Factor Analysis Example QianLi Xue Biostatistics Program Harvard

Factor Analysis Example Qian-Li Xue Biostatistics Program Harvard Catalyst | The Harvard Clinical & Translational Science Center Short course, October 28, 2016 1

Example: Frailty § Frailty is “a biologic syndrome of decreased reserve and resistance to stressors, resulting from cumulative declines across multiple physiologic systems, and causing vulnerability to adverse outcomes” (Fried et al. 2001) § Common phenotypes of “frailty” in geriatrics include “weakness, fatigue, weight loss, decreased balance, low levels of physical activity, slowed motor processing and performance, social withdrawal, mild cognitive changes, and increased vulnerability to stressors” (Walston et al. 2006) 2

Example: Frailty Manifest Variables of Frailty: Body composition: Arm circumference Tricep skinfold thickness Body mass index Slowed motor processing and performance: Speed of fast walk Speed of Pegboard test Speed of usual walk Time to do chair stands Muscle Strength: Grip strength Knee extension Hip extension 3

Recap of Basic Characteristics of Exploratory Factor Analysis (EFA) § Most EFA extract orthogonal factors, which may not be a reasonable assumption § Distinction between common and unique variances § EFA is underidentified (i. e. no unique solution) § Remember rotation? Equally good fit with different rotations! § All measures are related to each factor 4

Major steps in EFA 1. Data collection and preparation 2. Choose number of factors to extract 3. Extracting initial factors 4. Rotation to a final solution 5. Model diagnosis/refinement 6. Derivation of factor scales to be used in further analysis 5
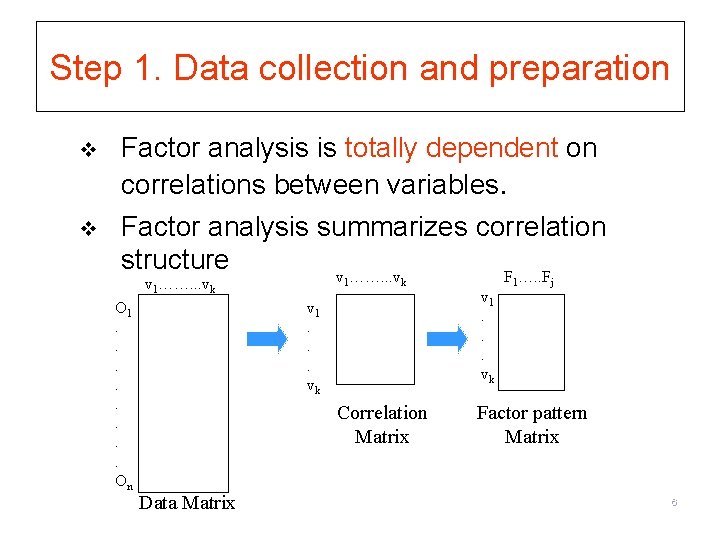
Step 1. Data collection and preparation v v Factor analysis is totally dependent on correlations between variables. Factor analysis summarizes correlation structure v ……. . . v F …. . F v 1……. . . vk O 1. . . . On 1 k v 1. . . vk Correlation Matrix Data Matrix v 1. . . vk 1 j Factor pattern Matrix 6
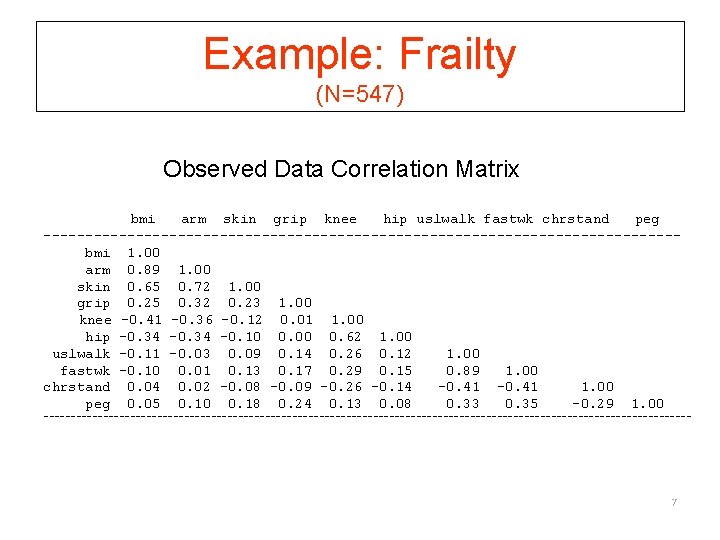
Example: Frailty (N=547) Observed Data Correlation Matrix bmi arm skin grip knee hip uslwalk fastwk chrstand peg --------------------------------------bmi 1. 00 arm 0. 89 1. 00 skin 0. 65 0. 72 1. 00 grip 0. 25 0. 32 0. 23 1. 00 knee -0. 41 -0. 36 -0. 12 0. 01 1. 00 hip -0. 34 -0. 10 0. 00 0. 62 1. 00 uslwalk -0. 11 -0. 03 0. 09 0. 14 0. 26 0. 12 1. 00 fastwk -0. 10 0. 01 0. 13 0. 17 0. 29 0. 15 0. 89 1. 00 chrstand 0. 04 0. 02 -0. 08 -0. 09 -0. 26 -0. 14 -0. 41 1. 00 peg 0. 05 0. 10 0. 18 0. 24 0. 13 0. 08 0. 33 0. 35 -0. 29 1. 00 ------------------------------------------------------------ 7

Step 2. Choose number of factors v Intuitively: The number of uncorrelated constructs that are jointly measured by the Y’s. v Only useful if number of factors is less than number of Y’s (recall “data reduction”). v Estimability: Is there enough information in the data to estimate all of the parameters in the factor analysis? May be constrained to a certain number of factors. 8

Step 2. Choosing number of factors Use Principal Components Analysis (PCA) to help decide § Similar to “factor” analysis, but conceptually quite different! § number of “factors” is equivalent to number of variables § each “factor” or principal component is a weighted combination of the input variables Y 1 …. Yn: P 1 = a 11 Y 1 + a 12 Y 2 + …. a 1 n. Yn § Principal components ARE NOT latent variable § Does not differentiate between common and unique variances 9

Choosing Number of Factors /* Principal Components analysis */ Proc factor data=frailty METHOD=PRIN outstat=abc. pca_all plots=(scree); var bmi arm skin grip knee hip uslwalk fastwk chrstand peg; %parallel(data=frailty, niter=1000, statistic=Median); run; 10
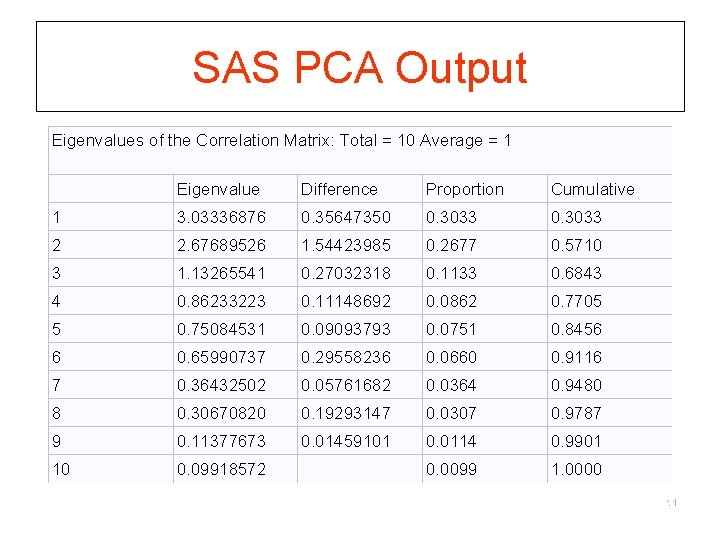
SAS PCA Output Eigenvalues of the Correlation Matrix: Total = 10 Average = 1 Eigenvalue Difference Proportion Cumulative 1 3. 03336876 0. 35647350 0. 3033 2 2. 67689526 1. 54423985 0. 2677 0. 5710 3 1. 13265541 0. 27032318 0. 1133 0. 6843 4 0. 86233223 0. 11148692 0. 0862 0. 7705 5 0. 75084531 0. 09093793 0. 0751 0. 8456 6 0. 65990737 0. 29558236 0. 0660 0. 9116 7 0. 36432502 0. 05761682 0. 0364 0. 9480 8 0. 30670820 0. 19293147 0. 0307 0. 9787 9 0. 11377673 0. 01459101 0. 0114 0. 9901 10 0. 09918572 0. 0099 1. 0000 11

Step 2. Choosing number of factors § To select how many factors to use, evaluate eigenvalues from PCA § Two interpretations: § eigenvalue equivalent number of variables which the factor represents § eigenvalue amount of variance in the data described by the factor. § Criteria to go by: § § § number of eigenvalues > 1 (Kaiser-Guttman Criterion) scree plot parallel analysis % variance explained comprehensibility 12
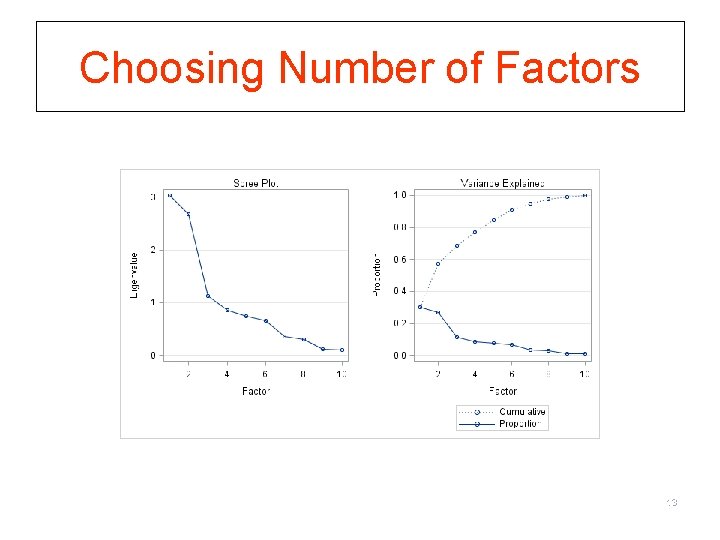
Choosing Number of Factors 13

Parallel Analysis (Hayton, Allen, & Scarpello (2004) § Eigenvalues (EV) that would be expected from random data are compared to those produced by the data § If EV(random data) > EV(real data), the derived factors are mostly random noise § How to do this in SAS http: //www 2. sas. com/proceedings/sugi 28/090 -28. pdf § How to do this in STATA Type “findit fapara” in STATA to locate the program for free download Reference: http: //www. ats. ucla. edu/stata/faq/parallel. htm 14
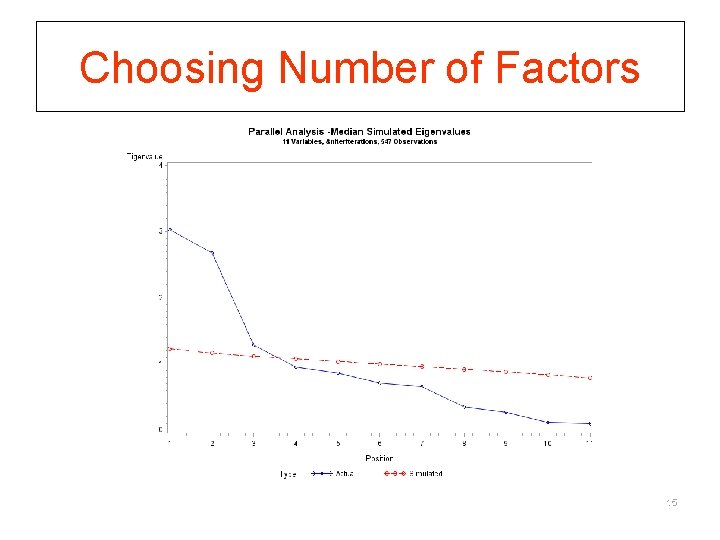
Choosing Number of Factors 15

Accuracy of Retention Criteria § EV > 1 § Tends to always over estimate number of factor § Accuracy increase with small number variables & communalities are high § Scree Test § More accurate than EV>1 § Subjective and sometimes ambiguous § Parallel Test § Most accurate § Becoming the standard 16

Step 3. Extracting initial factors Using MLE Proc factor data=frailty METHOD=ML priors=smc msa residual rotate=varimax reorder outstat=abc. fa_all plots=(scree initloadings); var bmi arm skin grip knee hip uslwalk fastwk chrstand peg; run; 17
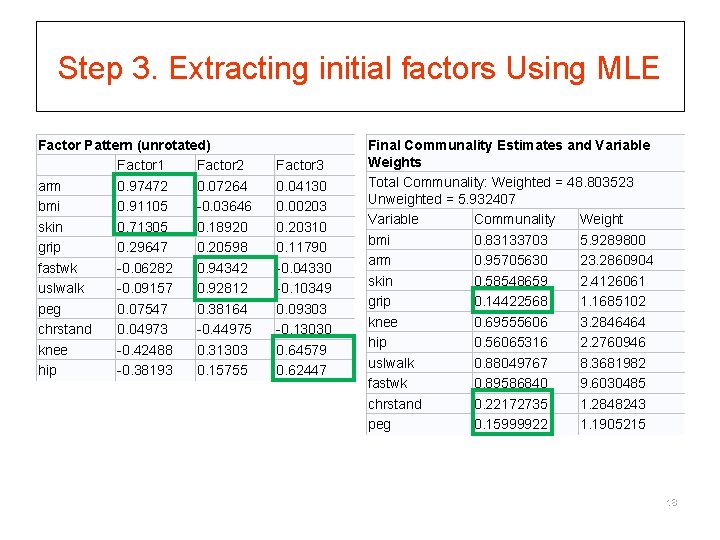
Step 3. Extracting initial factors Using MLE Factor Pattern (unrotated) Factor 1 Factor 2 arm 0. 97472 0. 07264 bmi 0. 91105 -0. 03646 skin 0. 71305 0. 18920 grip 0. 29647 0. 20598 fastwk -0. 06282 0. 94342 uslwalk -0. 09157 0. 92812 peg 0. 07547 0. 38164 chrstand 0. 04973 -0. 44975 knee -0. 42488 0. 31303 hip -0. 38193 0. 15755 Factor 3 0. 04130 0. 00203 0. 20310 0. 11790 -0. 04330 -0. 10349 0. 09303 -0. 13030 0. 64579 0. 62447 Final Communality Estimates and Variable Weights Total Communality: Weighted = 48. 803523 Unweighted = 5. 932407 Variable Communality Weight bmi 0. 83133703 5. 9289800 arm 0. 95705630 23. 2860904 skin 0. 58548659 2. 4126061 grip 0. 14422568 1. 1685102 knee 0. 69555606 3. 2846464 hip 0. 56065316 2. 2760946 uslwalk 0. 88049767 8. 3681982 fastwk 0. 89586840 9. 6030485 chrstand 0. 22172735 1. 2848243 peg 0. 15999922 1. 1905215 18

Step 4. Factor Rotation § Steps 2 and 3 determines the minimum number of factors needed to account for observed correlations § After obtaining initial orthogonal factors, we want to find more easily interpretable factors via rotations § While keeping the number of factors and communalities of Ys fixed!!! § Rotation does NOT improve fit! 19

Step 4. Factor Rotation § All solutions are relatively the same § Goal is simple structure § Most construct validation assumes simple (typically rotated) structure. § Rotation does NOT improve fit! 20
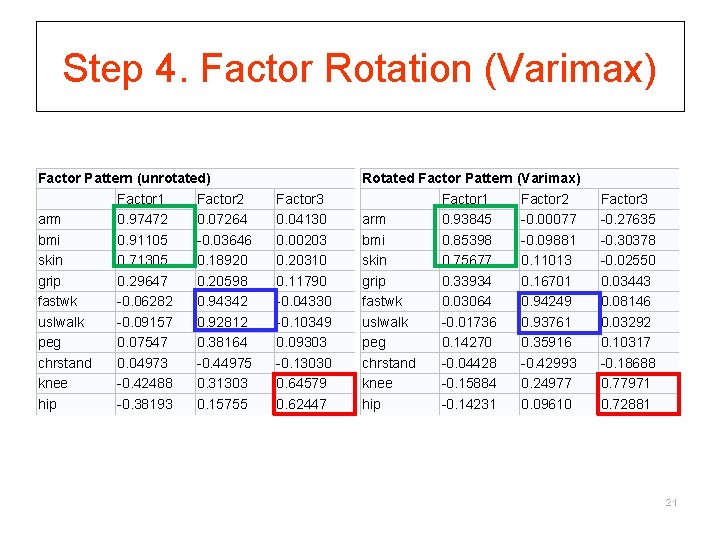
Step 4. Factor Rotation (Varimax) Factor Pattern (unrotated) Factor 1 Factor 2 arm 0. 97472 0. 07264 bmi 0. 91105 -0. 03646 skin 0. 71305 0. 18920 grip 0. 29647 0. 20598 fastwk -0. 06282 0. 94342 uslwalk -0. 09157 0. 92812 peg 0. 07547 0. 38164 chrstand 0. 04973 -0. 44975 knee -0. 42488 0. 31303 hip -0. 38193 0. 15755 Factor 3 0. 04130 0. 00203 0. 20310 0. 11790 -0. 04330 -0. 10349 0. 09303 -0. 13030 0. 64579 0. 62447 Rotated Factor Pattern (Varimax) Factor 1 Factor 2 arm 0. 93845 -0. 00077 bmi 0. 85398 -0. 09881 skin 0. 75677 0. 11013 grip 0. 33934 0. 16701 fastwk 0. 03064 0. 94249 uslwalk -0. 01736 0. 93761 peg 0. 14270 0. 35916 chrstand -0. 04428 -0. 42993 knee -0. 15884 0. 24977 hip -0. 14231 0. 09610 Factor 3 -0. 27635 -0. 30378 -0. 02550 0. 03443 0. 08146 0. 03292 0. 10317 -0. 18688 0. 77971 0. 72881 21
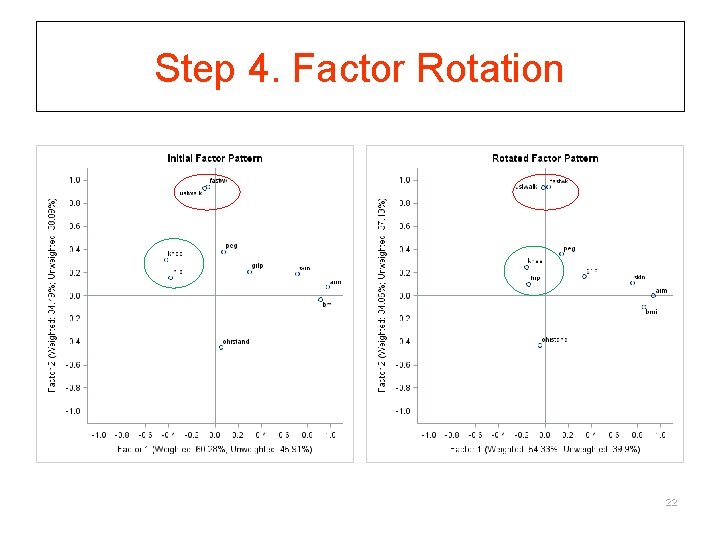
Step 4. Factor Rotation 22

Step 4. Factor Rotation (Promax) Promax Rotated Factor Pattern (Standardized) Factor 1 Factor 2 Factor 3 arm 0. 93580 0. 02339 -0. 09848 bmi 0. 84202 -0. 07008 -0. 13502 skin 0. 79870 0. 09709 0. 12514 grip 0. 36553 0. 15612 0. 09186 uslwalk -0. 03965 0. 95788 -0. 08761 fastwk 0. 02314 0. 95283 -0. 02381 peg 0. 16611 0. 34652 0. 09988 chrstand -0. 07896 -0. 40683 -0. 16182 knee 0. 01027 0. 12041 0. 79711 hip 0. 02043 -0. 02875 0. 76406 Rotated Factor Pattern (Varimax) Factor 1 Factor 2 arm 0. 93845 -0. 00077 bmi 0. 85398 -0. 09881 skin 0. 75677 0. 11013 grip 0. 33934 0. 16701 fastwk 0. 03064 0. 94249 uslwalk -0. 01736 0. 93761 peg 0. 14270 0. 35916 chrstand -0. 04428 -0. 42993 knee -0. 15884 0. 24977 hip -0. 14231 0. 09610 Factor 3 -0. 27635 -0. 30378 -0. 02550 0. 03443 0. 08146 0. 03292 0. 10317 -0. 18688 0. 77971 0. 72881 Inter-Factor Correlations Factor 1 Factor 2 Factor 3 Factor 1 1. 00000 -0. 02794 -0. 39917 Factor 2 -0. 02794 1. 00000 0. 27183 Factor 3 -0. 39917 0. 27183 1. 00000 23
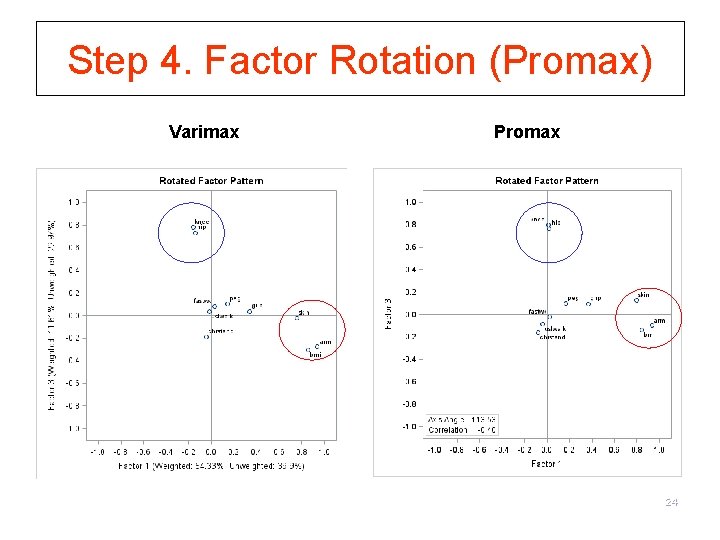
Step 4. Factor Rotation (Promax) Varimax Promax 24
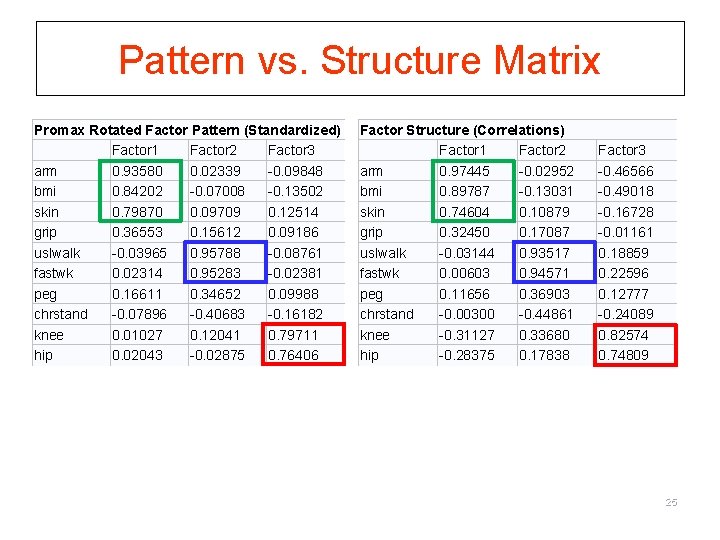
Pattern vs. Structure Matrix Promax Rotated Factor Pattern (Standardized) Factor 1 Factor 2 Factor 3 arm 0. 93580 0. 02339 -0. 09848 bmi 0. 84202 -0. 07008 -0. 13502 skin 0. 79870 0. 09709 0. 12514 grip 0. 36553 0. 15612 0. 09186 uslwalk -0. 03965 0. 95788 -0. 08761 fastwk 0. 02314 0. 95283 -0. 02381 peg 0. 16611 0. 34652 0. 09988 chrstand -0. 07896 -0. 40683 -0. 16182 knee 0. 01027 0. 12041 0. 79711 hip 0. 02043 -0. 02875 0. 76406 Factor Structure (Correlations) Factor 1 Factor 2 arm 0. 97445 -0. 02952 bmi 0. 89787 -0. 13031 skin 0. 74604 0. 10879 grip 0. 32450 0. 17087 uslwalk -0. 03144 0. 93517 fastwk 0. 00603 0. 94571 peg 0. 11656 0. 36903 chrstand -0. 00300 -0. 44861 knee -0. 31127 0. 33680 hip -0. 28375 0. 17838 Factor 3 -0. 46566 -0. 49018 -0. 16728 -0. 01161 0. 18859 0. 22596 0. 12777 -0. 24089 0. 82574 0. 74809 25

Step 5. Model Diagnostics: Goodness-of-Fit Significance Tests Based on 547 Observations Test DF Chi-Square Pr > Chi. Sq H 0: No common factors 45 2875. 6589 <. 0001 HA: At least one common factor H 0: 3 Factors are sufficient 18 45. 1733 0. 0004 HA: More factors are needed 26

Step 5. Model Diagnostics: Residual Correlations With Uniqueness on the Diagonal bmi arm skin grip knee bmi 0. 16866 -0. 00004 0. 00437 -0. 01782 -0. 01083 arm -0. 00004 0. 04294 -0. 00055 0. 00649 0. 00358 skin 0. 00437 -0. 00055 0. 41451 -0. 04196 -0. 00979 grip -0. 01782 0. 00649 -0. 04196 0. 85577 -0. 00439 knee -0. 01083 0. 00358 -0. 00979 -0. 00439 0. 30444 hip 0. 01499 -0. 00468 0. 01556 0. 00224 0. 00151 uslwalk 0. 00359 -0. 00047 -0. 00157 -0. 00833 0. 00177 fastwk -0. 00355 0. 00071 0. 00134 0. 00163 -0. 00199 chrstand -0. 02409 0. 00652 -0. 00236 0. 00008 -0. 01761 peg -0. 00339 -0. 00528 0. 03339 0. 12758 -0. 02066 hip 0. 01499 -0. 00468 0. 01556 0. 00224 0. 00151 0. 43935 -0. 00076 0. 00327 0. 03101 -0. 00831 uslwalk 0. 00359 -0. 00047 -0. 00157 -0. 00833 0. 00177 -0. 00076 0. 11950 0. 00044 -0. 00513 -0. 00928 fastwk -0. 00355 0. 00071 0. 00134 0. 00163 -0. 00199 0. 00327 0. 00044 0. 10413 0. 00904 -0. 00060 chrstand -0. 02409 0. 00652 -0. 00236 0. 00008 -0. 01761 0. 03101 -0. 00513 0. 00904 0. 77827 -0. 11063 peg -0. 00339 -0. 00528 0. 03339 0. 12758 -0. 02066 -0. 00831 -0. 00928 -0. 00060 -0. 11063 0. 84000 Root Mean Square Off-Diagonal Residuals: Overall = 0. 02799287 bmi arm skin grip knee hip uslwalk fastwk chrstand peg 0. 0120016 0. 0040602 0. 0189818 0. 0453301 0. 0104969 0. 0130721 0. 0047294 0. 0035657 0. 0397855 0. 0579768 27
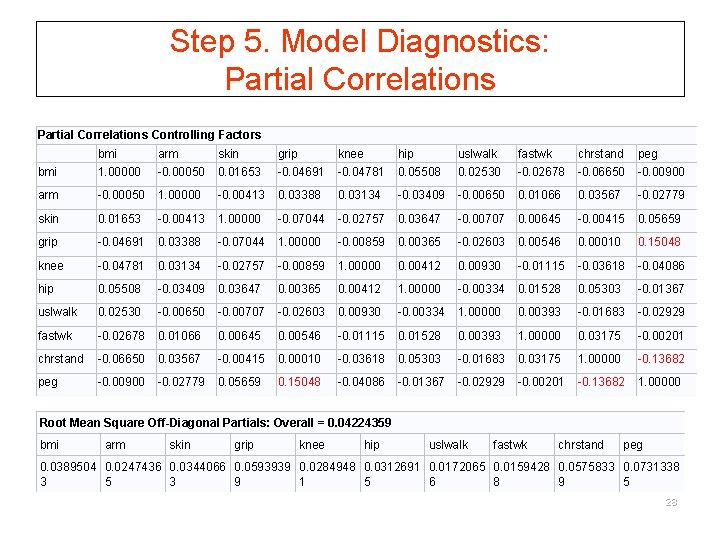
Step 5. Model Diagnostics: Partial Correlations Controlling Factors bmi arm skin grip knee hip uslwalk fastwk chrstand peg bmi 1. 00000 -0. 00050 0. 01653 -0. 04691 -0. 04781 0. 05508 0. 02530 -0. 02678 -0. 06650 -0. 00900 arm -0. 00050 1. 00000 -0. 00413 0. 03388 0. 03134 -0. 03409 -0. 00650 0. 01066 0. 03567 -0. 02779 skin 0. 01653 -0. 00413 1. 00000 -0. 07044 -0. 02757 0. 03647 -0. 00707 0. 00645 -0. 00415 0. 05659 grip -0. 04691 0. 03388 -0. 07044 1. 00000 -0. 00859 0. 00365 -0. 02603 0. 00546 0. 00010 0. 15048 knee -0. 04781 0. 03134 -0. 02757 -0. 00859 1. 00000 0. 00412 0. 00930 -0. 01115 -0. 03618 -0. 04086 hip 0. 05508 -0. 03409 0. 03647 0. 00365 0. 00412 1. 00000 -0. 00334 0. 01528 0. 05303 -0. 01367 uslwalk 0. 02530 -0. 00650 -0. 00707 -0. 02603 0. 00930 -0. 00334 1. 00000 0. 00393 -0. 01683 -0. 02929 fastwk -0. 02678 0. 01066 0. 00645 0. 00546 -0. 01115 0. 01528 0. 00393 1. 00000 0. 03175 -0. 00201 chrstand -0. 06650 0. 03567 -0. 00415 0. 00010 -0. 03618 0. 05303 -0. 01683 0. 03175 1. 00000 -0. 13682 peg -0. 00900 -0. 02779 0. 05659 0. 15048 -0. 04086 -0. 01367 -0. 02929 -0. 00201 -0. 13682 1. 00000 Root Mean Square Off-Diagonal Partials: Overall = 0. 04224359 bmi arm skin grip knee hip uslwalk fastwk chrstand peg 0. 0389504 0. 0247436 0. 0344066 0. 0593939 0. 0284948 0. 0312691 0. 0172065 0. 0159428 0. 0575833 0. 0731338 3 5 3 9 1 5 6 8 9 5 28

Step 6. Model Refinement: Analysis of Cronbach Alpha /* Cronbach Alpha */ proc corr data=frailty nomiss alpha plots; var grip knee hip; run; proc corr data=frailty nomiss alpha plots; var uslwalk fastwk chrstand 2 peg; run; 29

Step 6. Model Refinement: Item Deletion? Cronbach Coefficient Alpha Variables Alpha Raw 0. 033632 Standardized 0. 439547 Variables Alpha Raw 0. 584027 Standardized 0. 570175 Cronbach Coefficient Alpha with Deleted Variable Raw Variables Standardized Variables Corr. with Total Alpha uslwalk 0. 859578 -. 004977 0. 616827 0. 262410 Raw Variables Standardized Variables Corr. with Total Alpha grip 0. 003089 0. 762508 0. 002980 0. 762663 fastwk 0. 832333 -. 011250 0. 634211 0. 245257 knee 0. 091319 -. 002565 0. 444197 -. 009871 chrstand 2 -. 037600 0. 681264 -. 028611 0. 765927 hip 0. 078178 0. 005476 0. 430167 0. 020331 peg 0. 349941 0. 649959 0. 317186 0. 526887 Uniqueness of Grip = 0. 85577 Uniqueness of Chair Stand = 0. 77827 30

Step 7. Derivation of Factor Scores § Each object (e. g. each person) gets a factor score for each factor: § The factors themselves are variables § “Object’s” score is weighted combination of scores on input variables § § These weights are NOT the factor loadings! Different approaches exist for estimating (e. g. regression method) Factor scores are not unique Using factors scores instead of factor indicators can reduce measurement error, but does NOT remove it. § Therefore, using factor scores as predictors in conventional regressions leads to inconsistent coefficient estimators! 31

Step 7. Derivation of Factor Scores Proc factor data=frailty method=ML score outstat=fact priors=smc msa residual rotate=varimax reorder outstat=abc. fa_all plots=(scree initloadings); var bmi arm skin grip knee hip uslwalk fastwk chrstand peg; run; /* Calculate factor scores */ proc score data=frailty score=fact out=abc. scores; var bmi arm skin grip knee hip uslwalk fastwk chrstand peg; run; 32

Exploratory vs. Confirmatory Factor Analysis § Exploratory: § summarize data § describe correlation structure between variables § generate hypotheses § Confirmatory § Testing correlated measurement errors § Redundancy test of one-factor vs. multi-factor models § Measurement invariance test comparing a model across groups § Orthogonality tests 33
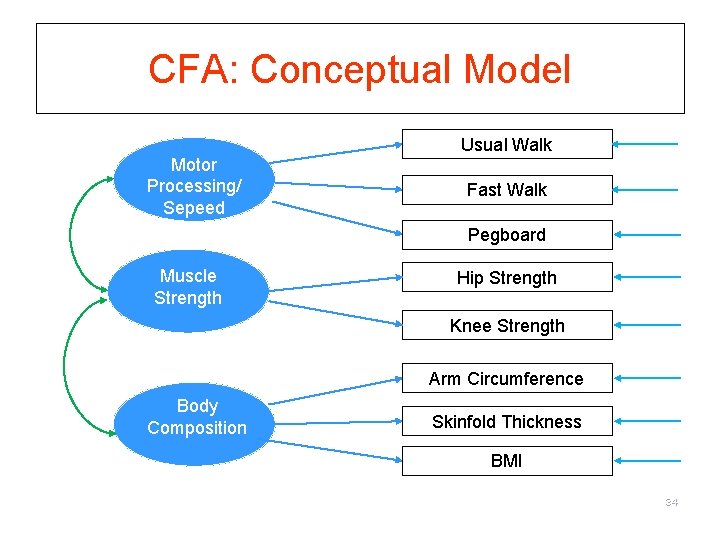
CFA: Conceptual Model Motor Processing/ Sepeed Usual Walk Fast Walk Pegboard Muscle Strength Hip Strength Knee Strength Arm Circumference Body Composition Skinfold Thickness BMI 34

SAS Code /* Confirmatory factor analysis */ proc calis data=frailty modification; factor Body_Factor ---> bmi arm skin = load 1 -load 3, Speed_Factor ---> uslwalk fastwk peg = load 4 load 6, Strength_Factor ---> knee hip = load 7 -load 8; pvar Body_Factor Speed_Factor Strength_Factor = 3*1; cov Body_Factor Speed_Factor = 0. ; run; 35
SAS Output: Standardized Loadings 36
SAS Output: Factor Correlations 37

Model Fit Statistics § Goodness-of-fit tests based on predicted vs. observed covariances: 1. 2 tests § § § d. f. =(# non-redundant components in S) – (# unknown parameters in the model) Null hypothesis: lack of significant difference between ( ) and S Sensitive to sample size Sensitive to the assumption of multivariate normality 2 tests for difference between NESTED models 2. Root Mean Square Error of Approximation (RMSEA) § § A population index, insensitive to sample size Test a null hypothesis of poor fit Availability of confidence interval <0. 10 “good”, <0. 05 “very good” (Steiger, 1989, p. 81) 3. Standardized Root Mean Residual (SRMR) § Squared root of the mean of the squared standardized residuals § SRMR = 0 indicates “perfect” fit, <. 05 “good” fit, <. 08 adequate fit 38

Model Fit Statistics § Goodness-of-fit tests comparing the given model with an alternative model Comparative Fit Index (CFI; Bentler 1989) 1. § § § 2. The Tucker-Lewis Index (TLI) or Non-Normed Fit Index (NNFI) § § § compares the existing model fit with a null model which assumes uncorrelated variables in the model (i. e. the "independence model") Interpretation: CFI× 100=% of the covariation in the data can be explained by the given model CFI ranges from 0 to 1, with 1 indicating a very good fit; acceptable fit if CFI>0. 9 Relatively independent of sample size (Marsh et al. 1988, 1996) NNFI >=. 95 indicates a good model fit, <0. 9 poor fit More about these later 39
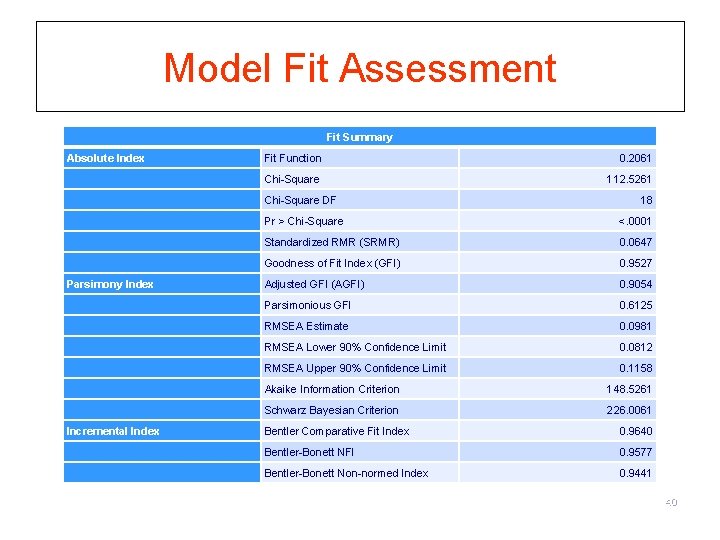
Model Fit Assessment Fit Summary Absolute Index Fit Function 0. 2061 Chi-Square 112. 5261 Chi-Square DF Pr > Chi-Square <. 0001 Standardized RMR (SRMR) 0. 0647 Goodness of Fit Index (GFI) 0. 9527 Parsimony Index Adjusted GFI (AGFI) 0. 9054 Parsimonious GFI 0. 6125 RMSEA Estimate 0. 0981 RMSEA Lower 90% Confidence Limit 0. 0812 RMSEA Upper 90% Confidence Limit 0. 1158 Akaike Information Criterion 148. 5261 Schwarz Bayesian Criterion 226. 0061 Incremental Index Bentler Comparative Fit Index 0. 9640 Bentler-Bonett NFI 0. 9577 Bentler-Bonett Non-normed Index 0. 9441 18 40

Lagrangian Multiplier Test (LMT) § For comparison of nested models § Only requires fitting of the restricted model § Based on the score s( u)= log. L( u)/ u, where L( u) is the unrestricted likelihood function § s( u)=0 when evaluated at the MLE of u § The Idea: substitute the MLE of r, assess departure from 0 § LM ~ 2 with d. f. =difference in the d. f. of the two nested models § Modification index (MI): expected drop in chi-square if the parameter that is fixed or constrained to be equal to other parameters is freely estimated 41
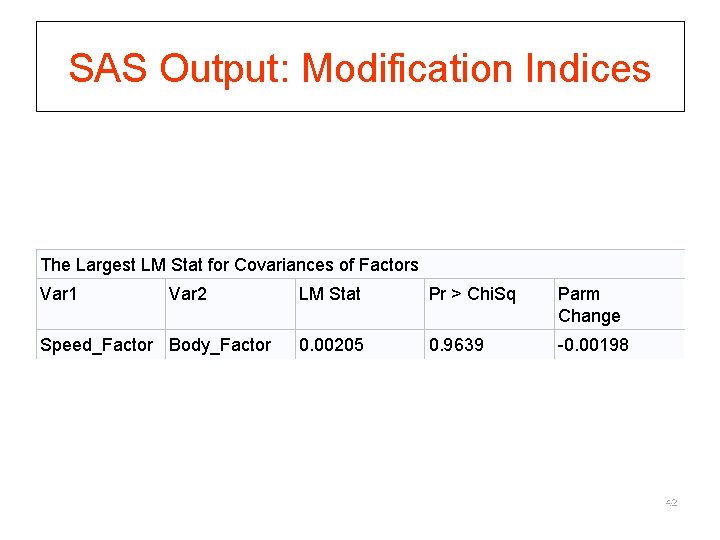
SAS Output: Modification Indices The Largest LM Stat for Covariances of Factors Var 1 Var 2 Speed_Factor Body_Factor LM Stat Pr > Chi. Sq Parm Change 0. 00205 0. 9639 -0. 00198 42
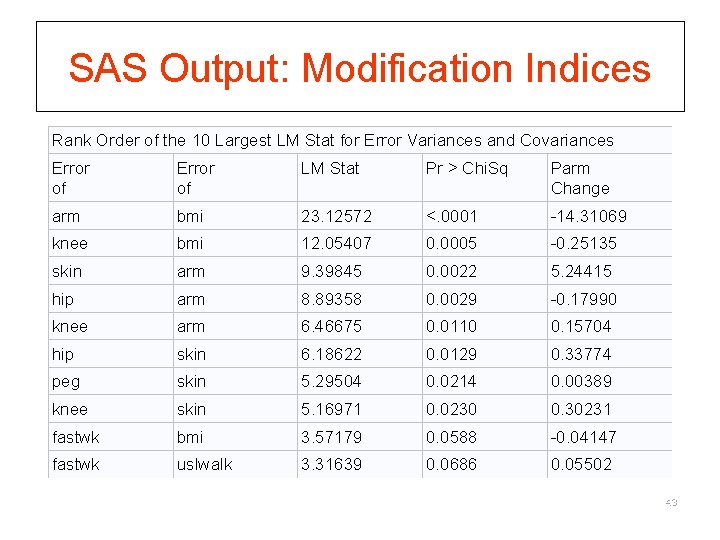
SAS Output: Modification Indices Rank Order of the 10 Largest LM Stat for Error Variances and Covariances Error of LM Stat Pr > Chi. Sq Parm Change arm bmi 23. 12572 <. 0001 -14. 31069 knee bmi 12. 05407 0. 0005 -0. 25135 skin arm 9. 39845 0. 0022 5. 24415 hip arm 8. 89358 0. 0029 -0. 17990 knee arm 6. 46675 0. 0110 0. 15704 hip skin 6. 18622 0. 0129 0. 33774 peg skin 5. 29504 0. 0214 0. 00389 knee skin 5. 16971 0. 0230 0. 30231 fastwk bmi 3. 57179 0. 0588 -0. 04147 fastwk uslwalk 3. 31639 0. 0686 0. 05502 43
- Slides: 43