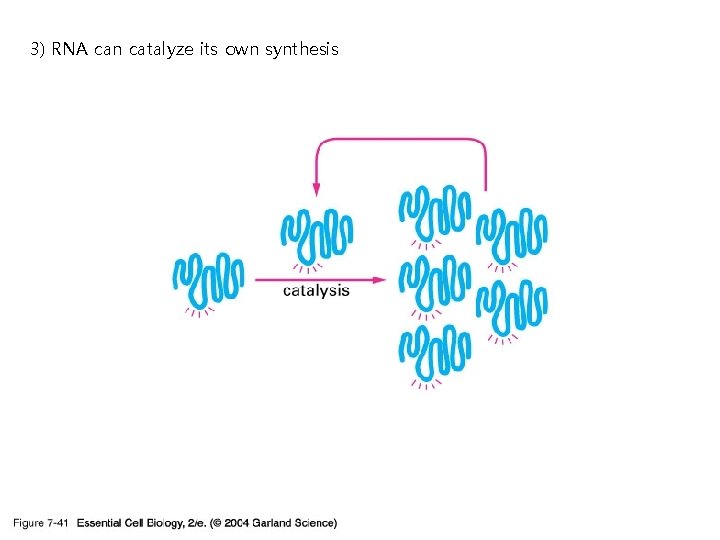
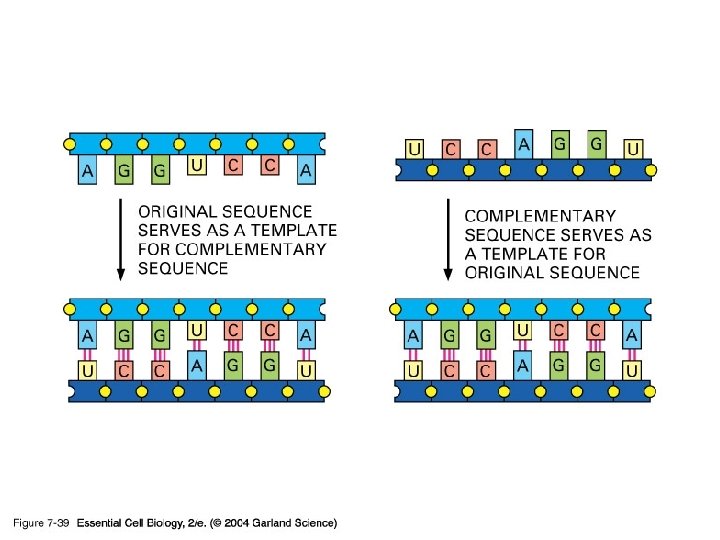
3) RNA can catalyze its own synthesis
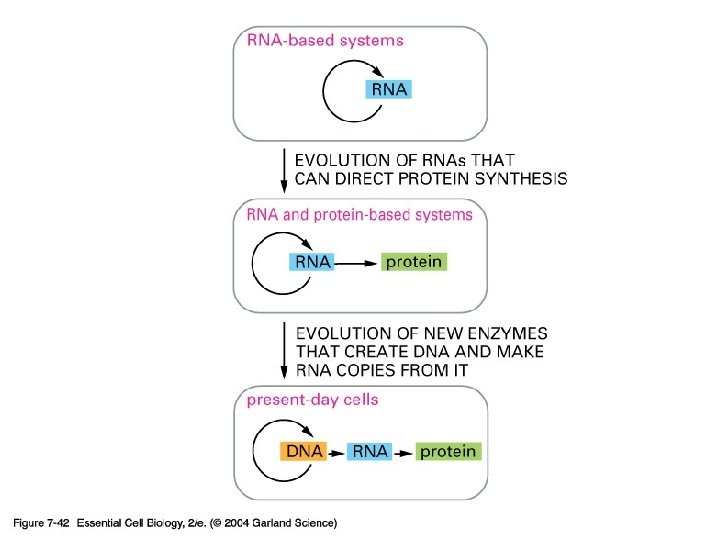
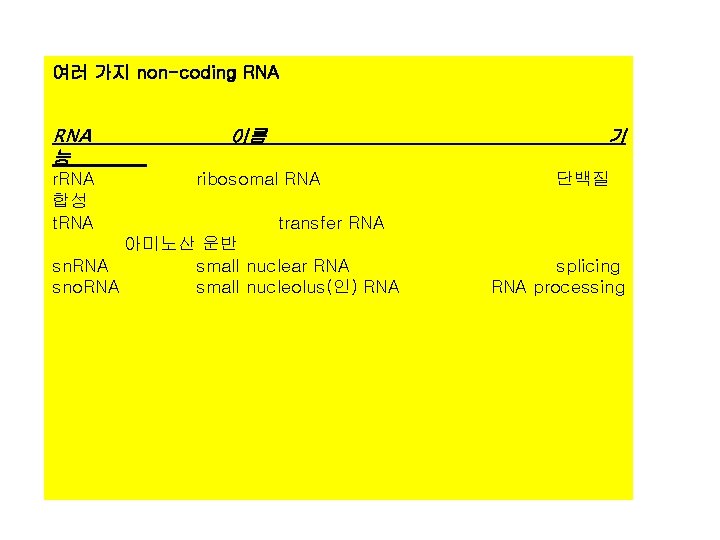
여러 가지 non-coding RNA 능 이름 기 r. RNA ribosomal RNA 단백질 합성 t. RNA transfer RNA 아미노산 운반 sn. RNA small nuclear RNA splicing sno. RNA small nucleolus(인) RNA processing
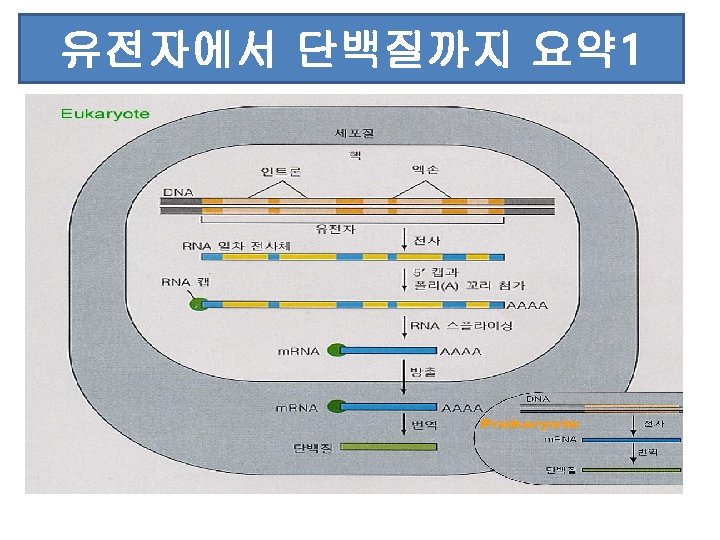
- Slides: 133