Forensic Science to Investigate a Foodborne Outbreak Foodborne

Forensic Science to Investigate a Foodborne Outbreak

Foodborne Outbreak Mini. Lab Outline • Background: What is foodborne illness and how is it investigated using forensic methods? • Part I: Observing Phenomena – Organize and analyze data from a cohort study – Construct a hypothesis about the source of a foodborne outbreak • Part II: Experimenting – Review molecular methods for detecting pathogen in clinical samples and food – Use gel electrophoresis to analyze samples and test your hypothesis • Communicate results and conclusions

What is a Foodborne Illness? • An infection or irritation of the gastrointestinal (GI) tract due to consuming contaminated food or beverage • The Centers for Disease Control and Prevention (CDC) estimates that in the United States • 1 in 6 Americans will come down with a foodborne illness each year • 128, 000 people will be hospitalized • 3, 000 people will die of the infection Source (CDC, 2014) • When two or more people develop a similar illness resulting from the consumption of the same product, this is considered an outbreak

Foodborne Outbreaks That Made the News • Shigella Outbreak at Mexican Restaurant in San Jose, Oct. 2015 • Chipotle E. coli Outbreak, Oct. 2015 • Costco Chicken Salad E. coli Outbreak, Nov. 2015 • Salmonella in imported cucumbers, Nov. 2015 • Dole Salad Listeriosis Outbreak, Jan. 2016

Foodborne illness: defined as foodborne infections or foodborne intoxication • Foodborne infections are caused by live microorganisms commonly transmitted by the fecal-oral route (poor hygienic practices). • Foodborne intoxication is caused by microbial toxins that cause illness. • Different types of microorganisms can cause foodborne illness: fungi (Aspergillus), protozoa (Cyclospora), bacteria (Salmonella, E. coli), viruses (Norovirus)

Investigation of a foodborne outbreak: the science of epidemiology • Epidemiologists use an investigative process to determine the source(s) of foodborne illness. • In this lab you will apply the scientific method to determine the source of an outbreak that affected party attendees. • Outbreaks are like a crime scene, but can be much more complex – – – “The scene” may involve multiple states or countries Often involves multiple “suspects” Evidence is usually difficult to find and collect Time is always in short supply Must understand the complete matrix of food production and distribution

Part I: Observing Phenomena Bad Food at a Good Party

Scientific approach to solving a problem Observation and Purpose – Ask questions, then form an objective based on your observation Background Research – Gain background knowledge about subject Hypothesis – Make an informed and testable prediction Experiment – Test your hypothesis by doing experiments and collecting data Results and Conclusion – Analyze your data and compare to hypothesis – were you right? Communicate – share your hypothesis, experiments, results and conclusions with others, seek feedback

The Case: A multi-state outbreak of foodborne infection has been detected. Which food is contaminated? Epidemiological approach: a cohort study Cohort: A group of people who share a common experience or characteristic.

Cohort study: several people attended a party, some became ill 1. 2. 3. 4. What foods were served? Question partygoers: who became ill? Who ate which foods? Find a common link between who became ill and which foods they consumed 5. Those who were sick had a common pathogen in their stool sample 6. Analyze suspected food for presence of pathogen 7. Connect the dots- find a logical link between clinical sample and contaminated food
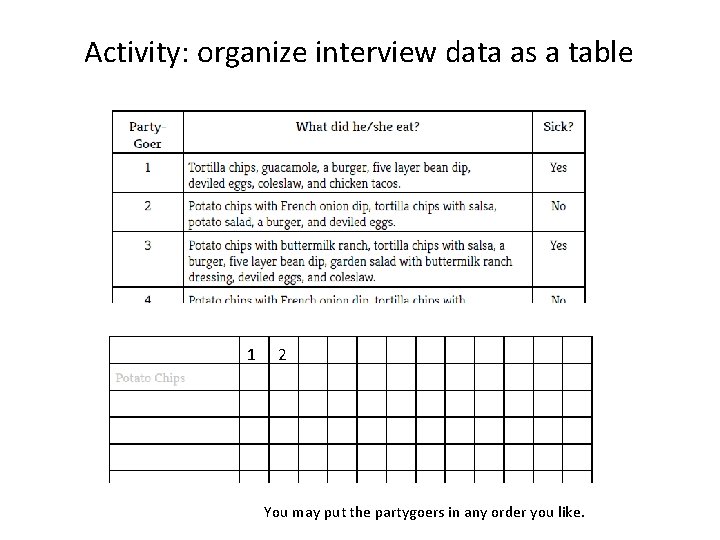
Activity: organize interview data as a table 1 2 You may put the partygoers in any order you like.

Scientific approach to solving a problem Observation and Purpose – Ask a question then form an objective based on your observation Background Research – Gain background knowledge about subject Hypothesis – Make an informed and testable prediction Experiment – Test your hypothesis by doing experiments and collecting data Results and Conclusion – Analyze your data and compare to hypothesis – were you right? Communicate – Share your hypothesis, experiments, results and conclusions with others, seek feedback

Form your hypothesis • Data known- Shigella sonnei, a bacterial pathogen, was isolated from stool samples from ill patients • Which foods were consumed by partygoers that became ill? • Which foods were consumed and no one became ill? • Narrow the list of suspected foods • Hypothesis: which specific food is contaminated with the pathogen?

Part II: Experimenting Tracing the Outbreak to its Source

Public Health Dispatch: Outbreak of Shigella sonnei Infections Associated with Eating a Nationally Distributed Dip -- California, Oregon, and Washington, January 2000 A multistate outbreak of Shigella sonnei infections with at least 30 cultureconfirmed cases in California, Oregon, and Washington has been linked to eating a nationally distributed five-layer dip. Symptom onsets occurred during January 1023, 2000; case-finding is ongoing. The implicated product is manufactured by Señor Felix's Mexican Foods (Baldwin Park, California) and distributed under the brand names Señor Felix's 5 -Layer Party Dip (sold in 16 -ounce, 20 -ounce, and 41 ounce containers), Delicioso 5 -Layer Party Dip (33 -ounce containers), and Trader Joe's 5 -Layer Party Dip (20 -ounce containers). The dip consists of layers of bean, salsa, guacamole, nacho cheese, and sour cream. (Source: http: //www. cdc. gov/mmwr/preview/mmwrhtml/mm 4903 a 4. htm)

Scientific approach to solving a problem Observation and Purpose – Ask a question then form an objective based on your observation Background Research – Gain background knowledge about subject Hypothesis – Make an informed and testable prediction Experiment – Test your hypothesis by doing experiments and collecting data Results and Conclusion – Analyze your data and compare to hypothesis – were you right? Communicate – Share your hypothesis, experiments, results and conclusions with others, seek feedback

Molecular methods for detecting pathogenic organisms in food • How do we collect and analyze food samples? • What are the best methods for detecting the pathogenic microbe? • What controls must be used to ensure correct results?

Detecting Shigella sonnei in foods • Shigella sonnei has LOW infectious dose of 10 -100 cells • Standard bacteriological culture may not grow organisms found in low numbers in food • Polymerase Chain Reaction (PCR) is simple, rapid, specific, and more sensitive than culture methods • PCR is an in vitro (test tube) method that amplifies (makes many copies of) specific target sequences in nucleic acids • Powerful tool to detect and identify organisms, from microbes to humans

Detecting Shigella sonnei in foods • Shigella-specific genes are amplified by PCR – 175 bp fragment from ipa. H gene – 1000 bp fragment from mxi. C gene • A Positive Control is used to determine true negative versus a PCR reaction failure due to inhibitory substances in the food. – Genetically modified lab strain Shigella mixed into all food samples – PCR fragments will be 175 bp and 1800 bp (not 1000 bp) • Stop and think: is a Negative Control necessary in PCR reactions?

Experimental procedure • Investigators first extracted DNA for PCR from a homogenized sample of all five layers • Initial PCR results were negative for Shigella • Investigators followed up by sampling each layer individually and repeating PCR • See “Bean Dip: General Production Information” to learn how each component was prepared • You will use agarose gel electrophoresis to analyze the PCR products of each reaction and determine the source of the pathogen
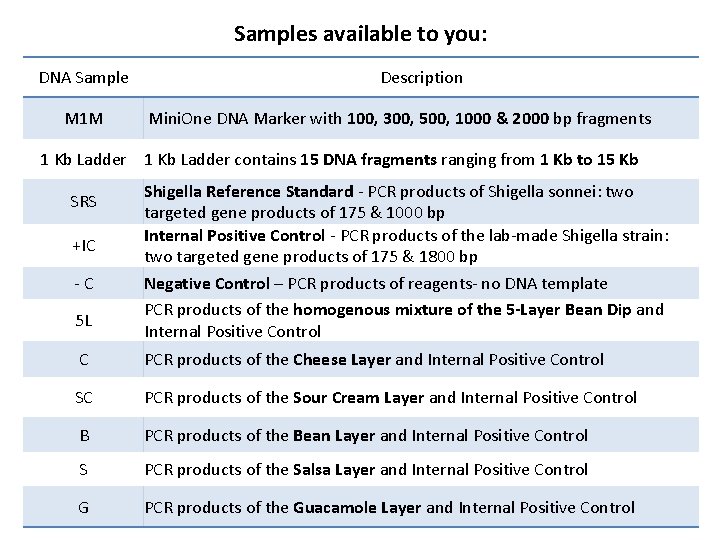
Samples available to you: DNA Sample M 1 M Description Mini. One DNA Marker with 100, 300, 500, 1000 & 2000 bp fragments 1 Kb Ladder contains 15 DNA fragments ranging from 1 Kb to 15 Kb SRS +IC - C 5 L Shigella Reference Standard - PCR products of Shigella sonnei: two targeted gene products of 175 & 1000 bp Internal Positive Control - PCR products of the lab-made Shigella strain: two targeted gene products of 175 & 1800 bp Negative Control – PCR products of reagents- no DNA template PCR products of the homogenous mixture of the 5 -Layer Bean Dip and Internal Positive Control C PCR products of the Cheese Layer and Internal Positive Control SC PCR products of the Sour Cream Layer and Internal Positive Control B PCR products of the Bean Layer and Internal Positive Control S PCR products of the Salsa Layer and Internal Positive Control G PCR products of the Guacamole Layer and Internal Positive Control
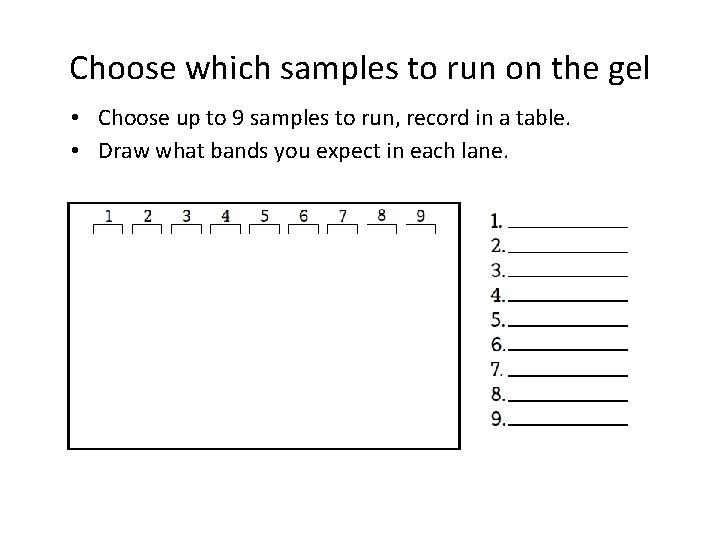
Choose which samples to run on the gel • Choose up to 9 samples to run, record in a table. • Draw what bands you expect in each lane.
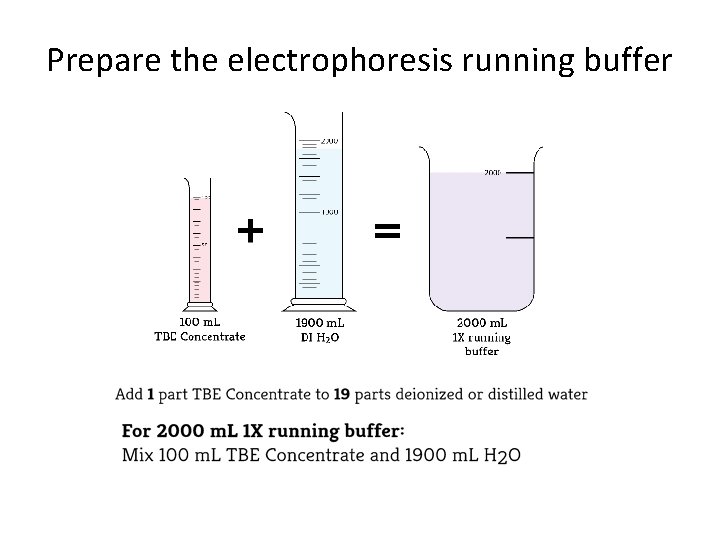
Prepare the electrophoresis running buffer

Cast the agarose gel

Cast the agarose gel

Prepare the Mini. One electrophoresis chamber

Prepare the Mini. One electrophoresis chamber

Load your samples

Run your gel

Document your results Include a labeled image of your gel in your lab report.

Analyze and Report • Which food item was the source of the pathogen? • Did the controls work properly to verify all data? • Were the dots connected between clinical and food samples? • Knowing what you do now, would you design the experiment differently? Explain how and why.

Conclusion In this lab you have: • Used a cohort study to determine which food sickened partygoers • Used sensitive molecular methods to identify the source of the contamination

Background Information on the PCR reaction

A PCR reaction requires the following components: • Template or target DNA, often genomic DNA isolated from an organism. (In the case of the Foodborne Outbreak Lab, the template is the bacterial DNA extracted from the contaminated bean dip. ) • Primers that are specific for the gene that is being amplified • d. NTPs, the building blocks for making DNA • DNA polymerase (heat-stable enzyme)-copies DNA • PCR buffer, a salt solution that maintains an ideal p. H • Mg 2+, a co-factor for the DNA polymerase

PCR Steps 1. Denaturation (separate DNA strands) temperature = 94 -95°C strand separation 2. Annealing of primers temperature variable < 72°C primers hybridize to template 3. Extension of template strands temperature = 72°C need heat stable polymerase, d. NTPs
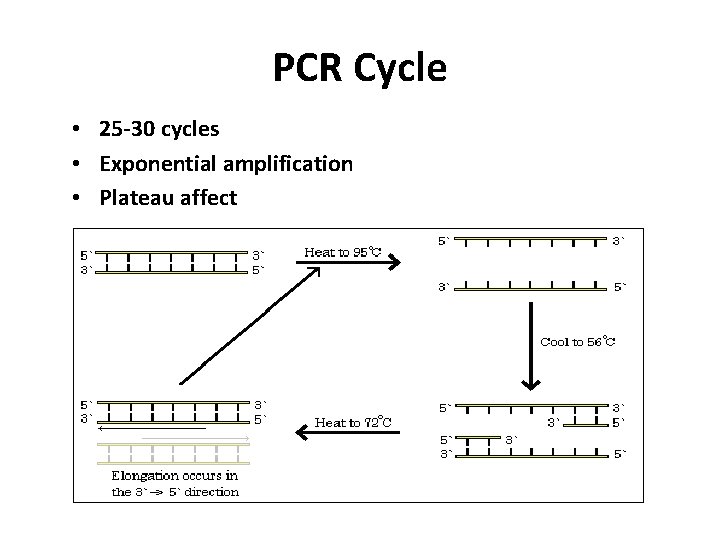
PCR Cycle • 25 -30 cycles • Exponential amplification • Plateau affect
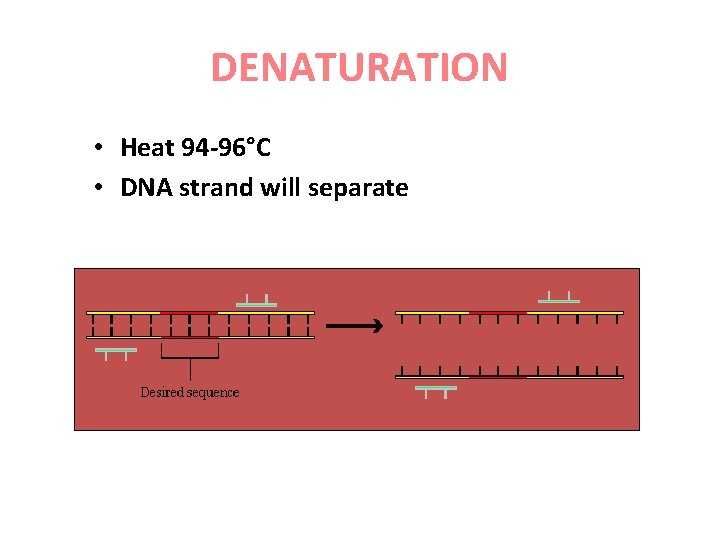
DENATURATION • Heat 94 -96°C • DNA strand will separate
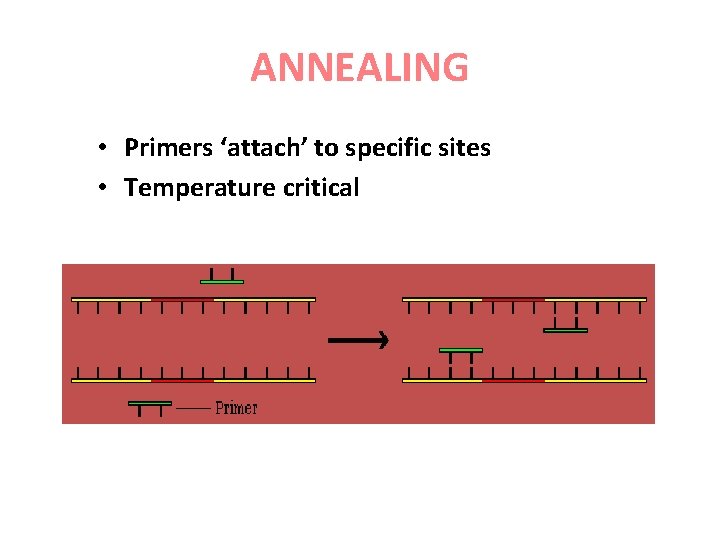
ANNEALING • Primers ‘attach’ to specific sites • Temperature critical
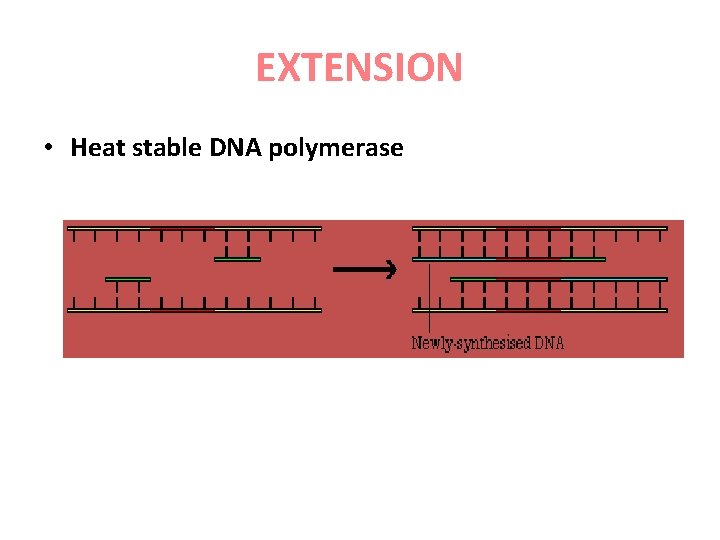
EXTENSION • Heat stable DNA polymerase
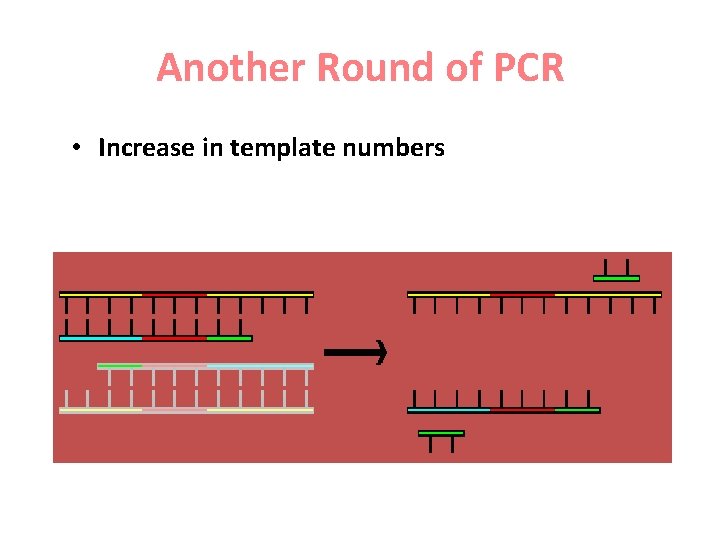
Another Round of PCR • Increase in template numbers

Successful PCR 1. REACTION CONDITIONS - Primer specificity - Mg 2+ concentration - Temperature - No carryover - Sensitivity 2. QUALITY OF TEMPLATE - No inhibitors - Intact vs sheared; degraded

Exponential Amplification • 1 copy of target gene after 30 cycles 230 ≈ 1 billion* • 5 copies of target gene after 30 cycles 5*230 ≈ 5 billion* • 10 copies of target gene after 30 cycles 10*230 ≈ 10 billion* *Due to the limitations of reagents, the amount of PCR products may not equal the calculated value.
- Slides: 42