Folding Experiments 2 UV absorbance of aromatic amino
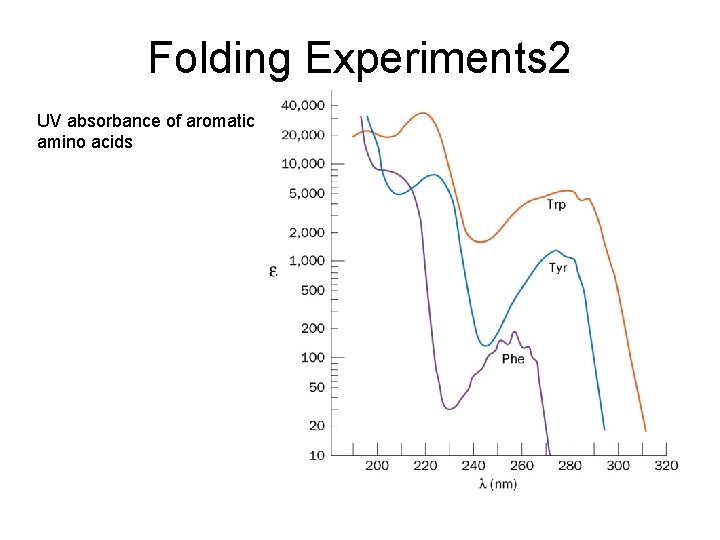
Folding Experiments 2 UV absorbance of aromatic amino acids
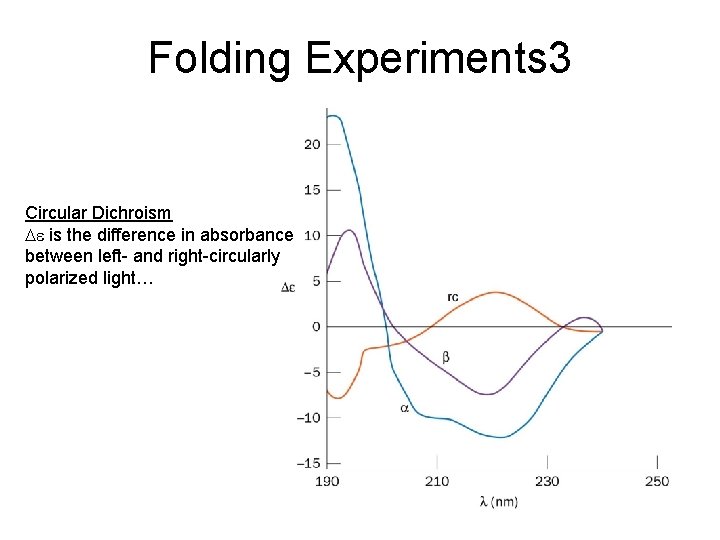
Folding Experiments 3 Circular Dichroism De is the difference in absorbance between left- and right-circularly polarized light…
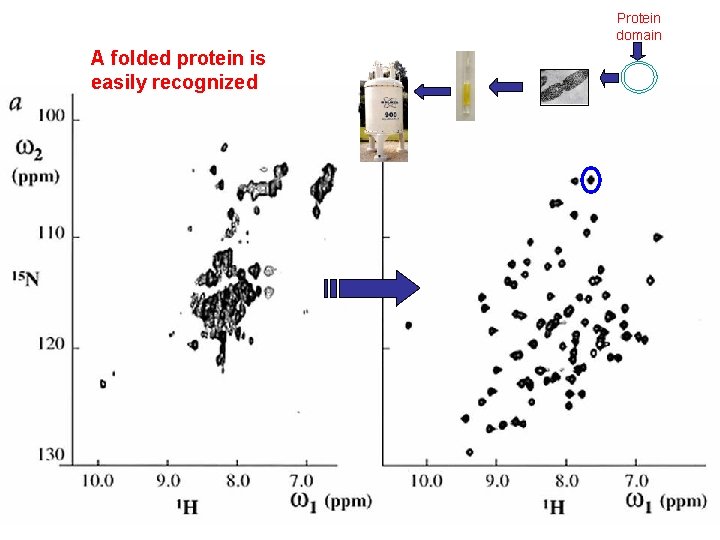
Protein domain A folded protein is easily recognized
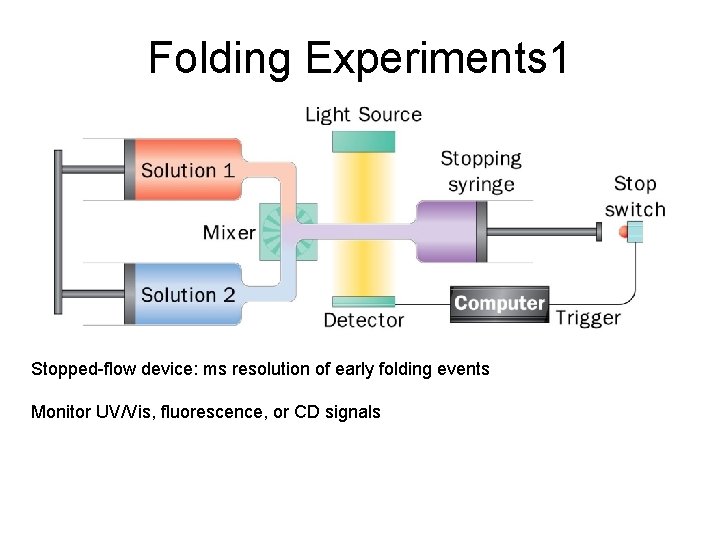
Folding Experiments 1 Stopped-flow device: ms resolution of early folding events Monitor UV/Vis, fluorescence, or CD signals

Folding Accessory Proteins 7 GRASP image of PDI (electrostatic surface potential: red=O- and blue=N+)
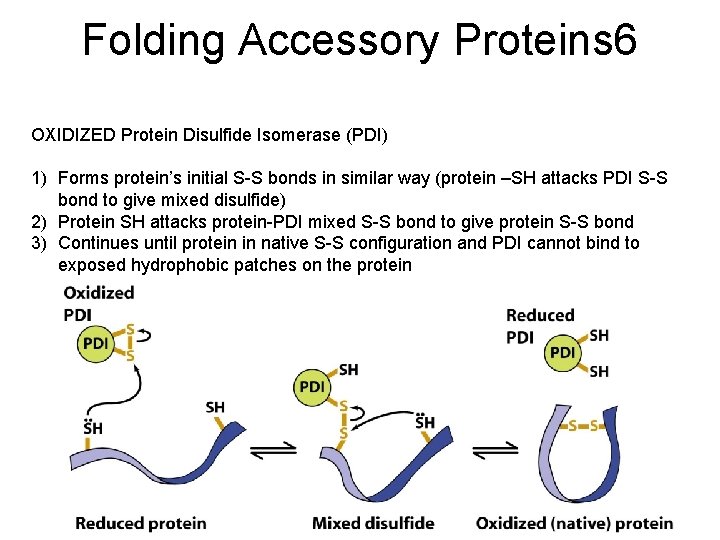
Folding Accessory Proteins 6 OXIDIZED Protein Disulfide Isomerase (PDI) 1) Forms protein’s initial S-S bonds in similar way (protein –SH attacks PDI S-S bond to give mixed disulfide) 2) Protein SH attacks protein-PDI mixed S-S bond to give protein S-S bond 3) Continues until protein in native S-S configuration and PDI cannot bind to exposed hydrophobic patches on the protein
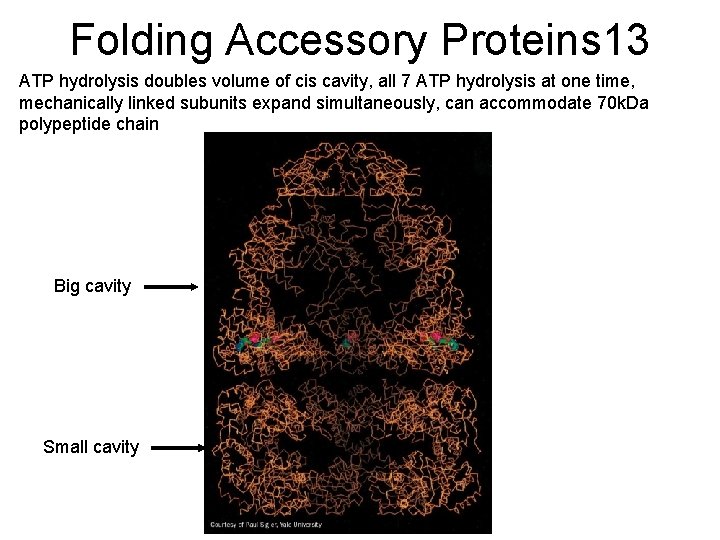
Folding Accessory Proteins 13 ATP hydrolysis doubles volume of cis cavity, all 7 ATP hydrolysis at one time, mechanically linked subunits expand simultaneously, can accommodate 70 k. Da polypeptide chain Big cavity Small cavity
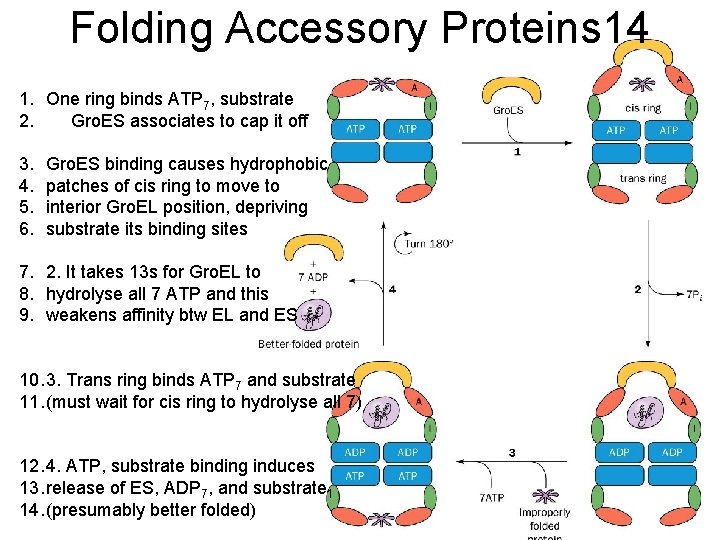
Folding Accessory Proteins 14 1. One ring binds ATP 7, substrate 2. Gro. ES associates to cap it off 3. 4. 5. 6. Gro. ES binding causes hydrophobic patches of cis ring to move to interior Gro. EL position, depriving substrate its binding sites 7. 2. It takes 13 s for Gro. EL to 8. hydrolyse all 7 ATP and this 9. weakens affinity btw EL and ES 10. 3. Trans ring binds ATP 7 and substrate 11. (must wait for cis ring to hydrolyse all 7) 12. 4. ATP, substrate binding induces 13. release of ES, ADP 7, and substrate 1 14. (presumably better folded)
- Slides: 8