FBox Containing Tubby Transcription Factor Family Daisy Robinton

F-Box Containing Tubby Transcription Factor Family Daisy Robinton Goldberg Lab Spring 2006

What is the TUBBY Protein Gene Family? • Studies have shown TUBBY-like proteins to play a vital role in the maitenance and function of neuronal cells during postdifferentiation and development in mammals • Members of this family have been identified in many multicellular organisms, but no single-celled organisms • The tubby domain is highly conserved • This data suggests that tubby proteins have fundamental biological functions • A conserved F-box-containing domain is present in plant TUBBY-like protein members

What is this F-box Protein? • Most F-box domains followed by specific amino acid sequences which are protein-protein interacting domains for recruiting specific proteins and targeting them for ubiquitin mediated proteolysis • F-box proteins regulate cellular processes (cell-cycle transition, transcriptional regulation and signal transduction) • F-box proteins regulate plant growth and development • Serve vital functions in plants as indicated by multiple TLPs, however this may mask the effects of an individual loss-of-function allele

How is this Family Related to Arabidopsis? • In the Arabidopsis plant there is a TUBBY-like protein gene family with 11 members (named At. TLP 1 -11) • All At. TLP members (except 8) have an F-box domain (researchers are not sure, this data was obtained solely via sequencing • Seven of the At. TLP genes are on the first chromosome • Two genes, one gene and one gene are on chromosomes II, III and V, respectively • All members of this family except At. TLP 8 contain a conserved F-box domain • F-box protein has been identified to play an important role during seed development
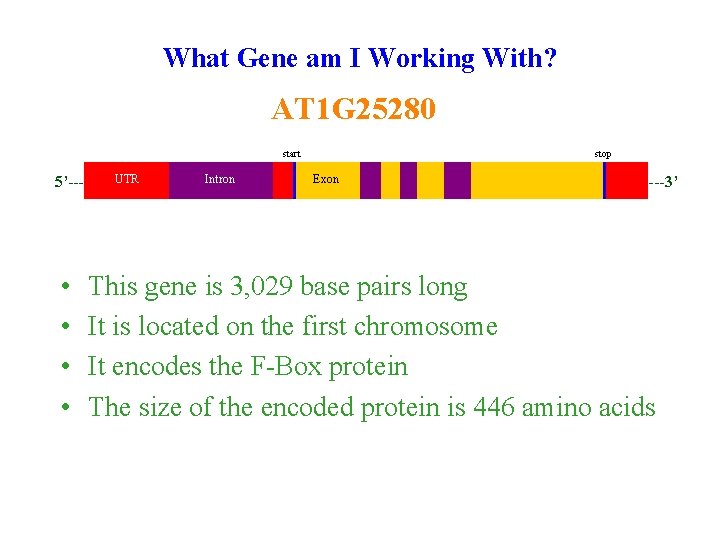
What Gene am I Working With? AT 1 G 25280 start 5’--- • • UTR Intron stop Exon ---3’ This gene is 3, 029 base pairs long It is located on the first chromosome It encodes the F-Box protein The size of the encoded protein is 446 amino acids

• • • Negative Control Positive Control Sil -RT Sil +RT Inflor -RT Inflor +RT Where is my Gene Active? Microarray data indicates that this gene is active in all of the above organs during seed development However, microarrays are not as accurate as the results obtained in RT-PCR because in RT-PCR we amplify the DNA so that the reaction is more sensitive to the presence of smaller DNA fragments My RT-PCR results confirm and agree with the indications from the genechip data. How do I know?

Negative Control Positive Control Sil -RT Sil +RT Inflor -RT Inflor +RT What do my RT-PCR Results Mean? Control Bands Gene-Specific Expected size: 160 bp • The gene-specific bands indicate that m. RNA was made in those organs during seed development, and so we know that our gene is being transcribed during that stage in that organ

Homozygous Mutants Plants 10 and 12 Predicted Size: 291 base pairs Melissa’s DNA w/ Insert Negative Control WT Plant 16 17 Plant 14 15 Plant 12* Plant 13 Plant 11 Plant 8 Plant 9 Plant 10* Plant 7 Positive Control Do I Have any Mutant Plants? • 19 Plants genotyped • 0 Heterozygous mutants • 2 Homozygous mutants • There are three bands in the positive control lane because Melissa’s primers and my primers are present.

Where is the Insert? Actual Insertion Site Via Sequencing Nucleotide 642 LBb 1 Rv 5’--- UTR Intron SALK prediction = insertion site at nucleotide 659 (difference of 17 nucleotides) start stop Exon • The T-DNA is inserted in the forward direction, interacting with the reverse primer • The LBb 1 sequencing reaction begins matching with my WT gene DNA at nucleotide 115 (of the sequencing reaction) ---3’

Does the Insert Disrupt Seed Development? Because I found homozygous mutants it is indicated that this knockout does not cause seed lethality. However, upon observation of seeds using the Nomarski microscope, interesting phenotypes were found. Wild Type Globular Mutant Heart Stage Torpedo Stage Cotyledon Stage NONE FOUND There were no other apparent phenotypic differences upon observing the actual plant.
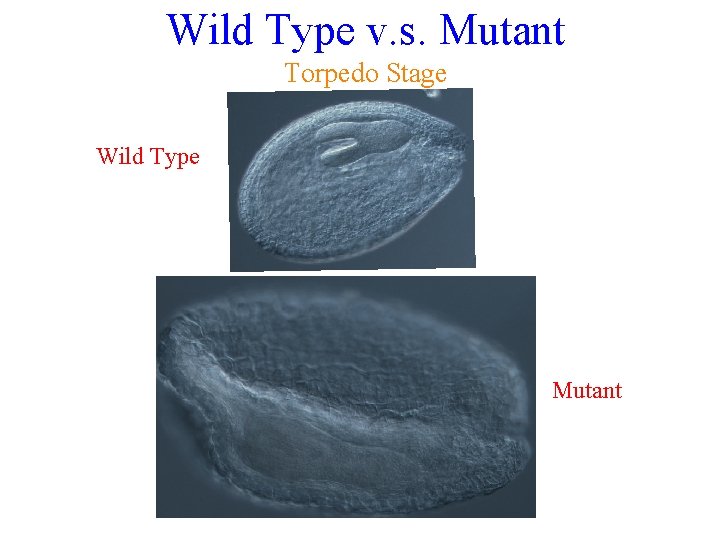
Wild Type v. s. Mutant Torpedo Stage Wild Type Mutant

When and Where does the “Promoter” Regulate the Transcription of my Gene? Colony 8 Colony 7 Colony 6 Colony 5 Negative Control Positive Control Predicted size of the cloned region: 3, 166 base pairs PCR Product • Compare to Sequencing • • • Of approxiamately 97 bacteria colonies, zero were found to be blue Verification of colonies containing recombinant plasmids by digestion with Eco R 1 yielded confirming results The T 7 and SP 6 sequencing seem to be reliable

What is my Second Gene? AT 1 G 76900 start 5’--- Intron • • stop Exon UTR This gene is 2, 506 base pairs long It is located on the first chromosome It encodes a Tubby-Like Protein The size of the encoded protein is 456 amino acids ---3’
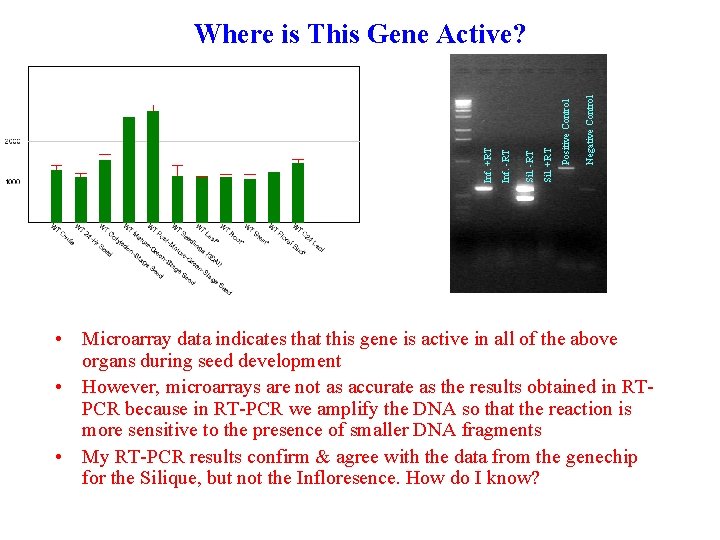
Negative Control Positive Control Sil. +RT Sil. -RT Inf. +RT Where is This Gene Active? • Microarray data indicates that this gene is active in all of the above organs during seed development • However, microarrays are not as accurate as the results obtained in RTPCR because in RT-PCR we amplify the DNA so that the reaction is more sensitive to the presence of smaller DNA fragments • My RT-PCR results confirm & agree with the data from the genechip for the Silique, but not the Infloresence. How do I know?
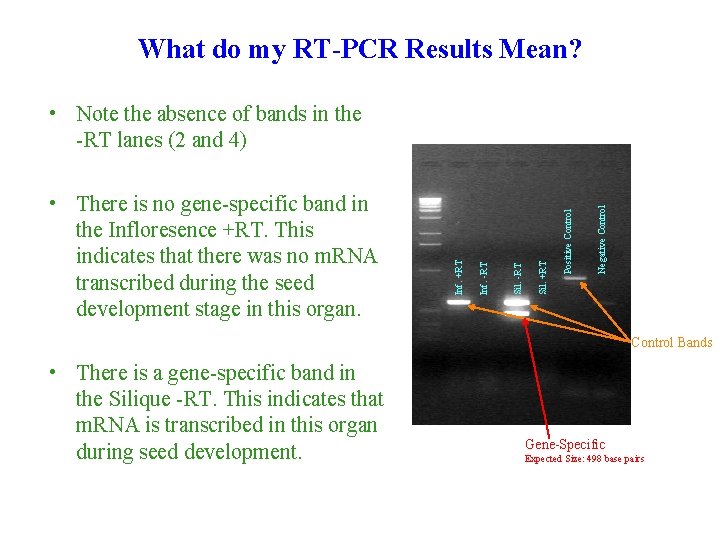
What do my RT-PCR Results Mean? Negative Control Positive Control Sil. +RT Sil. -RT Inf. -RT • There is no gene-specific band in the Infloresence +RT. This indicates that there was no m. RNA transcribed during the seed development stage in this organ. Inf. +RT • Note the absence of bands in the -RT lanes (2 and 4) Control Bands • There is a gene-specific band in the Silique -RT. This indicates that m. RNA is transcribed in this organ during seed development. Gene-Specific Expected Size: 498 base pairs

LB Test Negative Control Positive Control Plant 21 WT Plant 20 Plant 18 Plant 19 Plant 16 Plant 17 Plant 15 Plant 13 Plant 14 Plant 12 Plant 6 Plant 11 Plant 5 Plant 3 Plant 4 Plant 1 Plant 2 Are there any knockouts? Fw/Rv Multiplex • • Fw/LB Rv/LB Separating Primers 28 Plants genotyped 0 Heterozygous Mutants 0 Homozygous Mutants The separating primers genotyping shows that the LB primer was working (Thi’s DNA from her first gene SAIL line was used) Because there are no bands in the other LB reactions, no mutants of any kind were identified. Expected Wild Type Size: 2, 479 base pairs Expected Mutant Size: 2, 495 base pairs The separating primers attends to the problem of the similar size of expected mutant and WT bands

What is the Reason for the Absence of Knockouts? • SALK sent all Wild-Type seeds • The knockout causes seed lethality, and no seeds survived that contained mutant alleles (dominant mutation) • More plants need to be grown and genotyped • The LB was ineffective (for the last batch of genotyping)

Where was the Insert Predicted to Be? LB start FW 5’--- Intron stop Exon • SALK predicts the insert is in the third exon, at nucleotide 2, 151 • The insert is in the reverse direction of my gene, interacting with my forward primer • The lack of identified mutant plants may indicate disruption in seed development UTR ---3’

When and Where does the “Promoter” Regulate the Transcription of my Gene? • Predicted Size of Clone Region 2, 997 base pairs Negative Control Positive Control PCR Product • • Eco. R 1 Digestion gel photo unavailable Results showed three bands in each lane: one about 3. 5 k. B, one about 1. 3 k. B and one about 1. 05 k. B Verification of colonies containing recombinant plasmids by digestion with Eco R 1 yielded confirming results Of approxiamately 200 bacteria colonies, approx. 75 were found to be blue

What is the significance of these results? • • • Studies of the first gene, AT 1 G 25280, show a very interesting seed phenotype at the torpedo stage This phenotype may be an important clue as to the role this gene plays in the Arabidopsis plant Studies of the second gene, AT 1 G 76900, have not yet provided any significant results. What further research should be carried out? AT 1 G 25280 • • • More plants need to be grown and genotyped More siliques and seeds should be studied for phenotypic differences Because the insert is in an intron, RT-PCR of the mutant should be carried out AT 1 G 76900 • • • Get new seeds More plants need to be grown and genotyped Once/if an insert is found, the appropriate experiments should be conducted (as with the first gene)

Thanks! Thanks to SALK, who made our plants (and mixed things up so as to provide valuable learning experiences. Thanks to Ahnthu, Ria, John and Mike for testing, retesting, explaining, teaching, reteaching, etc etc thanks a million times. Thanks to Tomo, Javier, Brandon and Xingjun for behind the scenes work and Nomarski. = Thanks to Jessica for food and pictures haha What do you mean in the wrong place? And finally, thanks to Dr. Goldberg for making it all possible. And the pictures I borrowed for my presentation. : )
- Slides: 21