Fold Recognition Ole Lund Assistant professor CBS Fold

Fold Recognition Ole Lund, Assistant professor, CBS

Fold recognition l Find template for modeling – l 1 st step in comparative modeling Can be used to predict function OL

Template identification l Search with sequence – – – l l l Blast against proteins with known structure Psi-Blast against all proteins Fold recognition methods Use biological information Functional annotation in databases Active site/motifs OL

Blast derivatives: PDB-BLAST l Procedure 1. 2. l Build sequence profile by iterative PSI-BLAST search against a sequence database Use profile to search database of proteins with known structure Advantage – Makes sure hid to protein with known structure is not hidden behind a lot of hits to other proteins OL

BLAST derivatives: Transitive BLAST l Procedure 1. 2. 3. – l Find homologues to query (your) sequence Find homologues to these homologues Etc. Can be implemented with e. g. BLAST or PSIBLAST Also known as Intermediate Sequence Search (ISS) OL

CASP l CASP – – – Critical Assessment of Structure Predictions Every second year Sequences from about-to-be-solved-structures are given to groups who submit their predictions before the structure is published Modelers make prediction Meeting in Asilomar where correct answers are revealed OL

Target difficulty l l l CM: Comparative (homology) modeling CM/FR: not PSI-BLAST (but ISS) findable FR(H): Homologous fold recognition FR(A): Analogous fold recognition NF/FR: Partly New fold NF: New Fold (used to be called Ab Initio from first principles- prediction) OL
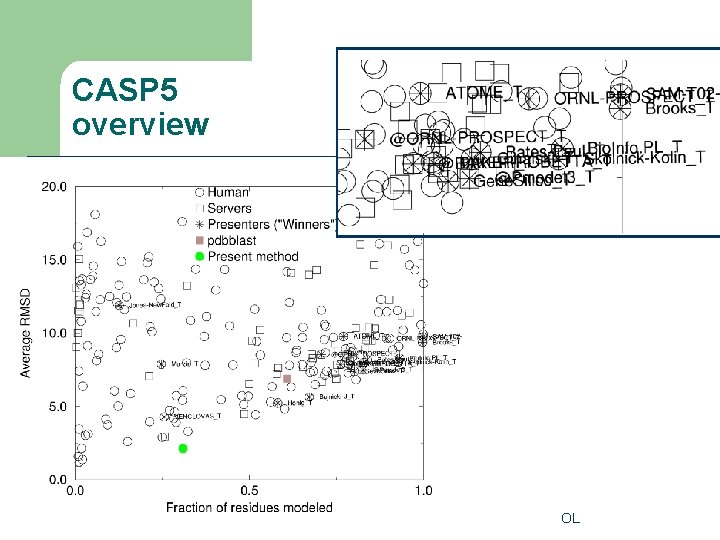
CASP 5 overview OL

Successful fold recognition groups at CASP 5 l l l 3 D-Jury (Leszek Rychlewski) 3 D-CAM (Krzysztof Ginalski) Template recombination (Paul Bates) HMAP (Barry Honig) PROSPECT (Ying Xu) ATOME (Gilles Labesse) OL

3 D-Jury (Rychlewski) l Inspired by Ab initio modeling methods – l Find most abundant high scoring models 1. 2. 3. 4. l Average of frequently obtained low energy structures is often closer to the native structure than the lowest energy structure Use output from a set of servers Superimpose all pairs of structures Similarity score Sij = # of Ca pairs within 3. 5Å (if #>40; else Sij=0) 3 D-Jury score = Si. Sij/(N+1) Similar methods developed by A Elofsson (Pcons) and D Fischer (3 D shotgun) Rychlewski. doc OL

Ginalski. doc 3 D-CAM (Krzysztof Ginalski) l 3 D-Consensus Alignment Method – – Structural alignment for all members of fold from FSSP Conservation of specific residues and contacts l l – – – – responsible for maintaining tertiary structure critical for substrate binding and/or catalysis Find homologues with iterative PSI-BLAST Align with Clustal. W – identify conserved residues Structural integrity of alignments Manual realignment Fold recognition for homologues Modelling Verification l l Visually Computationally (Verify 3 D, Prosa. II, WHAT_CHECK) OL

Abstract Paul A Bates - In Silico Recombination of Templates, Alignments and Models l Problems – – l Models rarely better than templates Manual intervention have marginal effect Possible solution – Recombination of models OL

Abstract Paul A Bates – Modelling Procedure l l Define domains Make models (FAMS/Pmodeller/Esy. Pred 3 D) – – – l Select pair of models – – l Superimpose Crossover or mutate (average coordinates) Select best proportion – – l Manual inspection/correction of alignments Alignment of annotated residues (PFAM) Preferably use alignment with >2 bits/aa Contact pair potentials Solvation energies (calculated from solvent accessible area) Convergence – Minimization and final refinements OL

Abstract Barry Honig l Sequence&structure profile-profile based alignment – Database of template profiles l l – Multiple structure alignment Sequence based profiles Position specific gap penalties derived from secondary structure Calibration to estimate statistical significance Query profile l l Sequence based profile Predicted secondary structure (consensus between PSIPRED, PHD, JNET) OL

Abstract Ying Xu l PROSPECT: optimal alignments for a given energy function with any combination of the following terms: mutation energy (including position-specific score matrix derived from multiple-sequence alignments), 2. singleton energy (including matching scores to the predicted secondary structures), 3. pairwise contact potential 4. alignment gap penalties. 1. OL

Abstract Gilles Labesse l Meta Server – l l 3 D-PSSM, PDB-BLAST, FUGUE, Gen. THREADER, SAM-T 99, JPRED-2 Tool for Incremental Threading optimization (T. I. T. O. ) Consensus ranking OL

Live. Bench l The Live Bench Project is a continuous benchmarking program. Every week sequences of newly released PDB proteins are being submitted to participating fold recognition servers. The results are collected and continuous evaluated using automated model assessment programs. A summary of the results is produced after several months of data collection. The servers must delay the updating of their structural template libraries by one week to participate. OL
Meta Server OL
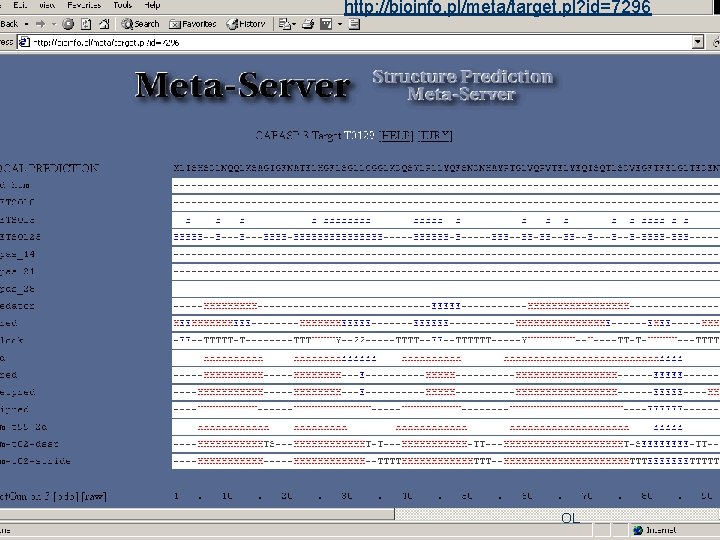
http: //bioinfo. pl/meta/target. pl? id=7296 Meta Server OL
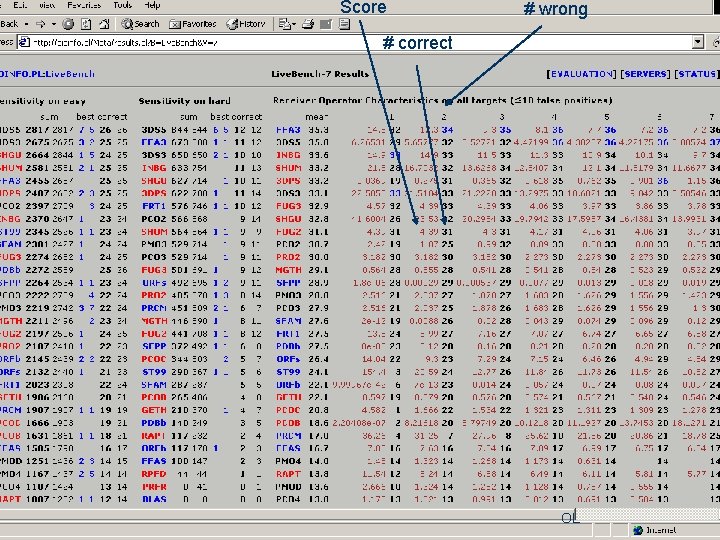
Score # wrong # correct OL

Best servers? l l l l FFA 3 3 DS 5 INBG SHUM 3 DPS 3 DS 3 FUG 3 SHGU FUG 2 PCO 2 PRO 2 MGTH SFPP PMO 3 OL

Links to fold recognition servers l Databases of links – – l Meta server – l http: //www. cse. ucsc. edu/research/compbio/HMM-apps/T 99 -query. html FOLD – l http: //www-cryst. bioc. cam. ac. uk/~fugue/prfsearch. html SAM – l http: //bioinf. cs. ucl. ac. uk/psipred/ FUGUE 2 – l http: //www. sbg. bio. ic. ac. uk/servers/3 dpssm/ Gen. THREADER – l http: //bioinfo. pl/meta/ (Example: http: //bioinfo. pl/meta/target. pl? id=7296 ) 3 DPSSM – good graphical output – l http: //bioinfo. pl/meta/servers. html http: //mmtsb. scripps. edu/cgi-bin/renderrelres? protmodel http: //fold. doe-mbi. ucla. edu/ FFAS/PDBBLAST – http: //bioinformatics. burnham-inst. org/ OL
- Slides: 22