F sarahfriMV Multivariate Linkage and Association Sarah Medland





















- Slides: 51

F: sarahfri_MV

Multivariate Linkage and Association Sarah Medland Manuel Ferreira -with heavy borrowing from Kate Morley and Frühling Rijsdijk

MV analyses can address ¢ Questions of common aetiology Same gene (snp) l Co-incidental covariation due to LD between two different genes l Co-variation due to shared social/environmental risk factors l Aust. Albatross

MV analyses can address ¢ Questions of common aetiology Same gene (snp) l Co-incidental covariation due to LD between two different genes l Co-variation due to shared social/environmental risk factors l ¢ Pleiotropy occurs when a single gene influences multiple phenotypic traits. Bilby

Studying multiple phenotypes… ¢ Run multiple univariate analyses on correlated traits Bandicoot


Studying multiple phenotypes… ¢ Run multiple univariate analyses l Correct for multiple testing… • Bonferroni • Correction for equivalent number of independent variables l Doesn’t really address the idea of common aetiology Blue ringed octopus

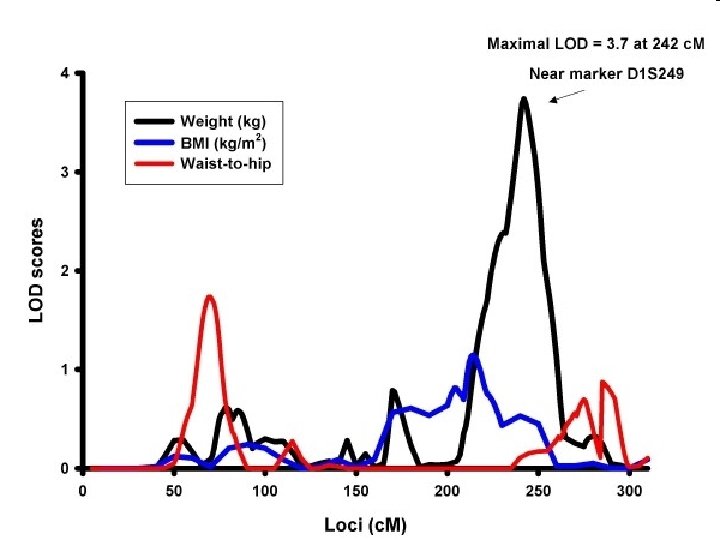
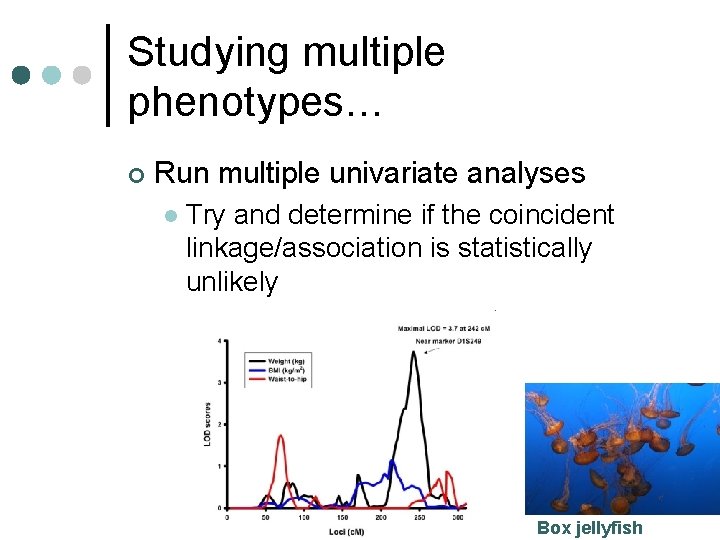
Studying multiple phenotypes… ¢ Run multiple univariate analyses l Try and determine if the coincident linkage/association is statistically unlikely Box jellyfish

Studying multiple phenotypes… ¢ Run multiple univariate analyses l Try and determine if the coincident linkage/association is statistically unlikely • Simulate/Permute data and assess how often this group of traits reaches this pattern of sig. by chance Brown Snake

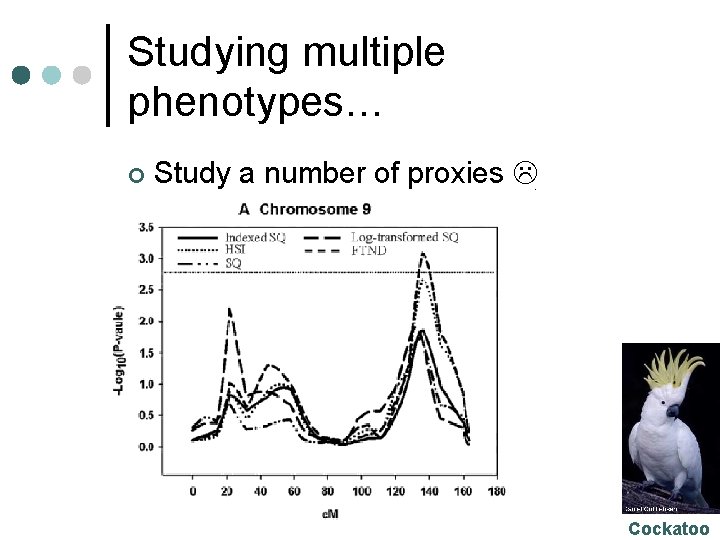
Studying multiple phenotypes… ¢ Study a number of proxies Cockatoo

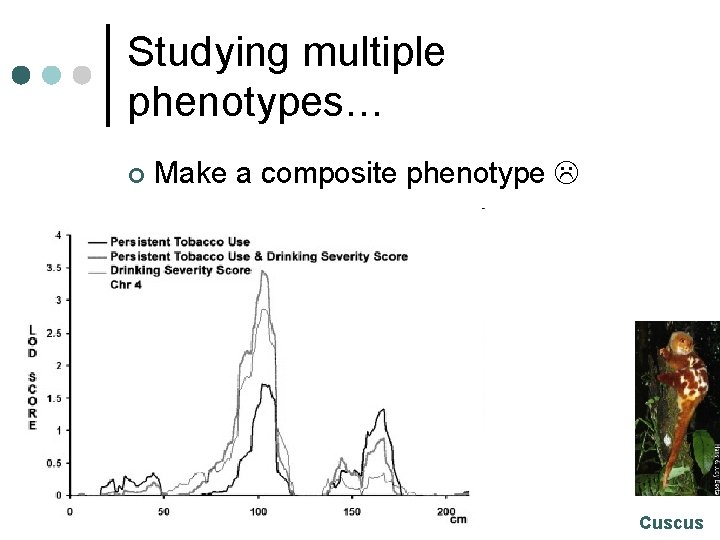
Studying multiple phenotypes… ¢ Make a composite phenotype Cuscus

Studying multiple phenotypes… ¢ Make a factor score combine both factor level and traitspecific effects l latent factor effects are inherently pleiotropic l residual effects are not l assumes factor loadings are constant across genome l Cuscus (again)

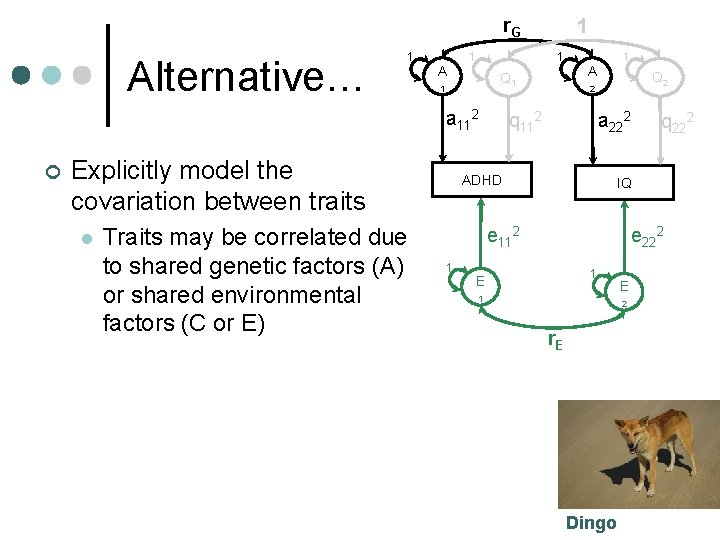
r. G Alternative… 1 A 1 1 Q 1 1 a 112 ¢ Explicitly model the covariation between traits l Traits may be correlated due to shared genetic factors (A) or shared environmental factors (C or E) 1 1 A Q 2 2 q 112 a 222 ADHD IQ e 112 1 q 222 e 222 1 E 2 r. E Dingo

Explicitly model the covariation between traits… Directly assess pleiotropy ¢ Increased power ¢ l Esp. when the pattern of QTL effects is different from the background covariation • ie positive r but gene effects are in opposite directions ¢ Reduced multiple testing Drop Bear

Multivariate analysis… ¢ In the context of linkage analysis when we have family data two traits measured in twin pairs l Interested in: l • Cross-trait covariance within individuals • Cross-trait covariance between twins • MZ: DZ ratio of cross-trait covariance between twins Echidna

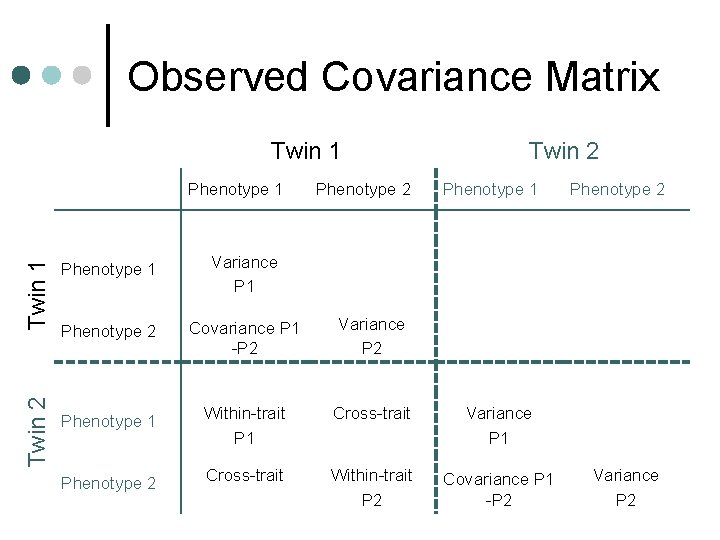
Observed Covariance Matrix Twin 1 Twin 2 Twin 1 Phenotype 2 Twin 2 Phenotype 1 Variance P 1 Phenotype 2 Covariance P 1 -P 2 Variance P 2 Phenotype 1 Within-trait P 1 Cross-trait Variance P 1 Phenotype 2 Cross-trait Within-trait P 2 Covariance P 1 -P 2 Phenotype 2 Variance P 2

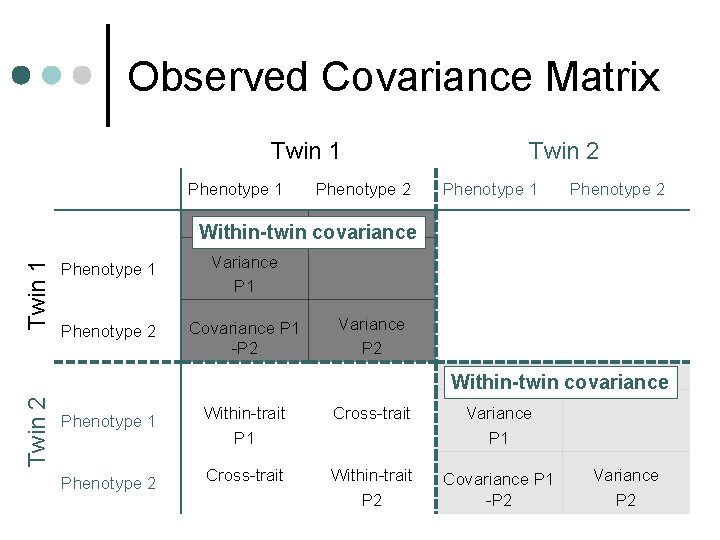
Observed Covariance Matrix Twin 1 Phenotype 2 Twin 2 Phenotype 1 Phenotype 2 Twin 1 Within-twin covariance Phenotype 1 Variance P 1 Phenotype 2 Covariance P 1 -P 2 Variance P 2 Twin 2 Within-twin covariance Phenotype 1 Within-trait P 1 Cross-trait Variance P 1 Phenotype 2 Cross-trait Within-trait P 2 Covariance P 1 -P 2 Variance P 2

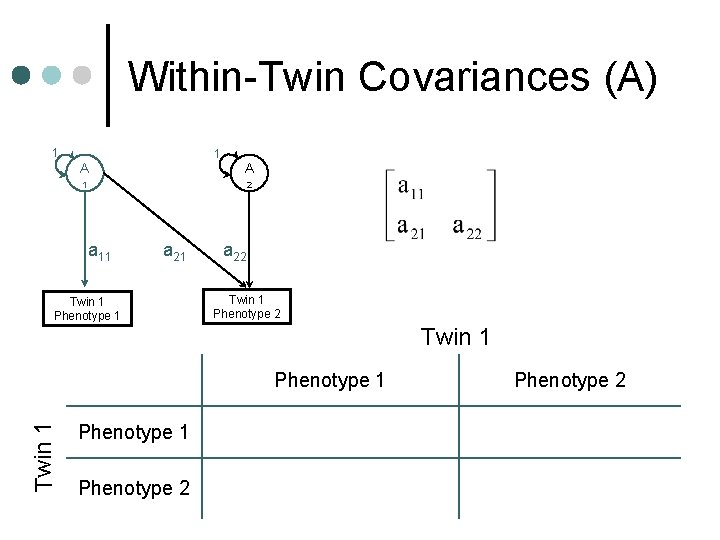
Within-Twin Covariances (A) 1 1 A A 1 2 a 11 a 21 Twin 1 Phenotype 1 a 22 Twin 1 Phenotype 1 a 112+c 112+e 112 Phenotype 2 a 11 a 21+c 11 c 21+e 11 e 21 Phenotype 2 a 222+a 212+c 222+c 212+e 2 2 22 +e 21

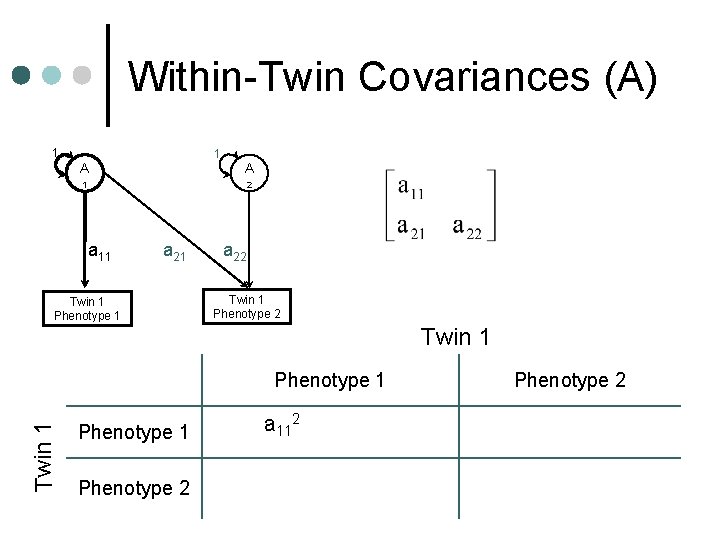
Within-Twin Covariances (A) 1 1 A A 1 2 a 11 a 21 Twin 1 Phenotype 1 a 22 Twin 1 Phenotype 1 a 112+c 112+e 112 Phenotype 2 a 11 a 21+c 11 c 21+e 11 e 21 Phenotype 2 a 222+a 212+c 222+c 212+e 2 2 22 +e 21

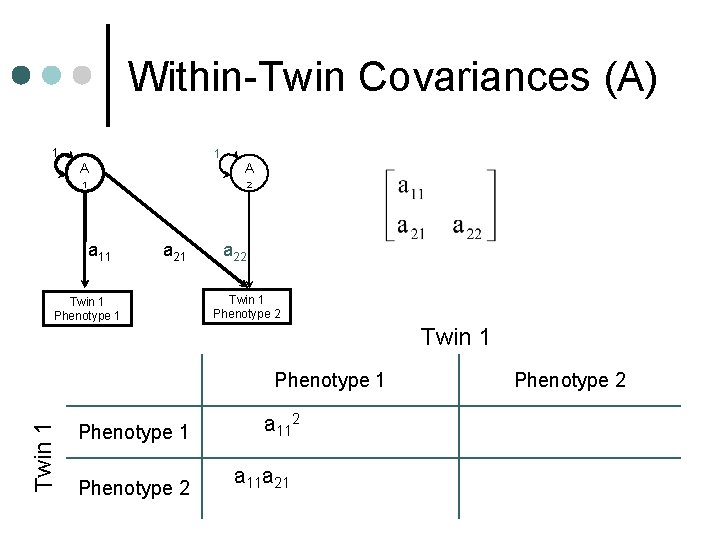
Within-Twin Covariances (A) 1 1 A A 1 2 a 11 a 21 Twin 1 Phenotype 1 a 22 Twin 1 Phenotype 1 a 112+c 112+e 112 Phenotype 2 a 11 a 21+c 11 c 21+e 11 e 21 Phenotype 2 a 222+a 212+c 222+c 212+e 2 2 22 +e 21

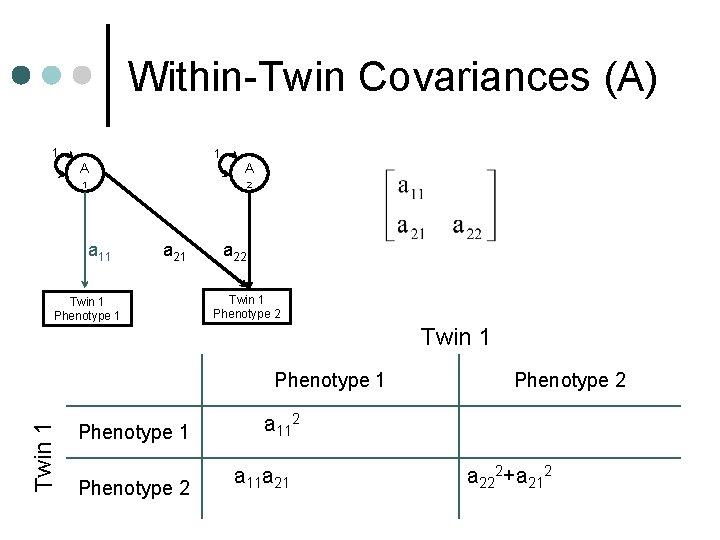
Within-Twin Covariances (A) 1 1 A A 1 2 a 11 a 21 Twin 1 Phenotype 1 a 22 Twin 1 Phenotype 1 a 112+c 112+e 112 Phenotype 2 a 11 a 21+c 11 c 21+e 11 e 21 Phenotype 2 a 222+a 212+c 222+c 212+e 2 2 22 +e 21

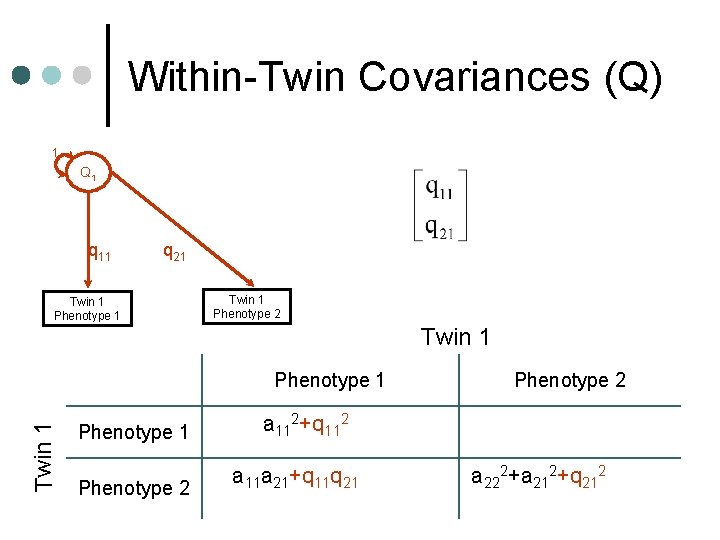
Within-Twin Covariances (Q) 1 Q 1 q 11 q 21 Twin 1 Phenotype 2 Twin 1 Phenotype 1 a 112+q 112+e 112 Phenotype 2 a 11 a 21+q 11 q 21+e 11 e 21 Phenotype 2 a 222+a 212+q 212+e 222+ e 212

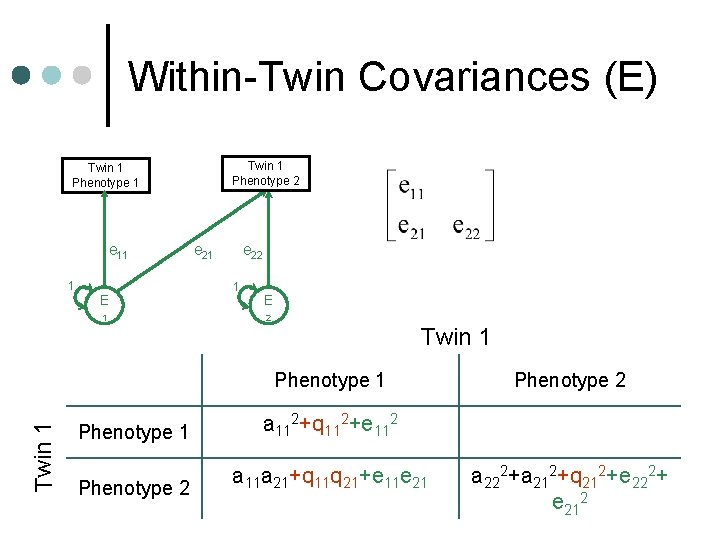
Within-Twin Covariances (E) Twin 1 Phenotype 2 Twin 1 Phenotype 1 e 11 1 E 1 e 22 1 E 2 Twin 1 Phenotype 1 a 112+q 112+e 112 Phenotype 2 a 11 a 21+q 11 q 21+e 11 e 21 Phenotype 2 a 222+a 212+q 212+e 222+ e 212

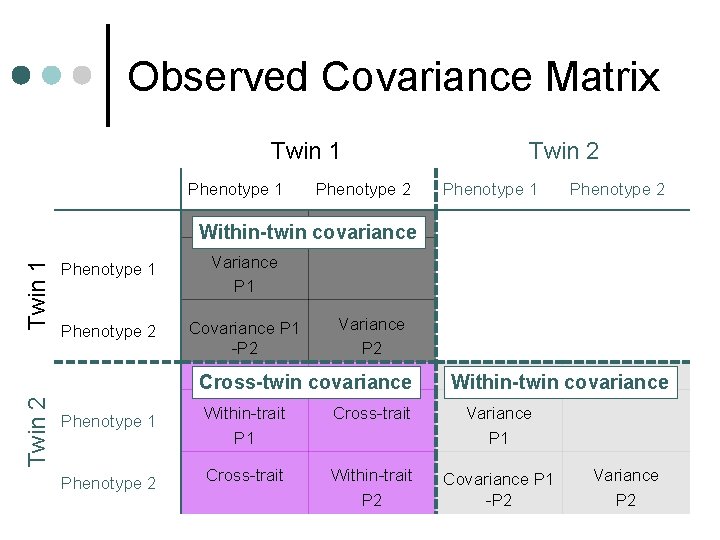
Observed Covariance Matrix Twin 1 Phenotype 2 Twin 2 Phenotype 1 Phenotype 2 Twin 1 Within-twin covariance Phenotype 1 Variance P 1 Phenotype 2 Covariance P 1 -P 2 Variance P 2 Twin 2 Cross-twin covariance Within-twin covariance Phenotype 1 Within-trait P 1 Cross-trait Variance P 1 Phenotype 2 Cross-trait Within-trait P 2 Covariance P 1 -P 2 Variance P 2

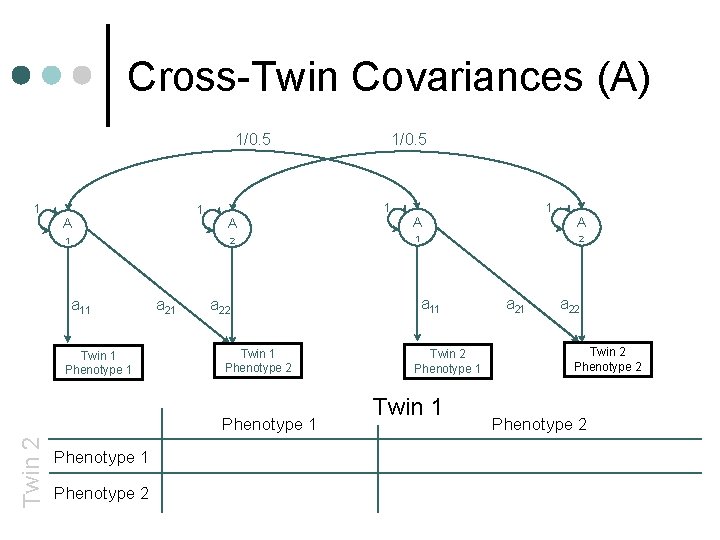
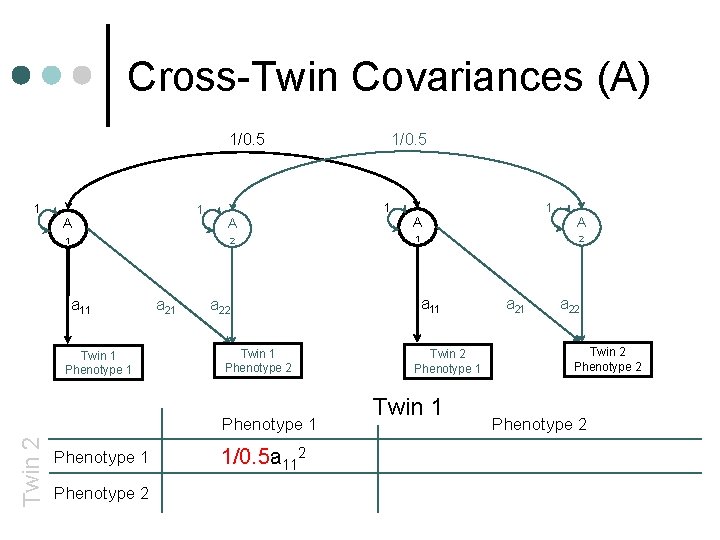
Cross-Twin Covariances (A) 1/0. 5 1 1 A 1 a 11 Twin 1 Phenotype 1 a 21 1 A A 2 1 Twin 1 Phenotype 2 1 A 2 a 11 a 22 Phenotype 1 Twin 2 1/0. 5 Twin 2 Phenotype 1 Twin 1 a 22 Twin 2 Phenotype 1 Phenotype 2 +c 11 c 21 +c 222+c 212

Cross-Twin Covariances (A) 1/0. 5 1 1 A 1 a 11 Twin 1 Phenotype 1 a 21 1 A A 2 1 Twin 1 Phenotype 2 Phenotype 1 1/0. 5 a 112+ Phenotype 2 +c 11 c 21 1 A 2 a 11 a 22 Phenotype 1 Twin 2 1/0. 5 Twin 2 Phenotype 1 Twin 1 a 22 Twin 2 Phenotype 2 +c 222+c 212

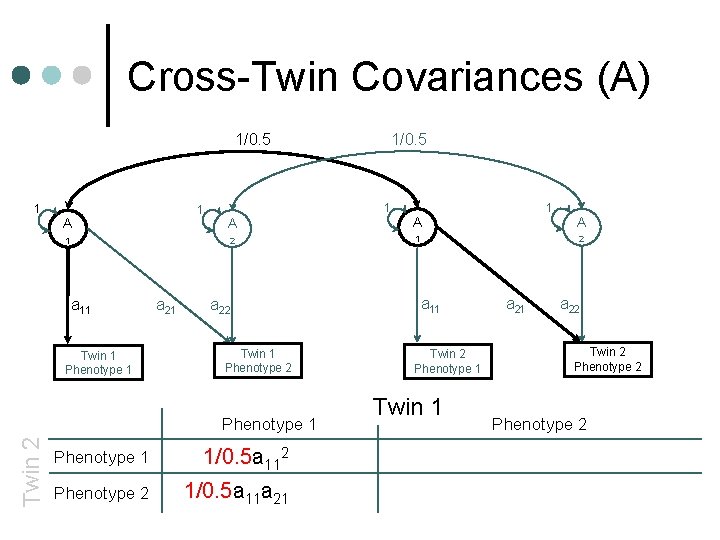
Cross-Twin Covariances (A) 1/0. 5 1 1 A 1 a 11 Twin 1 Phenotype 1 a 21 1 A A 2 1 a 22 Twin 1 Phenotype 2 Phenotype 1 Twin 2 1/0. 5 Phenotype 1 1/0. 5 a 112+c 112 Phenotype 2 1/0. 5 a 11 a 21+c 11 c 21 1 A 2 a 11 Twin 2 Phenotype 1 Twin 1 a 22 Twin 2 Phenotype 2 +c 222+c 212

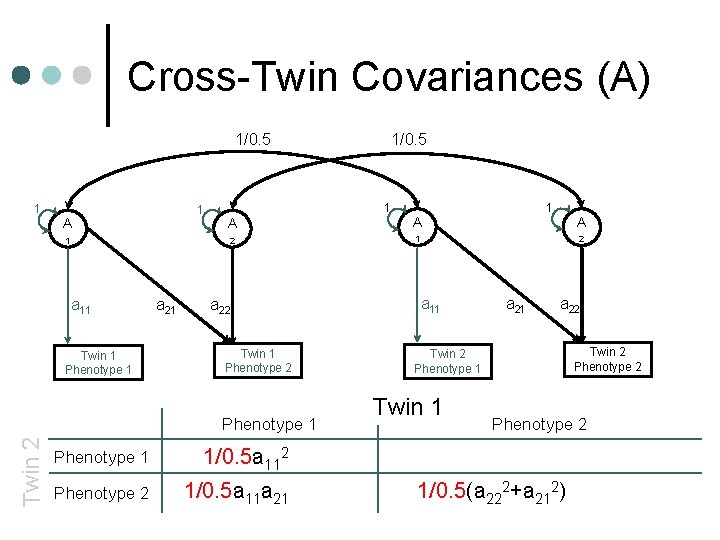
Cross-Twin Covariances (A) 1/0. 5 1 1 A 1 a 11 Twin 1 Phenotype 1 a 21 1 A A 2 1 a 22 Twin 1 Phenotype 2 Phenotype 1 Twin 2 1/0. 5 Phenotype 1 1/0. 5 a 112+c 112 Phenotype 2 1/0. 5 a 11 a 21+c 11 c 21 1 A 2 a 11 Twin 2 Phenotype 1 Twin 1 a 22 Twin 2 Phenotype 2 1/0. 5(a 222+a 212)+c 222+c 212

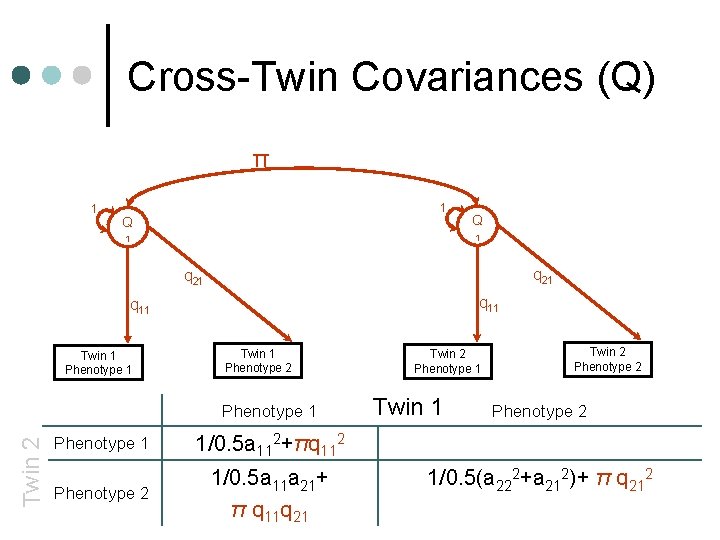
Cross-Twin Covariances (Q) π 1 1 Q Q 1 1 q 21 q 11 Twin 1 Phenotype 2 Twin 2 Phenotype 1 1/0. 5 a 112+πq 112 Phenotype 2 1/0. 5 a 11 a 21+ π q 11 q 21 Twin 2 Phenotype 2 1/0. 5(a 222+a 212)+ π q 212

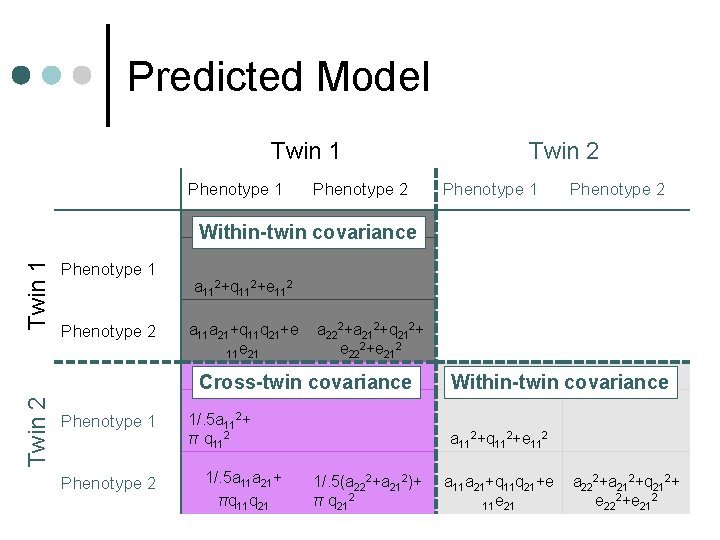
Predicted Model Twin 1 Phenotype 2 Twin 2 Phenotype 1 Phenotype 2 Twin 1 Within-twin covariance Phenotype 1 Phenotype 2 a 112+q 112+e 112 a 11 a 21+q 11 q 21+e 11 e 21 a 222+a 212+q 212+ e 222+e 212 Twin 2 Cross-twin covariance Phenotype 1 Phenotype 2 1/. 5 a 112+ π q 112 1/. 5 a 11 a 21+ πq 11 q 21 Within-twin covariance a 112+q 112+e 112 1/. 5(a 222+a 212)+ π q 212 a 11 a 21+q 11 q 21+e 11 e 21 a 222+a 212+q 212+ e 222+e 212

Running MV linkage analysis… ¢ Numerous programs l l Mx Solar Loki Merlin • Repeated measures ¢ ¢ Computationally intensive Multiple boundary issues l p-values difficult to obtain Emu (not e-moo)

MV association… ¢ Maximum likelihood – factor based approach l ¢ Canonical Correlation approach l ¢ Mx Plink Principal components approach l F-bat Funnel Web Spider

Maximum likelihood approach ¢ Unrelated individuals Shared variance due to a common factor l Residual non-shared variance l ¢ Family based data l ACE type models Galah

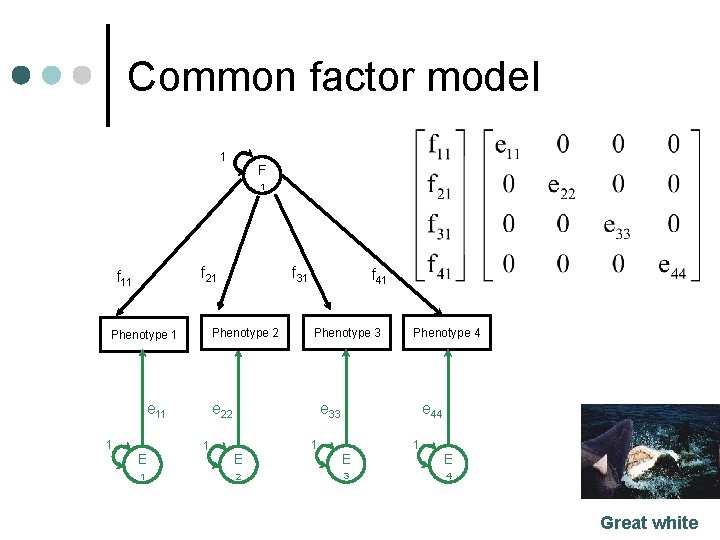
Common factor model 1 F 1 f 21 f 11 Phenotype 2 Phenotype 1 e 11 1 E 1 f 31 f 41 Phenotype 3 e 33 e 22 1 Phenotype 4 E 2 1 e 44 E 3 1 E 4 Great white

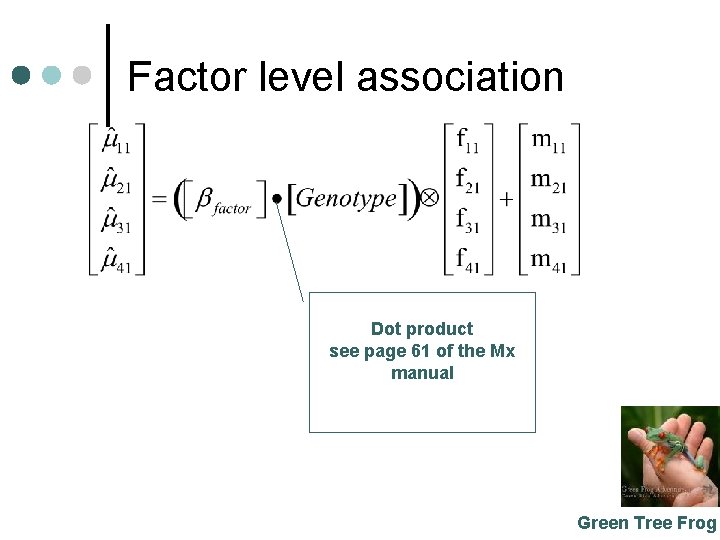
Factor level association ¢ ¢ Estimate a factor level beta Use the factor loadings as weights Add the uncorrected or grand mean 1 df Green Tree Frog

Factor level association Dot product see page 61 of the Mx manual Green Tree Frog

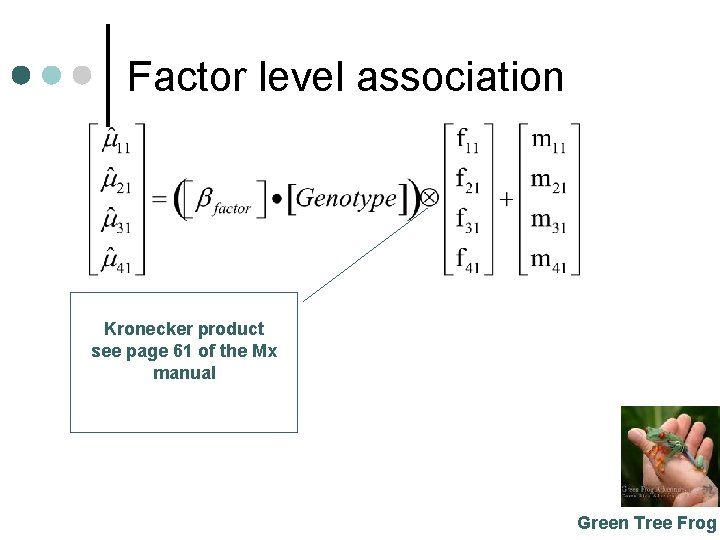
Factor level association Kronecker product see page 61 of the Mx manual Green Tree Frog

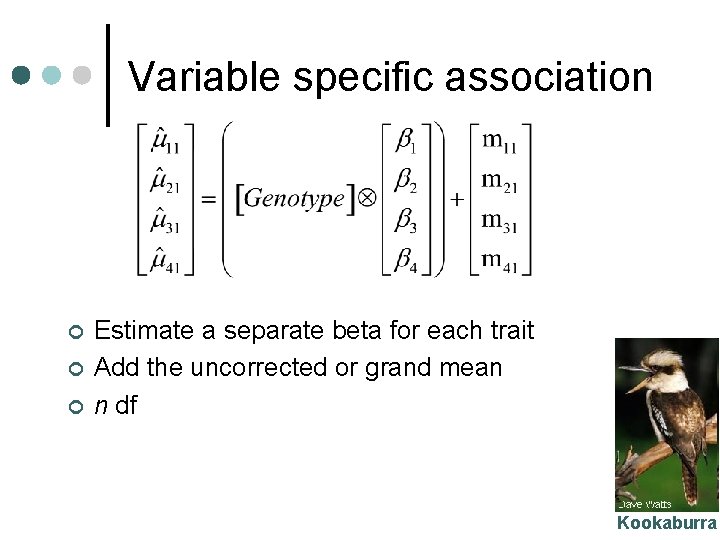
Variable specific association ¢ ¢ ¢ Estimate a separate beta for each trait Add the uncorrected or grand mean n df Kookaburra

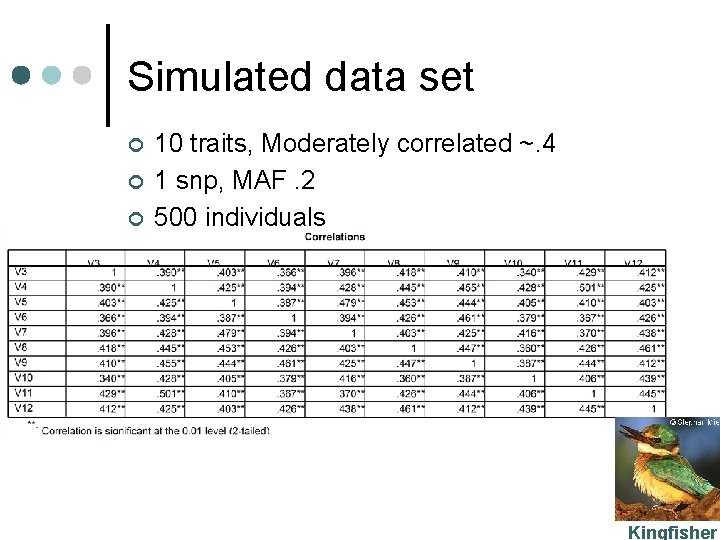
Simulated data set ¢ ¢ ¢ 10 traits, Moderately correlated ~. 4 1 snp, MAF. 2 500 individuals Kingfisher

factorlevel. mx… dataset. ped Example Factor level association script - Boulder 2009 Sarah Medland Data NI=15 NGroups=1 Rec file=dataset. ped Labels fid iid a 1 a 2 genotype V 1 V 2 V 3 V 4 V 5 V 6 V 7 V 8 V 9 V 10 Select genotype V 1 V 2 V 3 V 4 V 5 V 6 V 7 V 8 V 9 V 10 ; Definition genotype ; Begin Matrices; F full 10 1 free R diag 10 10 free M full 10 1 free B full 1 1 free G full 1 1 End Matrices; ! factor ! residuals ! grand means ! association beta ! genotype Numbat

factorlevel. mx… dataset. ped st. 6 F 1 1 to F 10 1 st. 4 R 1 1 to R 10 10 st 0 M 1 1 to M 10 1 sp G genotype Covariance (F*F')+(R*R') ; Means M + (B. G)@F ; Options multiple issat jiggle end drop B 1 1 1 end Perentie Monitor

variablespecific. mx… dataset. ped Example Factor level association script - Boulder 2009 Sarah Medland Data NI=15 NGroups=1 Rec file=dataset. ped Labels fid iid a 1 a 2 genotype V 1 V 2 V 3 V 4 V 5 V 6 V 7 V 8 V 9 V 10 Select genotype V 1 V 2 V 3 V 4 V 5 V 6 V 7 V 8 V 9 V 10 ; Definition genotype ; Begin Matrices; F full 10 1 free R diag 10 10 free M full 10 1 free B full 10 1 free G full 1 1 End Matrices; ! factor ! residuals ! grand means ! association beta ! genotype Quoll

variablespecific. mx… dataset. ped st. 6 F 1 1 to F 10 1 st. 4 R 1 1 to R 10 10 st 0 M 1 1 to M 10 1 sp G genotype Covariance (F*F')+(R*R') ; Means M + (G@B). F ; Options multiple issat jiggle end drop B 1 1 1 to B 1 10 1 end Red back spider

Your task ¢ Run both FL and VS tests in Mx for the first data set Edit the data file name l Calculate the p-value for the VS test using excel… l Which variables are associated? ¢ What is the mean for variable 1 by genotype? ¢

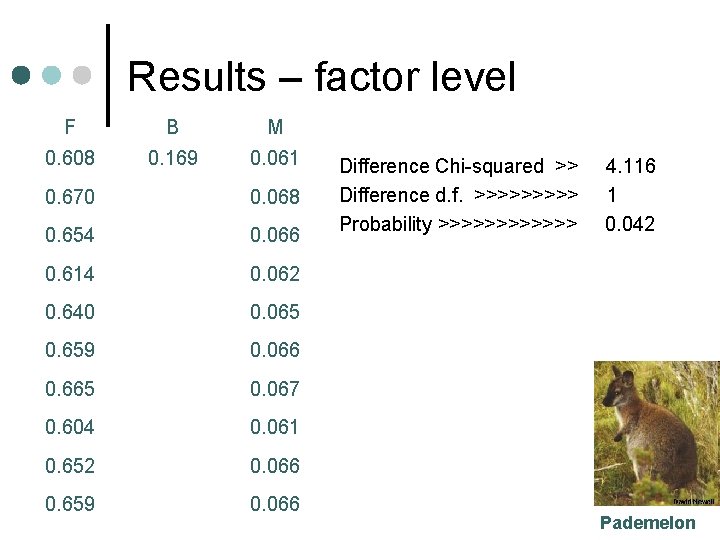
Results – factor level F B M 0. 608 0. 169 0. 061 0. 670 0. 068 0. 654 0. 066 0. 614 0. 062 0. 640 0. 065 0. 659 0. 066 0. 665 0. 067 0. 604 0. 061 0. 652 0. 066 0. 659 0. 066 Difference Chi-squared >> Difference d. f. >>>>> Probability >>>>>> 4. 116 1 0. 042 Pademelon

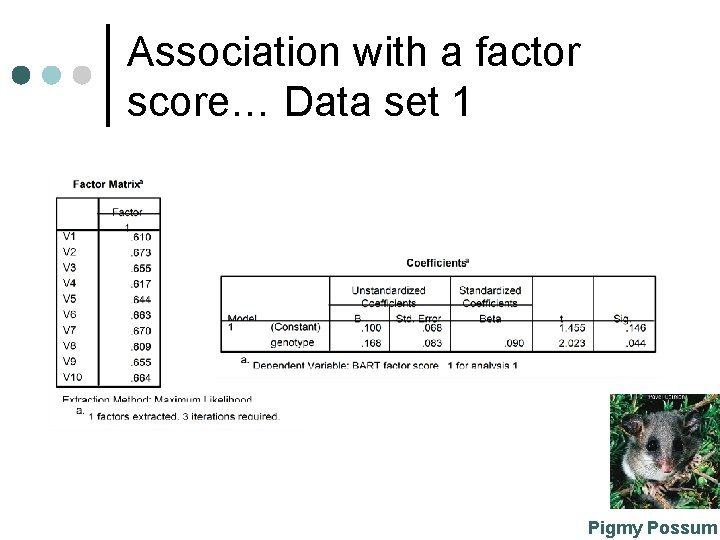
Association with a factor score… Data set 1 Pigmy Possum

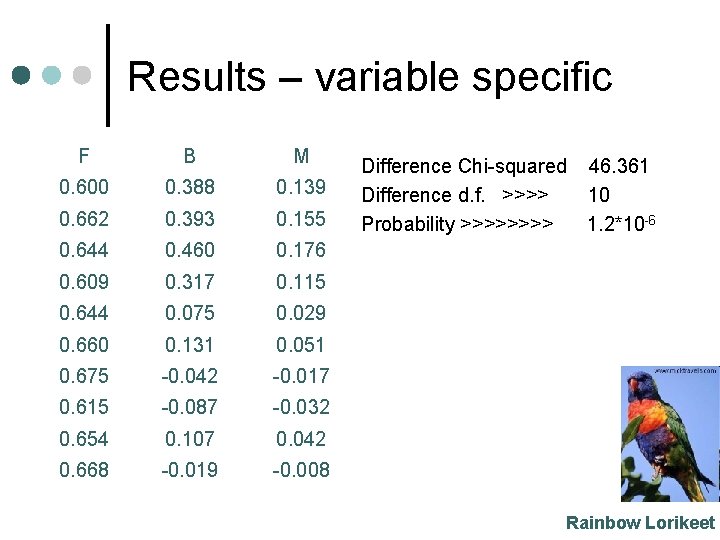
Results – variable specific F B M 0. 600 0. 388 0. 139 0. 662 0. 393 0. 155 0. 644 0. 460 0. 176 0. 609 0. 317 0. 115 0. 644 0. 075 0. 029 0. 660 0. 131 0. 051 0. 675 -0. 042 -0. 017 0. 615 -0. 087 -0. 032 0. 654 0. 107 0. 042 0. 668 -0. 019 -0. 008 Difference Chi-squared Difference d. f. >>>> Probability >>>> 46. 361 10 1. 2*10 -6 Rainbow Lorikeet

Mean under different genotypes -1 AA 0 AB 1 BB -0. 249 0. 139 0. 527 Salt water croc

Advantages to the ML approach ¢ Completely flexible l Can be applied to any model • Longitudinal models • Simplex/Autoregressive processed Easy to add dominance etc l Covariates l Extends to family data l Stone fish

Disadvantages Correction for multiple testing? ¢ FL & VS tests provide complementary information ¢ Inflation of type 1 error ¢ l use a Bonferroni correction if you use both Taipan

Tiger Snake Wombat Tassie Tiger Tree Kangaroo Thorny Devil Zebra finch Tassie Devil