f MRI Modelling Statistical Inference Guillaume Flandin Wellcome






![Contrasts [1 0 0 0 0] [0 1 -1 0 0 0] Contrasts [1 0 0 0 0] [0 1 -1 0 0 0]](https://slidetodoc.com/presentation_image_h/0b6c2cf62fe403a899848ef86dadfd20/image-22.jpg)



![Scaling issue Subject 1 [1 1 1 1 ] / 4 q The T-statistic Scaling issue Subject 1 [1 1 1 1 ] / 4 q The T-statistic](https://slidetodoc.com/presentation_image_h/0b6c2cf62fe403a899848ef86dadfd20/image-26.jpg)




- Slides: 41

f. MRI Modelling & Statistical Inference Guillaume Flandin Wellcome Trust Centre for Neuroimaging University College London SPM Course Chicago, 22 -23 Oct 2015

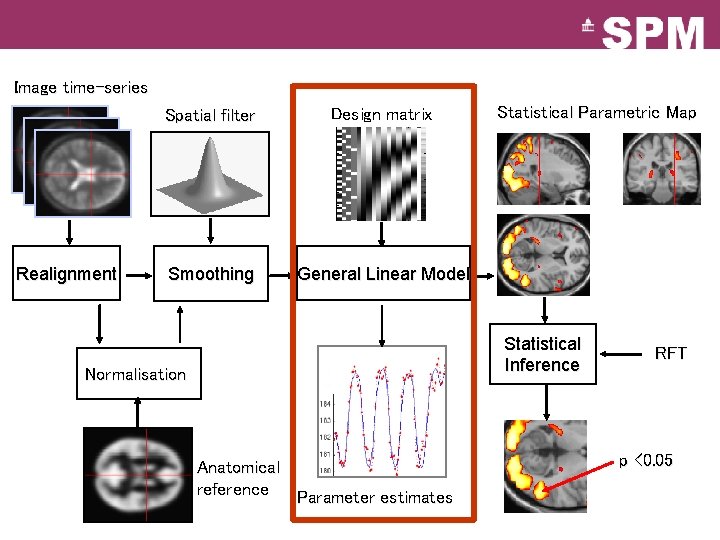
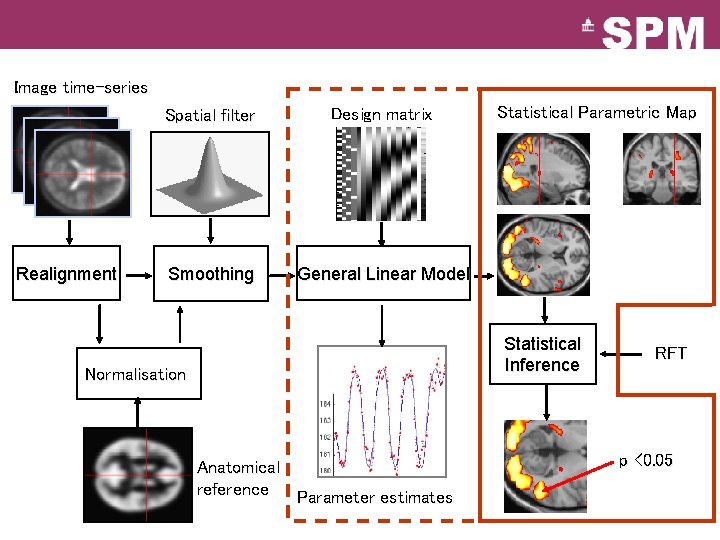
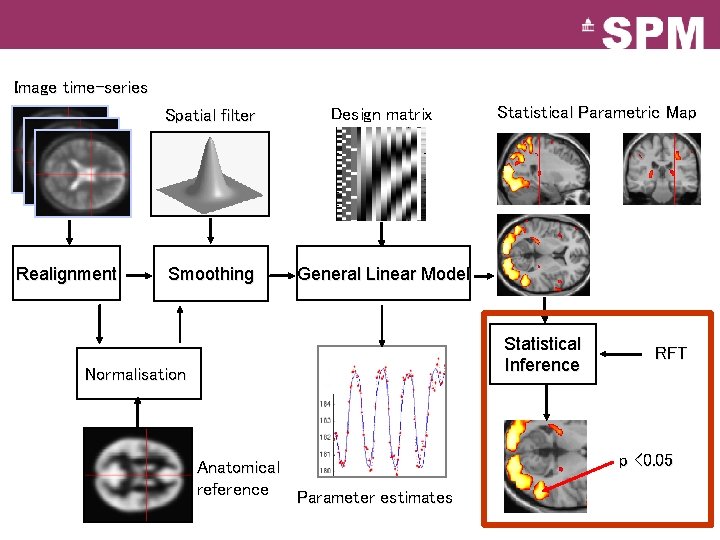
Image time-series Realignment Spatial filter Design matrix Smoothing General Linear Model Statistical Parametric Map Statistical Inference Normalisation Anatomical reference Parameter estimates RFT p <0. 05

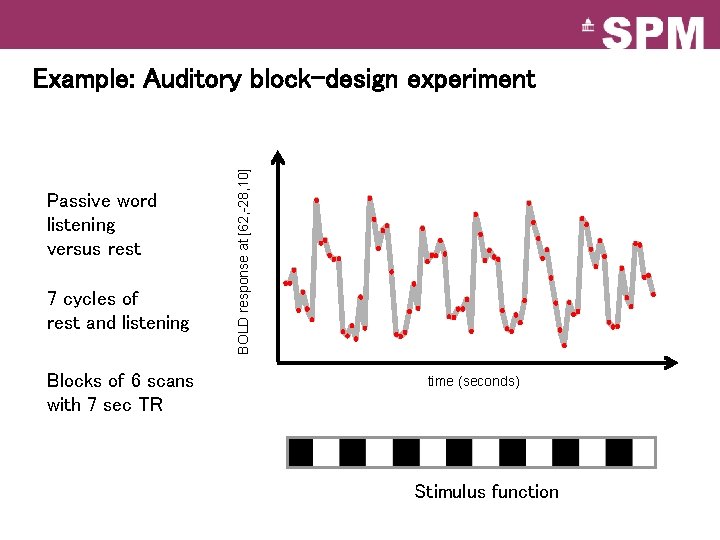
Passive word listening versus rest 7 cycles of rest and listening Blocks of 6 scans with 7 sec TR BOLD response at [62, -28, 10] Example: Auditory block-design experiment time (seconds) Stimulus function

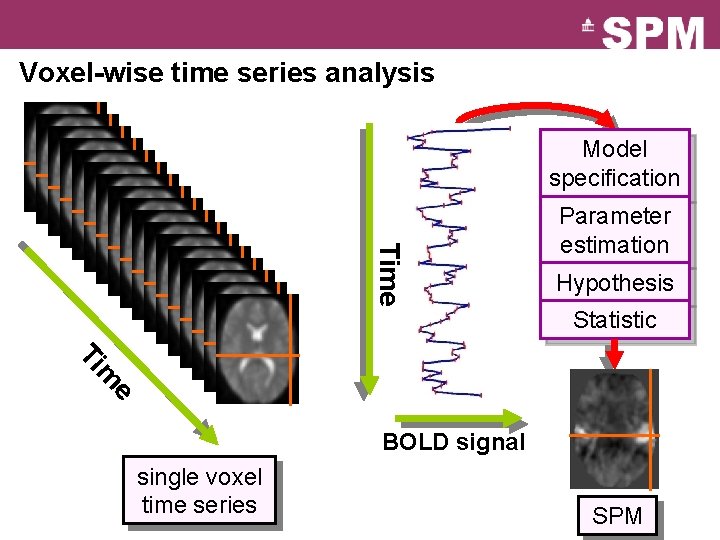
Voxel-wise time series analysis Model specification Time Parameter estimation Hypothesis Statistic e m Ti BOLD signal single voxel time series SPM

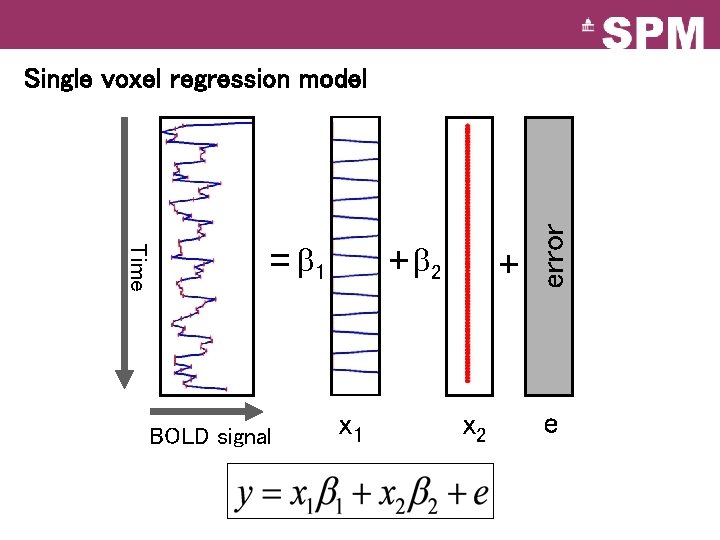
Time = 1 BOLD signal + 2 x 1 + x 2 error Single voxel regression model e

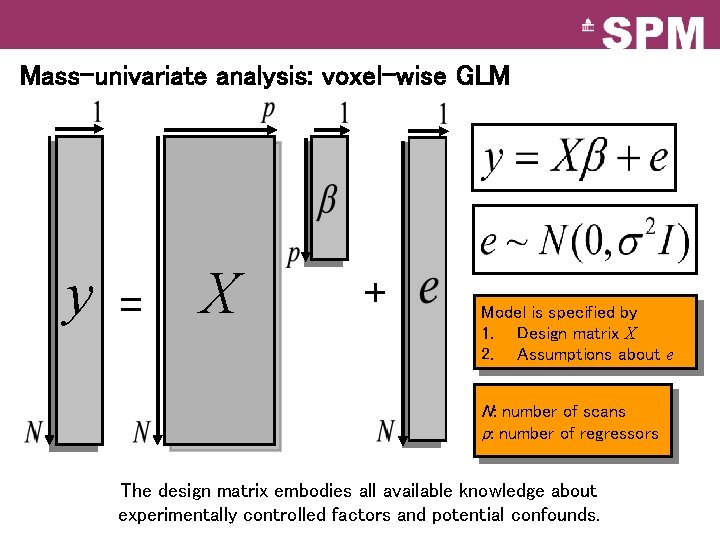
Mass-univariate analysis: voxel-wise GLM y = X + Model is specified by 1. Design matrix X 2. Assumptions about e N: number of scans p: number of regressors The design matrix embodies all available knowledge about experimentally controlled factors and potential confounds.

GLM: a flexible framework for parametric analyses • • one sample t-test two sample t-test paired t-test Analysis of Variance (ANOVA) Analysis of Covariance (ANCo. VA) correlation linear regression multiple regression

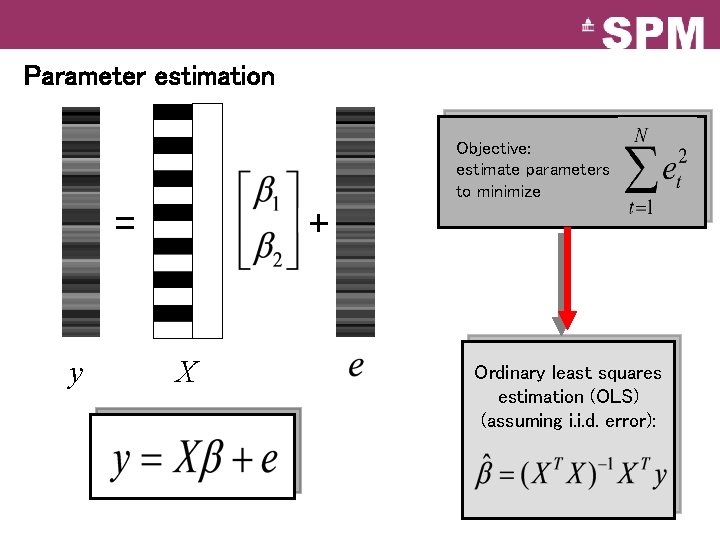
Parameter estimation Objective: estimate parameters to minimize = y + X Ordinary least squares estimation (OLS) (assuming i. i. d. error):

Problems of this model with f. MRI time series 1. The BOLD response has a delayed and dispersed shape.

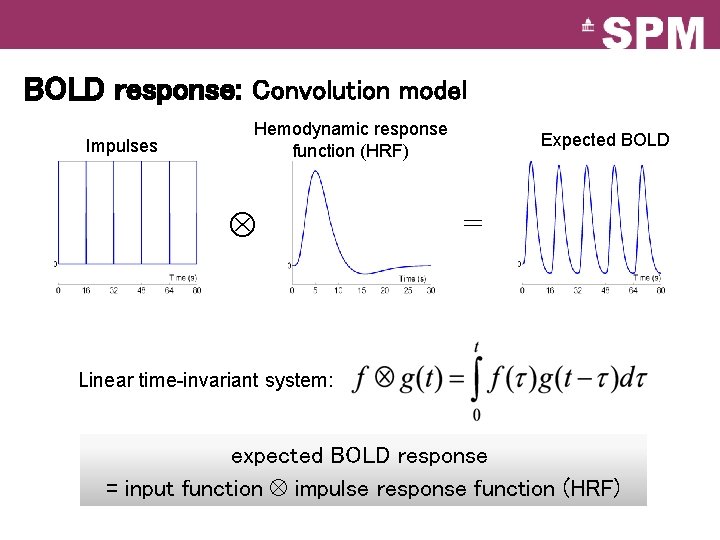
BOLD response: Convolution model Impulses Hemodynamic response function (HRF) Expected BOLD = Linear time-invariant system: expected BOLD response = input function impulse response function (HRF)

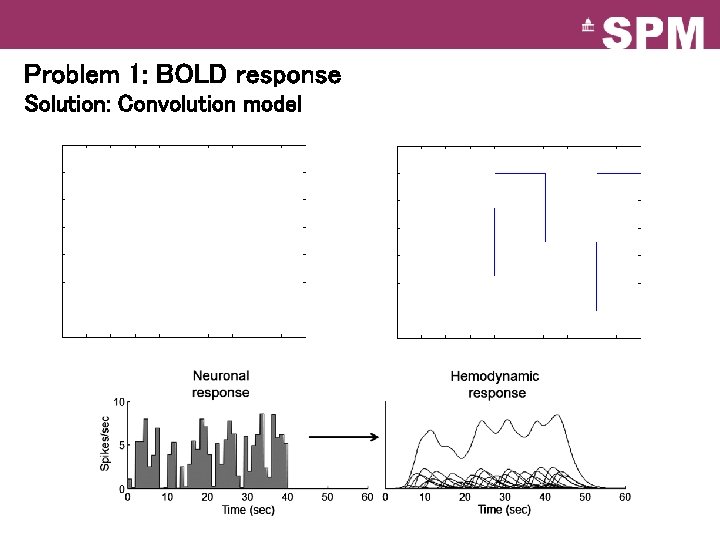
Problem 1: BOLD response Solution: Convolution model

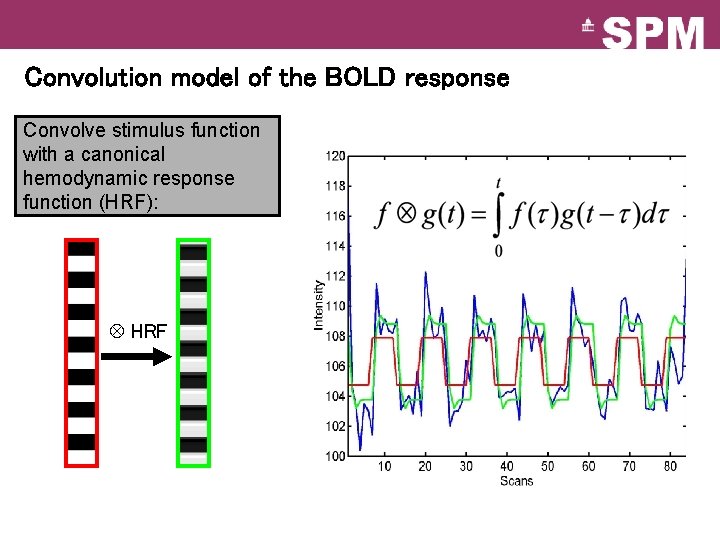
Convolution model of the BOLD response Convolve stimulus function with a canonical hemodynamic response function (HRF): HRF

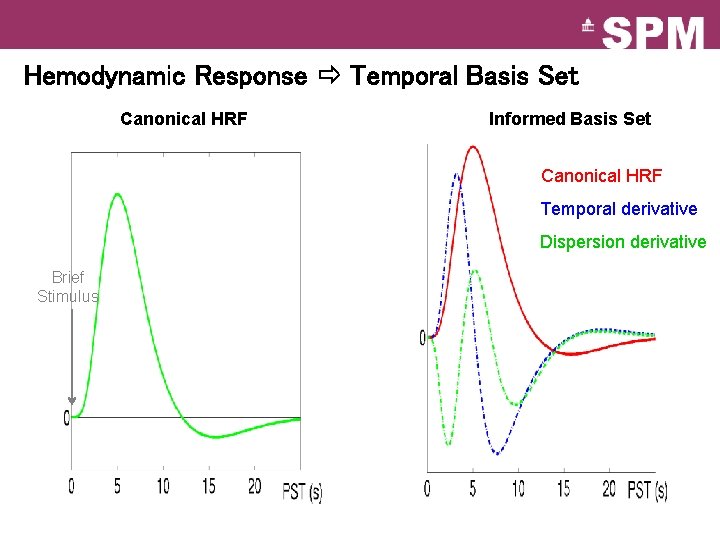
Hemodynamic Response Temporal Basis Set Canonical HRF Informed Basis Set Canonical HRF Temporal derivative Dispersion derivative Brief Stimulus

Problems of this model with f. MRI time series 1. The BOLD response has a delayed and dispersed shape. 2. The BOLD signal includes substantial amounts of lowfrequency noise (eg due to scanner drift).

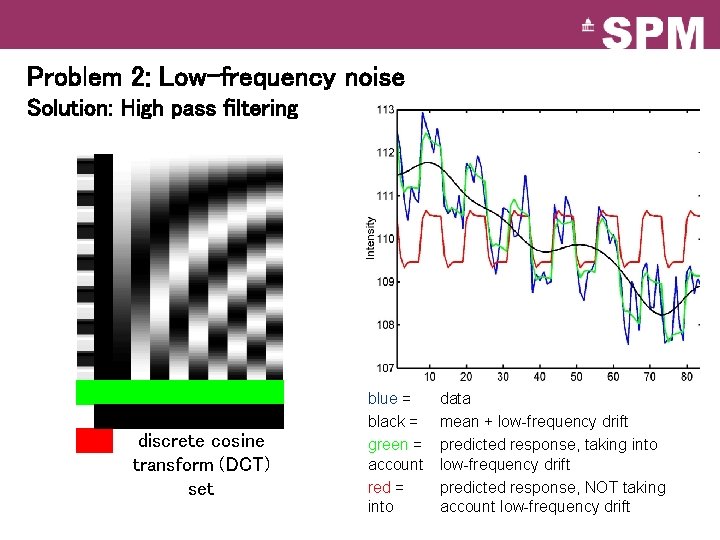
Problem 2: Low-frequency noise Solution: High pass filtering discrete cosine transform (DCT) set blue = black = green = account red = into data mean + low-frequency drift predicted response, taking into low-frequency drift predicted response, NOT taking account low-frequency drift

Problems of this model with f. MRI time series 1. The BOLD response has a delayed and dispersed shape. 2. The BOLD signal includes substantial amounts of lowfrequency noise (eg due to scanner drift). 3. Due to breathing, heartbeat & unmodeled neuronal activity, the errors are serially correlated. This violates the assumptions of the noise model in the GLM.

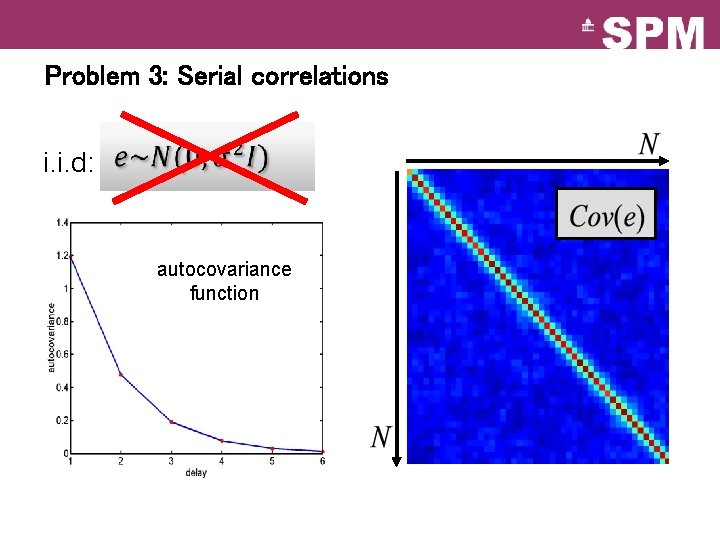
Problem 3: Serial correlations i. i. d: autocovariance function

Multiple covariance components enhanced noise model at voxel i error covariance components Q and hyperparameters V = 1 Q 1 + 2 Q 2 Estimation of hyperparameters with Re. ML (Restricted Maximum Likelihood).

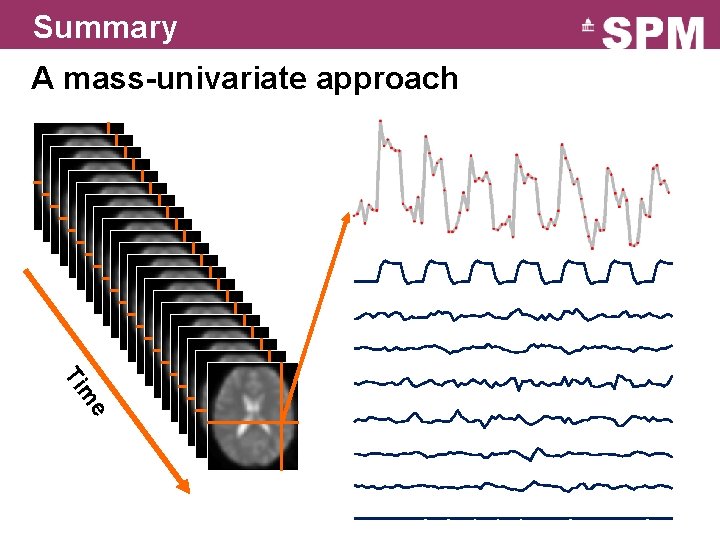
Summary A mass-univariate approach e m Ti

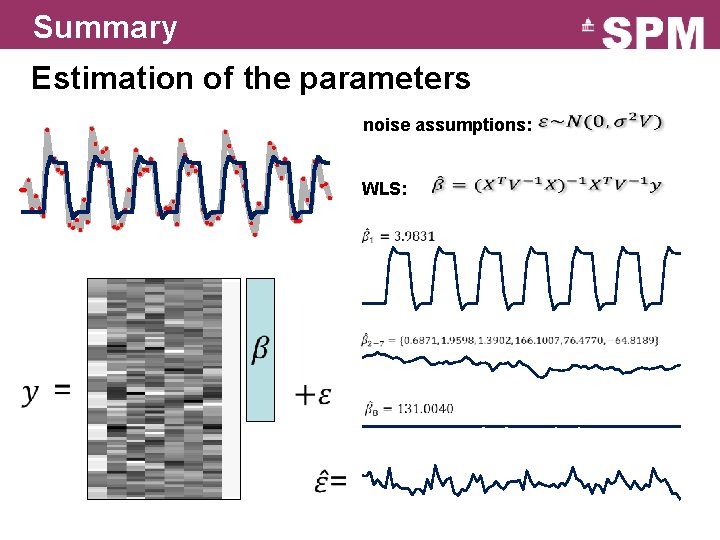
Summary Estimation of the parameters noise assumptions: WLS:

Image time-series Realignment Spatial filter Design matrix Smoothing General Linear Model Statistical Parametric Map Statistical Inference Normalisation Anatomical reference Parameter estimates RFT p <0. 05
![Contrasts 1 0 0 0 0 0 1 1 0 0 0 Contrasts [1 0 0 0 0] [0 1 -1 0 0 0]](https://slidetodoc.com/presentation_image_h/0b6c2cf62fe403a899848ef86dadfd20/image-22.jpg)
Contrasts [1 0 0 0 0] [0 1 -1 0 0 0]

Hypothesis Testing To test an hypothesis, we construct “test statistics”. q Null Hypothesis H 0 Typically what we want to disprove (no effect). The Alternative Hypothesis HA expresses outcome of interest. q Test Statistic T The test statistic summarises evidence about H 0. Typically, test statistic is small in magnitude when the hypothesis H 0 is true and large when false. We need to know the distribution of T under the null hypothesis. Null Distribution of T

Hypothesis Testing u q Significance level α: Acceptable false positive rate α. threshold uα Threshold uα controls the false positive rate Null Distribution of T q Conclusion about the hypothesis: We reject the null hypothesis in favour of the alternative hypothesis if t > uα q p-value: A p-value summarises evidence against H 0. This is the chance of observing value more extreme than t under the null hypothesis. t p-value Null Distribution of T

T-test - one dimensional contrasts – SPM{t} c. T =10000000 Question: Amplitude of cond 1 > 0 ? i. e. b 1 = c. Tb> 0 ? b 1 b 2 b 3 b 4 b 5. . . Null hypothesis: H 0: c. Tb = 0 HA: contrast of estimated parameters Test statistic: T= variance estimate
![Scaling issue Subject 1 1 1 1 1 4 q The Tstatistic Scaling issue Subject 1 [1 1 1 1 ] / 4 q The T-statistic](https://slidetodoc.com/presentation_image_h/0b6c2cf62fe403a899848ef86dadfd20/image-26.jpg)
Scaling issue Subject 1 [1 1 1 1 ] / 4 q The T-statistic does not depend on the scaling of the regressors. q The T-statistic does not depend on the scaling of the contrast. Subject 5 [1 1 1 ] / 3 q Contrast depends on scaling. Ø Be careful of the interpretation of the contrasts themselves (eg, for a second level analysis): sum ≠ average

F-test - the extra-sum-of-squares principle q Model comparison: Null Hypothesis H 0: True model is X 0 (reduced model) X 0 X 1 Test statistic: ratio of explained variability and unexplained variability (error) RSS 0 Full model ? or Reduced model? 1 = rank(X) – rank(X 0) 2 = N – rank(X)

F-test - multidimensional contrasts – SPM{F} q Tests multiple linear hypotheses: H 0: True model is X 0 X 1 (b 4 -9) H 0: b 4 = b 5 =. . . = b 9 = 0 X 0 c. T = test H 0 : c. Tb = 0 ? 0001000001000001000 000000100 000000010 00001 SPM{F 6, 322} Full model? Reduced model?

F-test: summary q F-tests can be viewed as testing for the additional variance explained by a larger model wrt a simpler (nested) model comparison. q Hypotheses: q In testing uni-dimensional contrast with an F-test, for example b 1 – b 2, the result will be the same as testing b 2 – b 1. It will be exactly the square of the t-test, testing for both positive and negative effects.

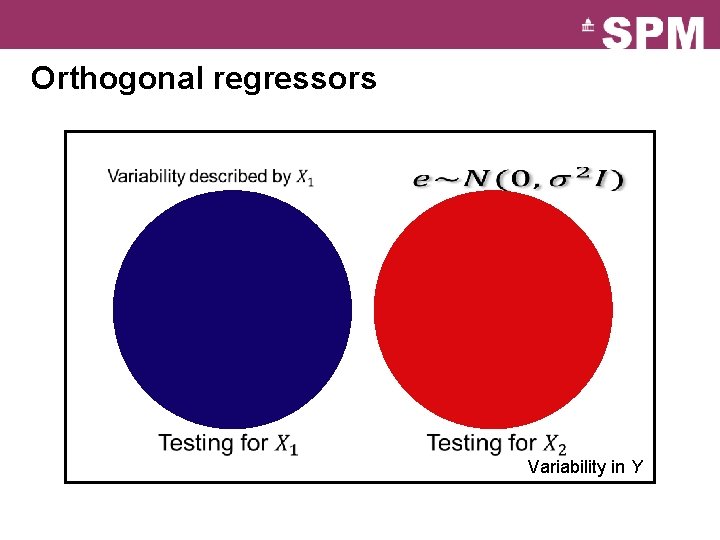
Orthogonal regressors Variability in Y

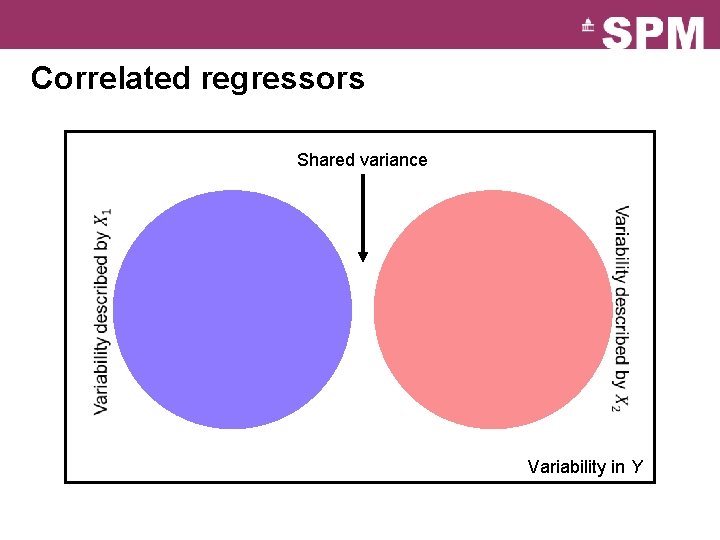
Correlated regressors Shared variance Variability in Y

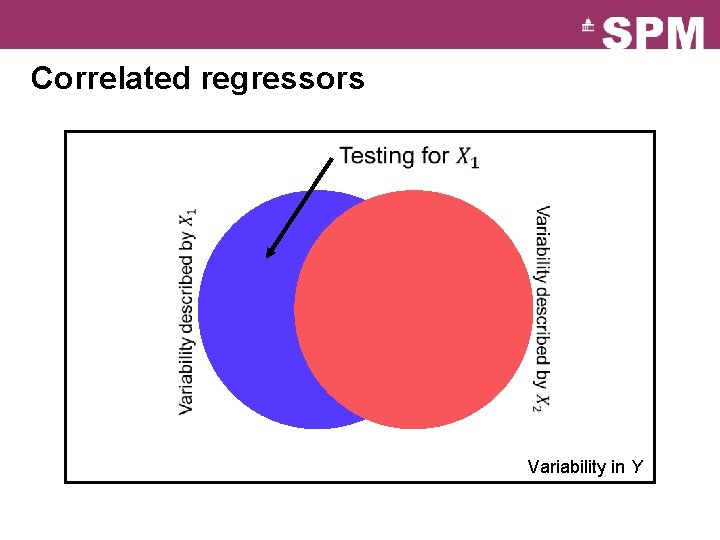
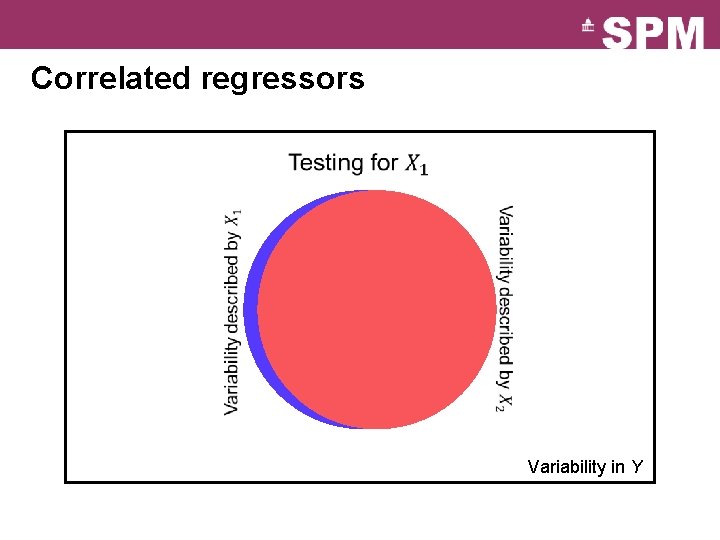
Correlated regressors Variability in Y

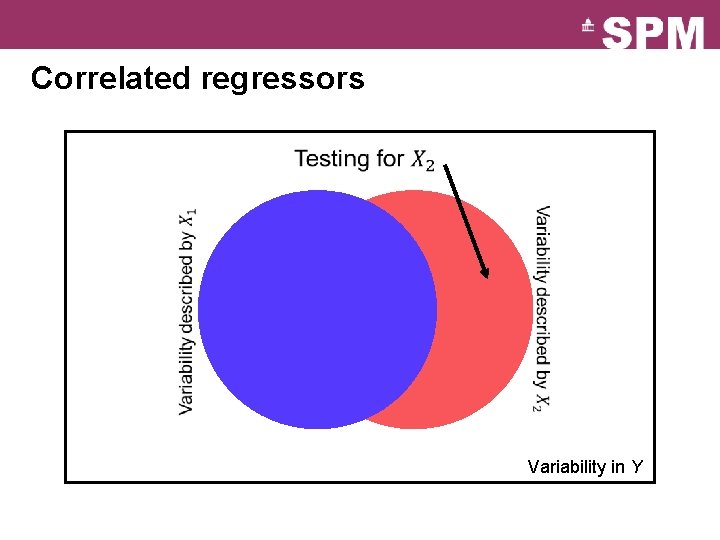
Correlated regressors Variability in Y

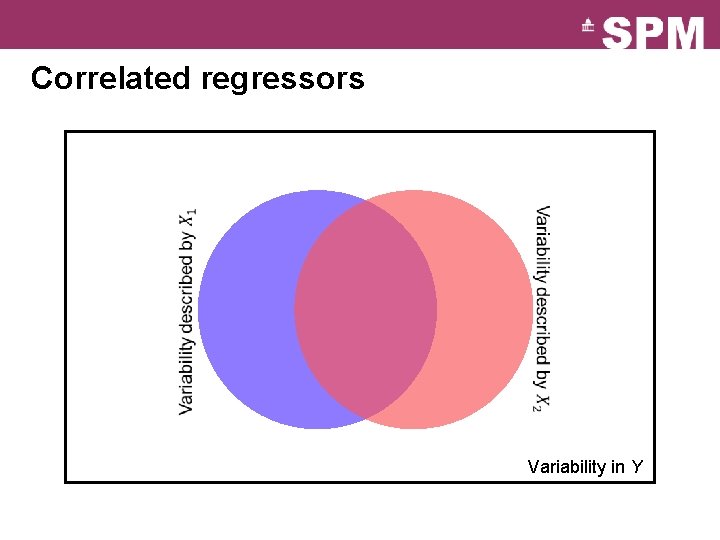
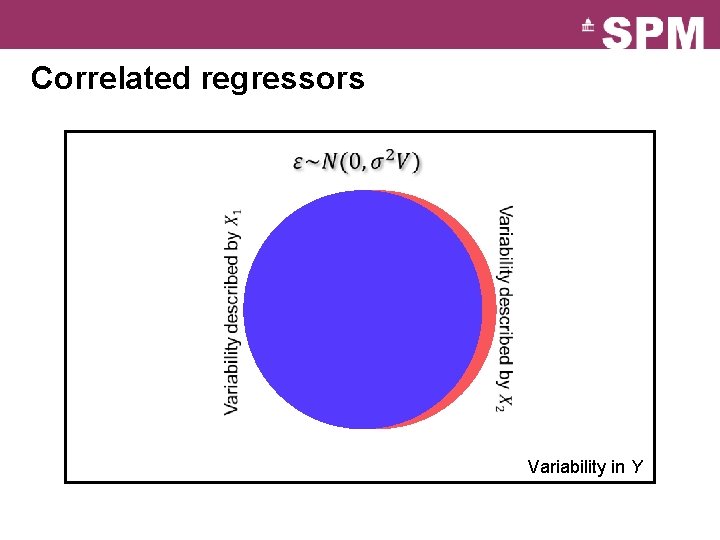
Correlated regressors Variability in Y

Correlated regressors Variability in Y

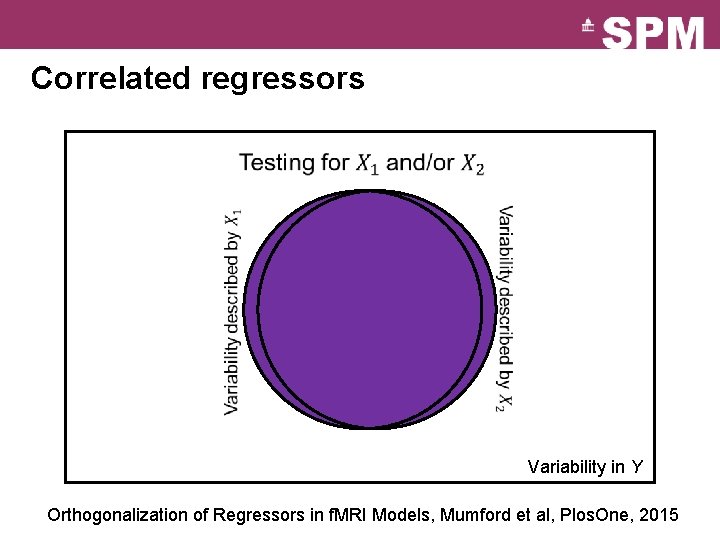
Correlated regressors Variability in Y

Correlated regressors Variability in Y Orthogonalization of Regressors in f. MRI Models, Mumford et al, Plos. One, 2015

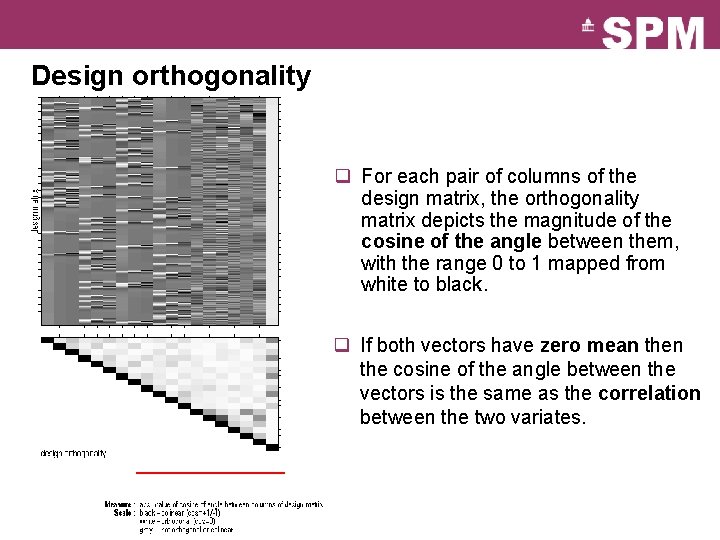
Design orthogonality q For each pair of columns of the design matrix, the orthogonality matrix depicts the magnitude of the cosine of the angle between them, with the range 0 to 1 mapped from white to black. q If both vectors have zero mean the cosine of the angle between the vectors is the same as the correlation between the two variates.

Design efficiency q The aim is to minimize the standard error of a t-contrast (i. e. the denominator of a t-statistic). q This is equivalent to maximizing the efficiency e: Noise variance Design variance q If we assume that the noise variance is independent of the specific design: q This is a relative measure: all we can really say is that one design is more efficient than another (for a given contrast).

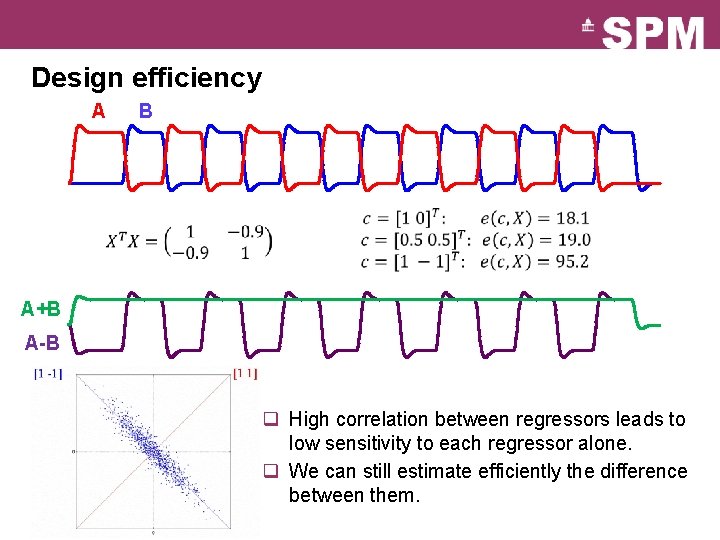
Design efficiency A B A+B A-B q High correlation between regressors leads to low sensitivity to each regressor alone. q We can still estimate efficiently the difference between them.

Image time-series Realignment Spatial filter Design matrix Smoothing General Linear Model Statistical Parametric Map Statistical Inference Normalisation Anatomical reference Parameter estimates RFT p <0. 05