Fluorescence in situ hybridization FISH an increasingly demanded

Fluorescence in situ hybridization (FISH): an increasingly demanded tool for biomarker research and personalized medicine Weimin Miao 1. Institute of Hematology and Blood Diseases Hospital, CAMS & PUMC 2. Tianjin key Laboratory of Blood Cell Therapy and Technology

Background Potential biomarkers for cancers and hematological diseases Recurrent genomic defects a. b. c. d. e. amplification, insertion deletion, recombination Mutation

personalized medicine (Precision medicine) Precision Diagnosis Precision biomarker detection

Detection methods for molecular biomarkers: 1. Cutting-edge high through-put a. Comparative Genomic Hybridization (CGH) array b. Single Nucleotide Polymorphism (SNP) array c. Next Generation Sequencing (NGS) 2. Traditional low through-put a. PCR b. Sanger sequencing c. Fluorescence In situ Hybridization (FISH)

Fluorescence in situ hybridization (FISH) FISH uses fluorescent DNA probes to target a specific chromosome location within a nucleus, resulting in coloured signals that can be detected by fluorescent microscope. 5 5

Topic 1: Fluorescence in situ hybridization (FISH) is increasingly demanded for biomarker detection and personalized medicine. Advantages: 1. Specific and sensitive 2. Simple and reliable 3. Results are visible Disavantages: 1. Low through-put
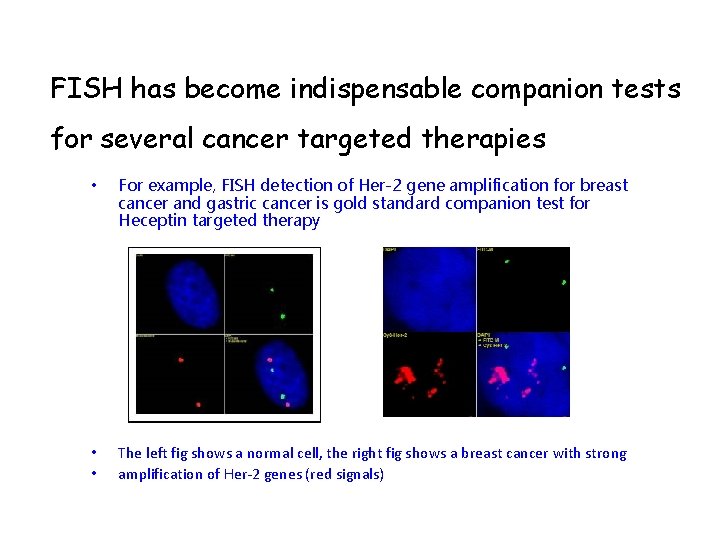
FISH has become indispensable companion tests for several cancer targeted therapies • For example, FISH detection of Her-2 gene amplification for breast cancer and gastric cancer is gold standard companion test for Heceptin targeted therapy • • The left fig shows a normal cell, the right fig shows a breast cancer with strong amplification of Her-2 genes (red signals)
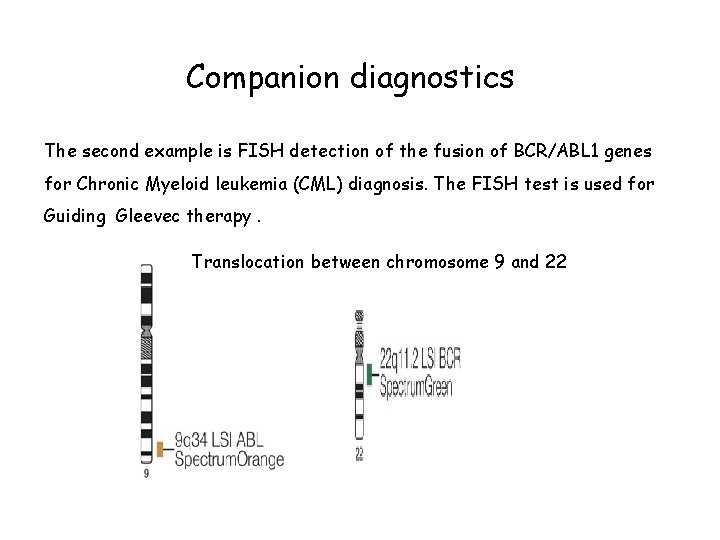
Companion diagnostics The second example is FISH detection of the fusion of BCR/ABL 1 genes for Chronic Myeloid leukemia (CML) diagnosis. The FISH test is used for Guiding Gleevec therapy. Translocation between chromosome 9 and 22
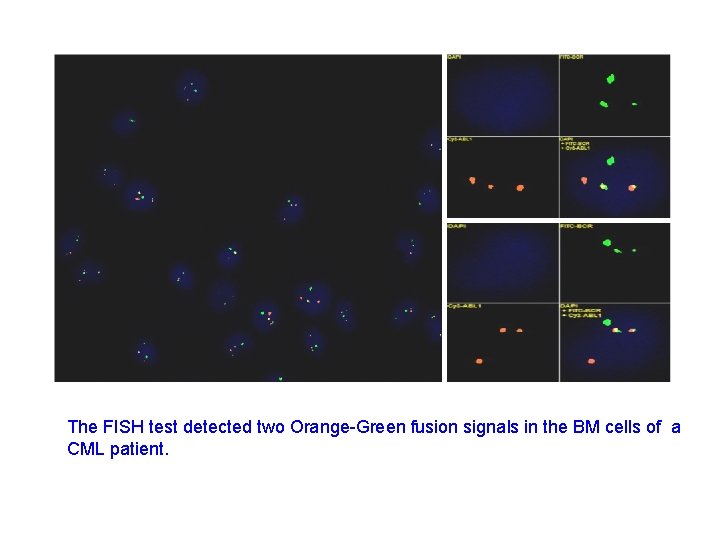
The FISH test detected two Orange-Green fusion signals in the BM cells of a CML patient.
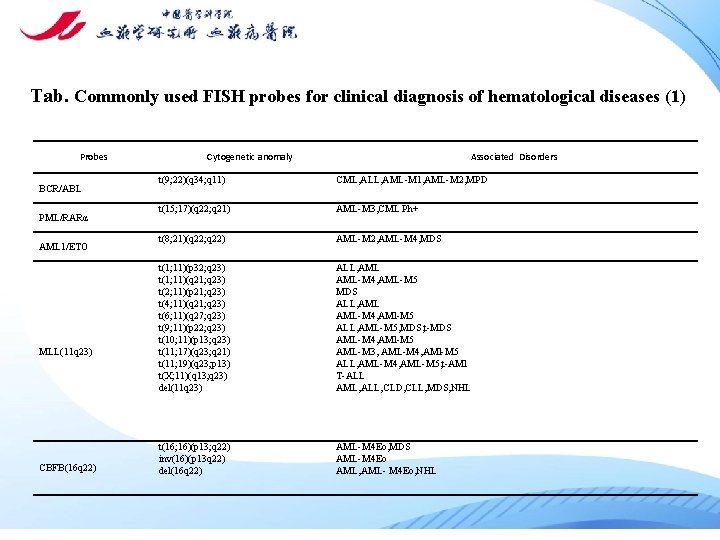
Tab. Commonly used FISH probes for clinical diagnosis of hematological diseases (1) Probes BCR/ABL PML/RARα AML 1/ETO MLL(11 q 23) CBFB(16 q 22) Cytogenetic anomaly Associated Disorders t(9; 22)(q 34; q 11) CML, ALL, AML-M 1, AML-M 2, MPD t(15; 17)(q 22; q 21) AML-M 3, CML Ph+ t(8; 21)(q 22; q 22) AML-M 2, AML-M 4, MDS t(1; 11)(p 32; q 23) t(1; 11)(q 21; q 23) t(2; 11)(p 21; q 23) t(4; 11)(q 21; q 23) t(6; 11)(q 27; q 23) t(9; 11)(p 22; q 23) t(10; 11)(p 13; q 23) t(11; 17)(q 23; q 21) t(11; 19)(q 23; p 13) t(X; 11)(q 13; q 23) del(11 q 23) ALL, AML-M 4, AML-M 5 MDS ALL, AML-M 4, AMl-M 5 ALL, AML-M 5, MDS, t-MDS AML-M 4, AMl-M 5 AML-M 3, AML-M 4, AMl-M 5 ALL, AML-M 4, AML-M 5, t-AMl T-ALL AML, ALL, CLD, CLL, MDS, NHL t(16; 16)(p 13; q 22) inv(16)(p 13 q 22) del(16 q 22) AML-M 4 Eo, MDS AML-M 4 Eo AML, AML- M 4 Eo, NHL
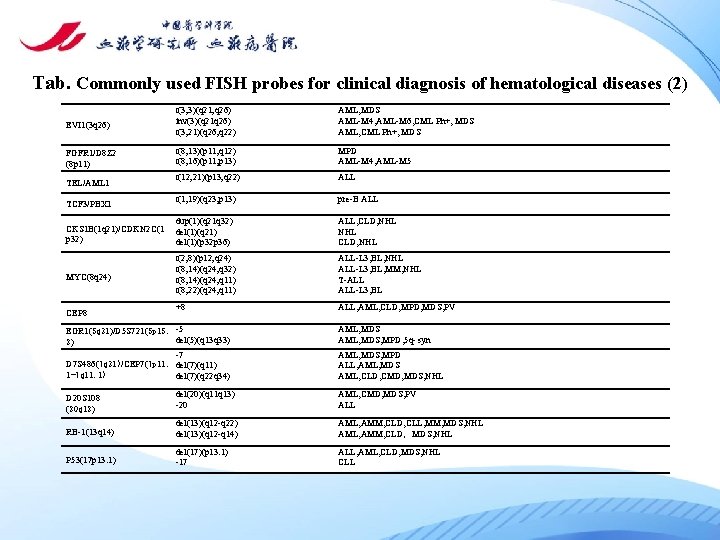
Tab. Commonly used FISH probes for clinical diagnosis of hematological diseases (2) EVI 1(3 q 26) t(3; 3)(q 21; q 26) inv(3)(q 21 q 26) t(3; 21)(q 26; q 22) AML, MDS AML-M 4, AML-M 6, CML Ph+, MDS AML, CML Ph+, MDS FGFR 1/D 8 Z 2 (8 p 11) t(8; 13)(p 11; q 12) t(8; 16)(p 11; p 13) MPD AML-M 4, AML-M 5 t(12; 21)(p 13; q 22) ALL t(1; 19)(q 23; p 13) pre-B ALL CKS 1 B(1 q 21)/CDKN 2 C(1 p 32) dup(1)(q 21 q 32) del(1)(q 21) del(1)(p 32 p 36) ALL, CLD, NHL MYC(8 q 24) t(2; 8)(p 12; q 24) t(8; 14)(q 24; q 32) t(8; 14)(q 24; q 11) t(8; 22)(q 24; q 11) ALL-L 3, BL, NHL ALL-L 3, BL, MM, NHL T-ALL ALL-L 3, BL +8 ALL, AML, CLD, MPD, MDS, PV TEL/AML 1 TCF 3/PBX 1 CEP 8 EGR 1(5 q 31)/D 5 S 721(5 p 15. -5 del(5)(q 13 q 33) 2) -7 D 7 S 486(7 q 31)/CEP 7(7 p 11. del(7)(q 11) 1 -7 q 11. 1) del(7)(q 22 q 34) AML, MDS, MPD, 5 q- syn AML, MDS, MPD ALL, AML, MDS AML, CLD, CMD, MDS, NHL D 20 S 108 (20 q 12) del(20)(q 11 q 13) -20 AML, CMD, MDS, PV ALL RB-1(13 q 14) del(13)(q 12 -q 22) del(13)(q 12 -q 14) AML, AMM, CLD, CLL, MM, MDS, NHL AML, AMM, CLD,MDS, NHL P 53(17 p 13. 1) del(17)(p 13. 1) -17 ALL, AML, CLD, MDS, NHL CLL

Tab. Commonly used FISH probes for clinical diagnosis of hematological diseases (3) ATM(11 q 22. 3) del(14 q 32) t(2; 14)(p 13; q 32) t(3; 14)(q 27; q 32) t(4; 14)(p 13; q 32) t(5; 14)(q 31; q 32) t(6; 14)(p 25; q 32) t(8; 14)(q 24; q 32) t(9; 14)(p 12 -13; q 32) t(11; 14)(q 13; q 32) t(14; 16)( q 32; q 23) t(14; 18)( q 32; q 21) t(14; 19)( q 32; q 13) t(14; 22)( q 32; q 11) del(11)(q 13 q 14 -q 23) CLD, NHL B-CLL DLCL, FL MM ALL B-NHL, LPL B-PLL, CLD, MM, MCL, MGUS, NHL MM CLD, FL, DLCL, MM, NHL CLD, CLL, NHL ALL AML, CLD, CLL, MDS, NHL FGFR 3/IGH t(4; 14)( (p 13; q 32)) MM MAF/IGH t(14; 16)( q 32; q 23) MM CCND 1/IGH t(11; 14) (q 13; q 32) B-PLL, CLD, MM, MCL, MGUS, NHL CEP 12 +12 AML, CLD, NHL t(3; 14) (q 27; q 32) t(3; 22) (q 27; q 11) t(11; 18) (q 21; q 21) t(14; 18) (q 32; q 21) del(18)(q 21) -18 +18 XY/XX -Y DLCL, FL MZL, NHL CLD, FL, DLCL, MM, NHL AML, CLD, NHL CLD ALL, CLD ALLO-SCT ALL, AML, CLD, MDS, MM, MPD, NHL, PV IGH(14 q 32) BCL 6(3 q 27) BCL 2(18 q 21) CEPX/Y
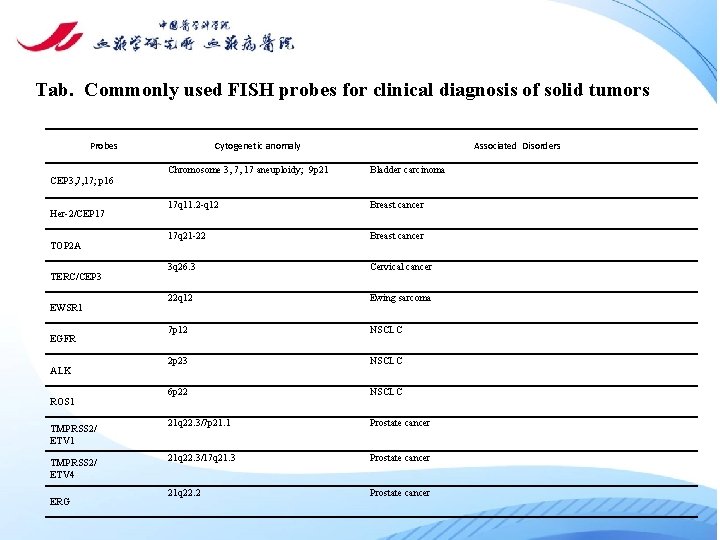
Tab. Commonly used FISH probes for clinical diagnosis of solid tumors Probes Cytogenetic anomaly Associated Disorders Chromosome 3, 7, 17 aneuploidy; 9 p 21 Bladder carcinoma 17 q 11. 2 -q 12 Breast cancer 17 q 21 -22 Breast cancer 3 q 26. 3 Cervical cancer 22 q 12 Ewing sarcoma 7 p 12 NSCLC 2 p 23 NSCLC 6 p 22 NSCLC 21 q 22. 3/7 p 21. 1 Prostate cancer 21 q 22. 3/17 q 21. 3 Prostate cancer 21 q 22. 2 Prostate cancer CEP 3, 7, 17; p 16 Her-2/CEP 17 TOP 2 A TERC/CEP 3 EWSR 1 EGFR ALK ROS 1 TMPRSS 2/ ETV 4 ERG
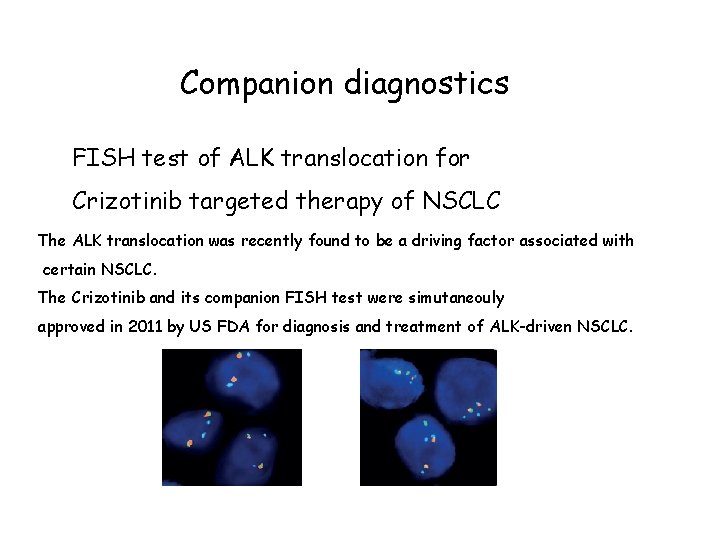
Companion diagnostics FISH test of ALK translocation for Crizotinib targeted therapy of NSCLC The ALK translocation was recently found to be a driving factor associated with certain NSCLC. The Crizotinib and its companion FISH test were simutaneouly approved in 2011 by US FDA for diagnosis and treatment of ALK-driven NSCLC.

Topic 2. Quantitative multi-gene FISH (5 -colour qm. FISH) Previously,most commercial FISH detection kits comprise one probe labeled with a single fluorochrome or two probes labeled with two distinct fluorochromes. With the rapid progress in disease gene discoveries, there is a need to detect multiple genes at a time. Thus, the FISH test employing multiple probes, called qm. FISH, have become popular recently. 15 15

We have prepared the FISH probes in our own laboratory

1. QM-FISH can be used to detect cancer cell heterogeity
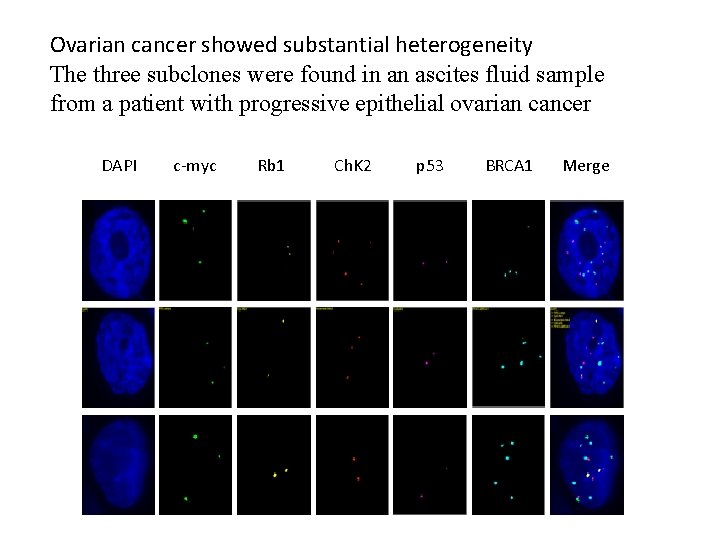
Ovarian cancer showed substantial heterogeneity The three subclones were found in an ascites fluid sample from a patient with progressive epithelial ovarian cancer DAPI c-myc Rb 1 Ch. K 2 p 53 BRCA 1 Merge

Tab. 1. The three subclones in an ascites fluid sample from a patient with progressive epithelial ovarian cancer Clone c-myc Rb 1 Chk 2 P 53 BRCA 1 1 4 2 5 2 4 2 3 2 4 3 3 2 2 2 5 Note: the numbers in Clone column indicate the coding number of the subclones; the numbers in FISH probe coloums (c-myc, Rb 1, Chk 2, p 53 and BRCA 1) indicate average number of signals of each probe within a single nucleus of the cancer cells (50 cells were counted).
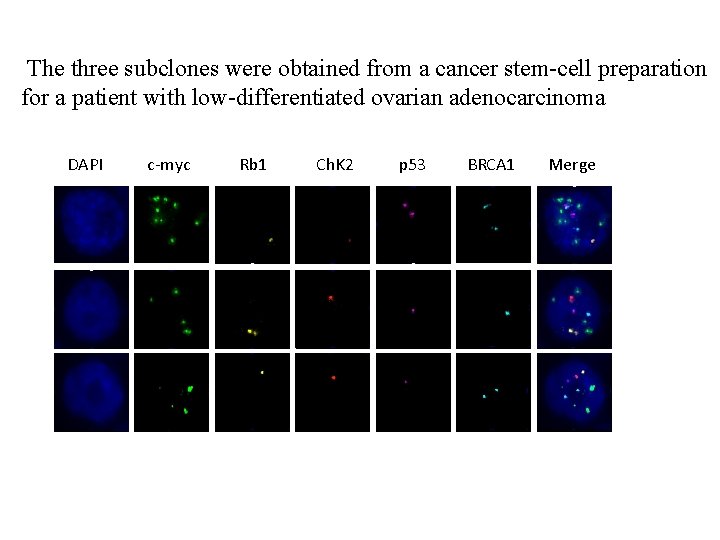
The three subclones were obtained from a cancer stem-cell preparation for a patient with low-differentiated ovarian adenocarcinoma DAPI c-myc Rb 1 Ch. K 2 p 53 BRCA 1 Merge
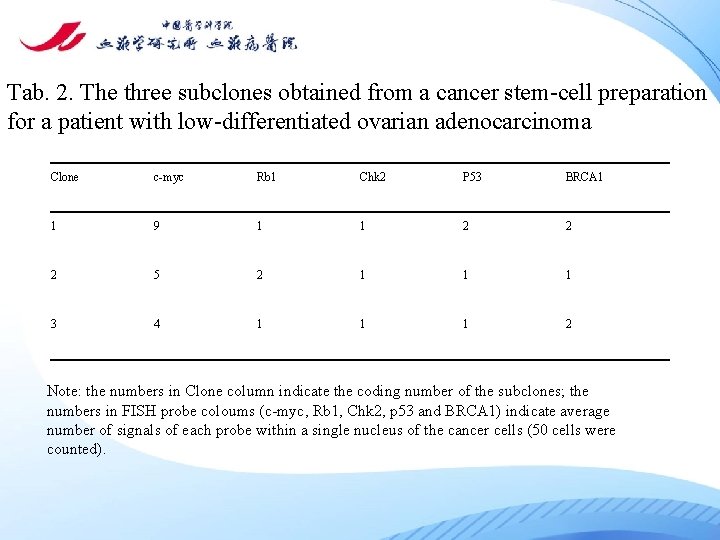
Tab. 2. The three subclones obtained from a cancer stem-cell preparation for a patient with low-differentiated ovarian adenocarcinoma Clone c-myc Rb 1 Chk 2 P 53 BRCA 1 1 9 1 1 2 2 2 5 2 1 1 1 3 4 1 1 1 2 Note: the numbers in Clone column indicate the coding number of the subclones; the numbers in FISH probe coloums (c-myc, Rb 1, Chk 2, p 53 and BRCA 1) indicate average number of signals of each probe within a single nucleus of the cancer cells (50 cells were counted).
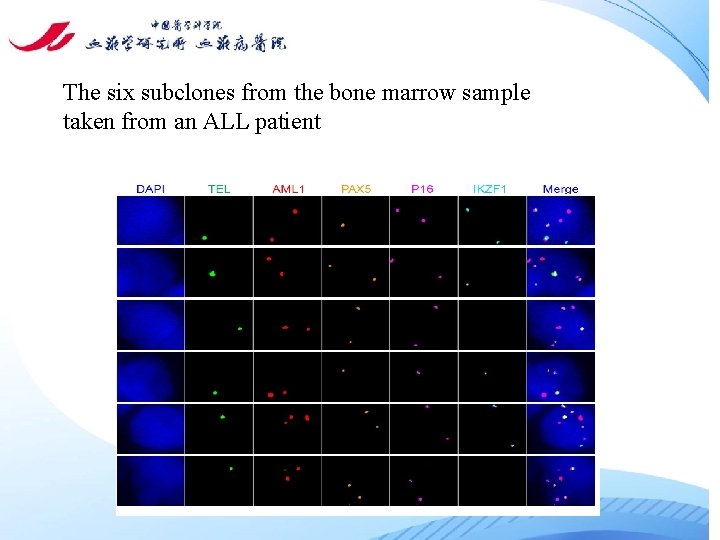
The six subclones from the bone marrow sample taken from an ALL patient
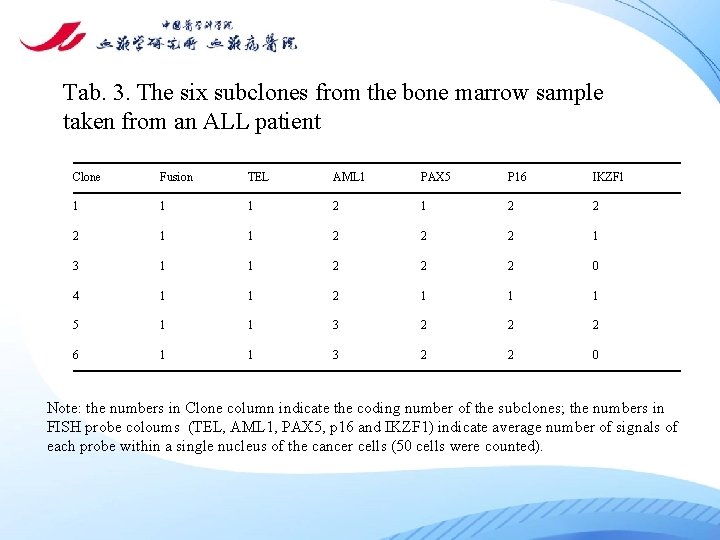
Tab. 3. The six subclones from the bone marrow sample taken from an ALL patient Clone Fusion TEL AML 1 PAX 5 P 16 IKZF 1 1 2 1 2 2 2 1 3 1 1 2 2 2 0 4 1 1 2 1 1 1 5 1 1 3 2 2 2 6 1 1 3 2 2 0 Note: the numbers in Clone column indicate the coding number of the subclones; the numbers in FISH probe coloums (TEL, AML 1, PAX 5, p 16 and IKZF 1) indicate average number of signals of each probe within a single nucleus of the cancer cells (50 cells were counted).
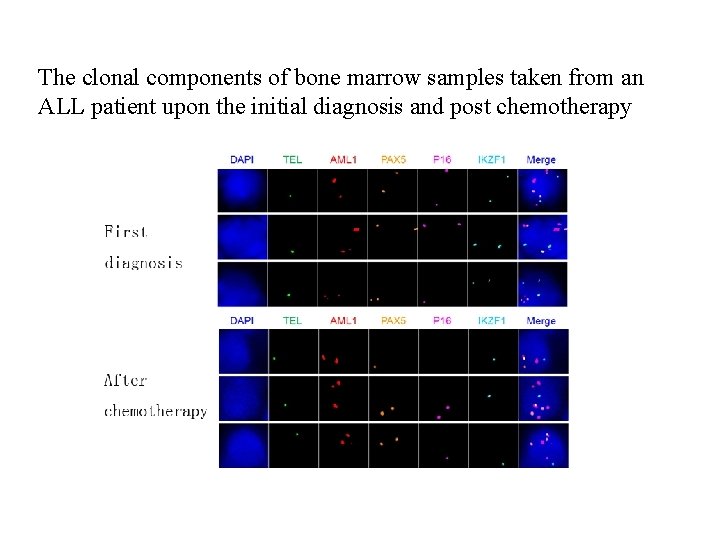
The clonal components of bone marrow samples taken from an ALL patient upon the initial diagnosis and post chemotherapy
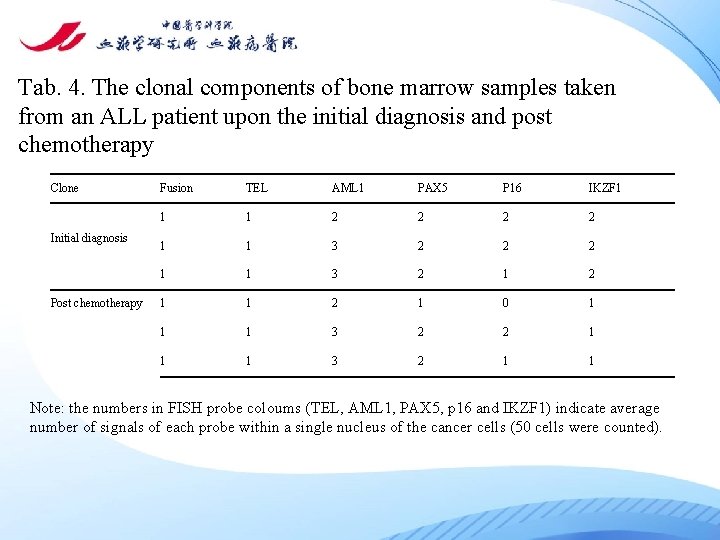
Tab. 4. The clonal components of bone marrow samples taken from an ALL patient upon the initial diagnosis and post chemotherapy Clone Initial diagnosis Post chemotherapy Fusion TEL AML 1 PAX 5 P 16 IKZF 1 1 1 2 2 1 1 3 2 1 1 2 1 0 1 1 1 3 2 2 1 1 1 3 2 1 1 Note: the numbers in FISH probe coloums (TEL, AML 1, PAX 5, p 16 and IKZF 1) indicate average number of signals of each probe within a single nucleus of the cancer cells (50 cells were counted).

2. QM-FISH can be used to study cancer genetic architecture and clonal evolution

Detection of heterogeneity and evolution of subclones in t(8; 21) AML by QM-FISH

The most obvious genetic changes in leukemia is the specific chromosomal translocation; Early in 1972, Dr. Janet Rowley discovered the first translocation between chromosomes 8 and 21 in acute myeloid leukemia (AML)—t(8; 21) , generating AML 1 -ETO fusion gene; Clinically, the t(8; 21) translocation represents the most frequent chromosomal abnormality in AML, occurring in approximately 4%~ 12% of adult and 12%~ 30% of pediatric patients.


36 newly diagnosed primary AML with t(8; 21) cases and 1 paired relapsed case were detected; Genomic copy number alterations (CNA) of AML 1, ETO, WT 1, p 27 and c-kit were investigated by qm. FISH; The clonal heterogeneity, subclonal architecture and putative ancestral evolution in AML 1 -ETO+ AML was assembled; The evolution of dominant clones at diagnosis and relapse were speculated.
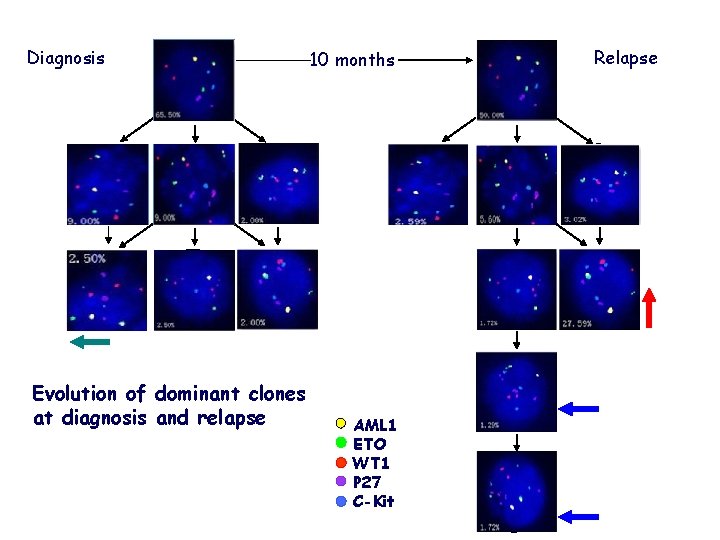
Diagnosis 1 1 1 2 2 2 F ETO AML 1 WT 1 p 27 c-Kit 1 1 1 2 2 2 10 months 1 1 1 3 2 2 F ETO AML 1 WT 1 p 27 c-Kit 1 1 1 2 3 2 F ETO AML 1 WT 1 p 27 c-Kit 1 1 1 2 2 3 F ETO AML 1 WT 1 p 27 c-Kit 1 1 1 3 3 2 F ETO AML 1 WT 1 p 27 c-Kit 1 1 1 2 4 2 F ETO AML 1 WT 1 p 27 c-Kit 1 1 1 2 3 3 F ETO AML 1 WT 1 p 27 c-kit Evolution of dominant clones at diagnosis and relapse 1 1 1 3 2 2 AML 1 ETO WT 1 P 27 C-Kit F ETO AML 1 WT 1 p 27 c-Kit Relapse 1 1 1 2 3 2 F ETO AML 1 WT 1 p 27 c-Kit 1 1 1 2 2 3 F ETO AML 1 WT 1 p 27 c-Kit 1 1 1 2 4 2 F ETO AML 1 WT 1 p 27 c-Kit 1 1 1 2 3 3 F ETO AML 1 WT 1 p 27 c-kit 1 1 1 2 4 3 F ETO AML 1 WT 1 p 27 c-Kit 1 1 1 2 5 3 F ETO AML 1 WT 1 p 27 c-Kit
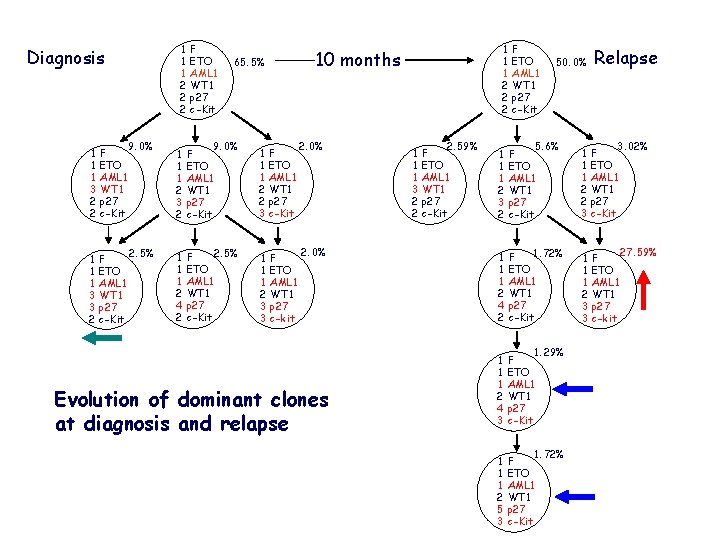
1 1 1 2 2 2 Diagnosis 1 1 1 3 2 2 F ETO AML 1 WT 1 p 27 c-Kit 1 1 1 3 3 2 F ETO AML 1 WT 1 p 27 c-Kit 9. 0% 2. 5% F ETO AML 1 WT 1 p 27 c-Kit 1 1 1 2 2 2 10 months 65. 5% 1 1 1 2 3 2 9. 0% F ETO AML 1 WT 1 p 27 c-Kit 1 1 1 2 2 3 F ETO AML 1 WT 1 p 27 c-Kit 1 1 1 2 4 2 2. 5% F ETO AML 1 WT 1 p 27 c-Kit 1 1 1 2 3 3 F ETO AML 1 WT 1 p 27 c-kit 2. 0% Evolution of dominant clones at diagnosis and relapse 1 1 1 3 2 2 2. 59% F ETO AML 1 WT 1 p 27 c-Kit 50. 0% Relapse 1 1 1 2 3 2 5. 6% F ETO AML 1 WT 1 p 27 c-Kit 1 1 1 2 2 3 3. 02% F ETO AML 1 WT 1 p 27 c-Kit 1 1 1 2 4 2 1. 72% F ETO AML 1 WT 1 p 27 c-Kit 1 1 1 2 3 3 F ETO AML 1 WT 1 p 27 c-kit 1 1 1 2 4 3 1. 29% F ETO AML 1 WT 1 p 27 c-Kit 1 1 1 2 5 3 1. 72% F ETO AML 1 WT 1 p 27 c-Kit 27. 59%

Summary Copy number alterations can be identified in all t(8; 21) AML patients by 5 -color qm. FISH; 2. The genetic architectures observed in t(8; 21) were very diverse, there is a marked subclonal heterogeneity, and the subclones could be aligned in a linear or branching ancestral tree; 3. The dominant subclone in relapse is originated from a major clone at diagnosis, continues to genetically diversify and acquires genetic lesions to generate secondary dominant subclone. 1.

Topic 3. Development of a Sequential FISH strategy up to 20 genes can be detected at the single-cell level

Background (1) While FISH is useful for detecting certain chromosomal abnormalities, the number of genes that can be simultaneously detected is limited. Even with qm. FISH, the number of detected genes is usually less than 5 due to the capacity of currently available fluorescence filter sets. (2) Multi-gene detection at the single-cell level is desirable to enable more precise genotyping of heterogeneous hematology and oncology samples

Sequential qm. FISH strategy a. Five fluorochromes were used to label each set of FISH gene probes, and 5 genes were detected each time using a five-color FISH protocol. b. After the first hybridization, the previous FISH probe set was stripped, and a second set of five-color FISH probes was used for rehybridization. After each hybridization, the fluorescence signals were recorded in 6 fluorescence filter channels that included DAPI, Spectrum Green™, Cy 3™ v 1, Texas Red, Cy 5, and PF-415. c. A digital automatic relocation procedure was used to ensure that exactly the same microscopic field was studied in each stripping and hybridization cycle. d. Therefore, up to 20 genes can be detected within a single nucleus
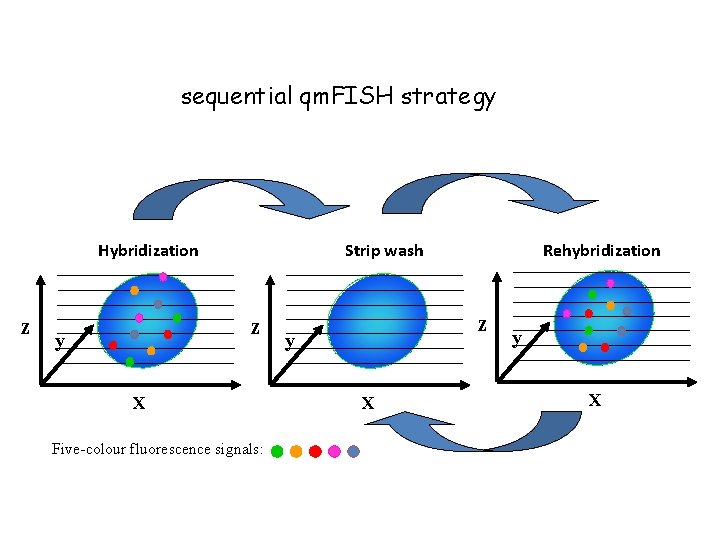
sequential qm. FISH strategy Hybridization z Strip wash z y X Five-colour fluorescence signals: Rehybridization z y X
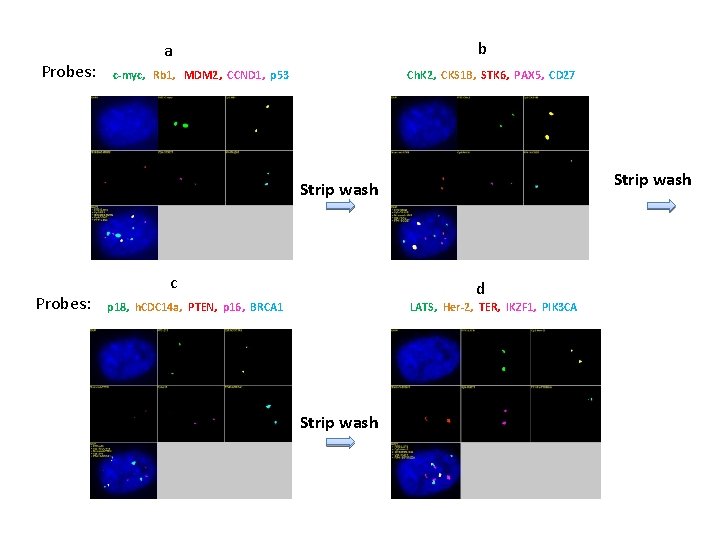
Probes: b a c-myc, Rb 1, MDM 2, CCND 1, p 53 Ch. K 2, CKS 1 B, STK 6, PAX 5, CD 27 Strip wash Probes: c d p 18, h. CDC 14 a, PTEN, p 16, BRCA 1 LATS, Her-2, TER, IKZF 1, PIK 3 CA Strip wash

This method can be used for more precise molecular subtyping or for clonal evolution studies of various types of diseases.

Future direction: 1. Discovery of novel cancer biomarkers 2. Development of novel FISH tests for personalized medicine 3. We welcome any kind of collaborations

Acknowledgement Co-workers: Linping Hu Chengwen Li Jing Ge Jiangman Sun Weixia Xue Xinchun Zhang Collaborators: Tao Cheng, M. D. Zetterberg Anders, M. D. , Ph. D. Wang Jianxiang, M. D. Kun Ru, MD. Ph. D. Ling Shi, Ph. D.

- Slides: 42