Extracting binary signals from microarray timecourse data Debashis

Extracting binary signals from microarray time-course data Debashis Sahoo 1, David L. Dill 2, Rob Tibshirani 3 and Sylvia K. Plevritis 4 1 Department of Electrical Engineering 2 Department of Computer Science 3 Department of Radiology and 4 Department of Health Research and Policy and Department of Statistics Stanford University Roli Shrivastava

Introduction • Problem Statement – To identify up and down regulated gene – To identify the time of transition • Experimental Technique – Microarray (Tens of thousands of distinct probes on an array to accomplish the equivalent number of genetic tests in parallel) • Computational Technique – A tool called Step. Miner to extract biologically meaningful result from large amounts of data
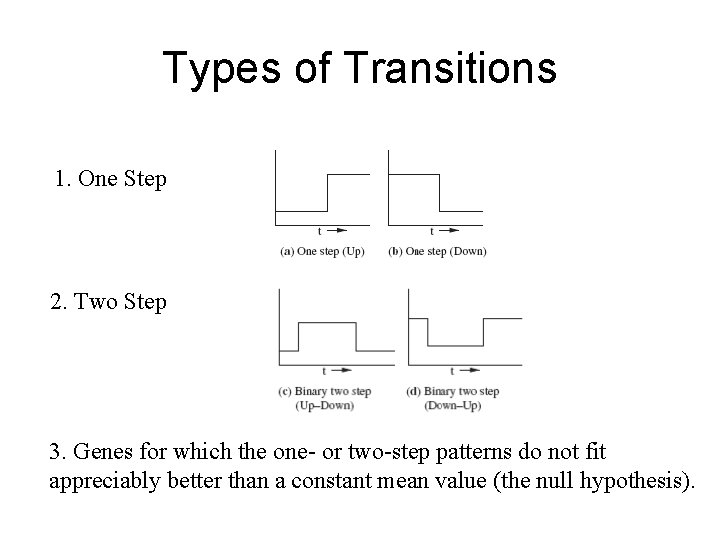
Types of Transitions 1. One Step 2. Two Step 3. Genes for which the one- or two-step patterns do not fit appreciably better than a constant mean value (the null hypothesis).

Fitting One or Two-Step Function • F 1 statistic: Computes how well the one-step model fits the data • F 2 statistic: Computes how well the two-step model fits the data • F 12 statistic: Compares the fit of one-step model and two-step model on same data • P-value: Low P-value represents a good fit of the model to the data Calculate the F statistic for the model and data set Calculate the P-value If P < Pthreshold The model fits Pthreshold = 0. 05 If P > Pthreshold The model does not fit

Step. Miner Algorithm one-step fits data AND one-step fits better than two-step fits data AND one-step does not fit it Neither one-step Nor two-step fits the data

Comparison of 4 Algorithms Step. Miner Algo Step height = 5σ. Number of timepoints = 15. A total of 2000 random data, 2000 one step data and 2000 two step data with random step positions.

Comparison of 4 Algorithms Step height = 5σ. Number of timepoints = 15. A total of 2000 random data, 2000 one step data and 2000 two step data with random step positions.

Generation of Simulated Data • Microarray data with 15 non-uniform time points • 4000 genes with 2000 one-step and 200 two-step patterns • Gaussian noise was added to the above data • P-value threshold of 0. 05 was used
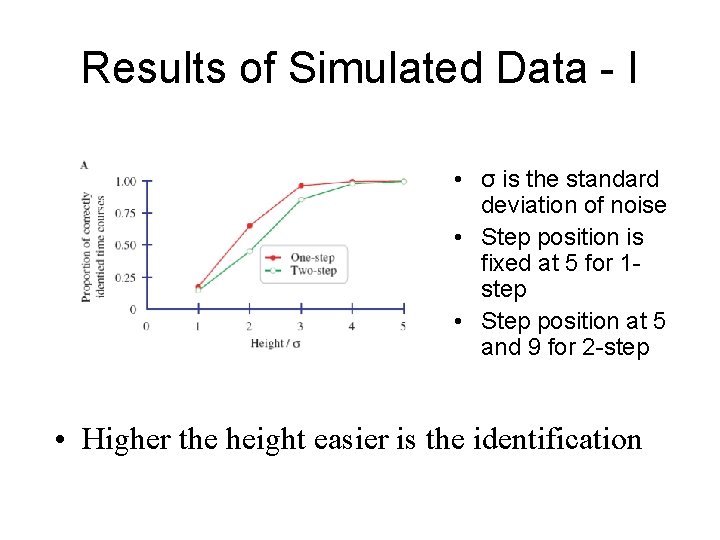
Results of Simulated Data - I • σ is the standard deviation of noise • Step position is fixed at 5 for 1 step • Step position at 5 and 9 for 2 -step • Higher the height easier is the identification
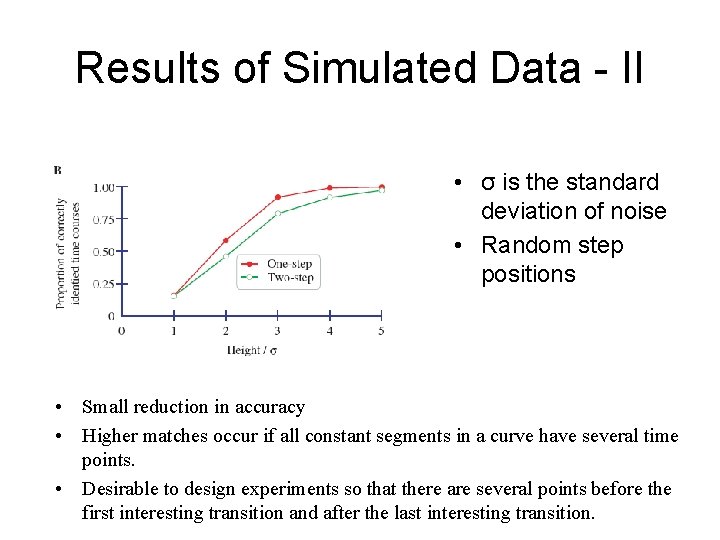
Results of Simulated Data - II • σ is the standard deviation of noise • Random step positions • Small reduction in accuracy • Higher matches occur if all constant segments in a curve have several time points. • Desirable to design experiments so that there are several points before the first interesting transition and after the last interesting transition.
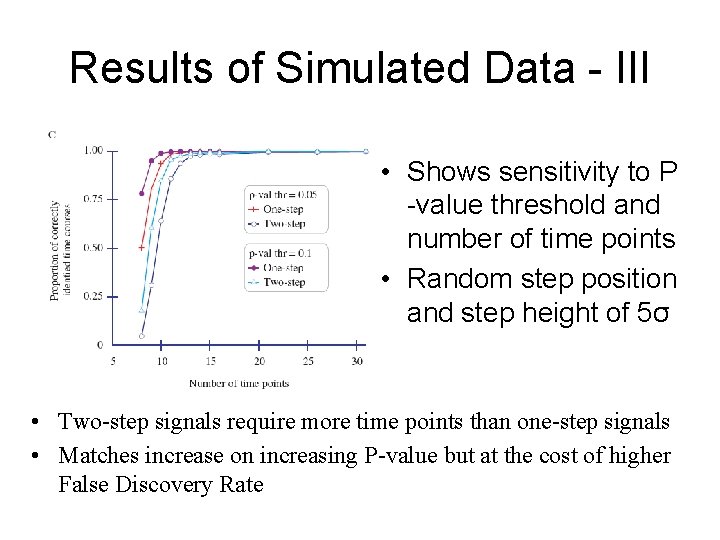
Results of Simulated Data - III • Shows sensitivity to P -value threshold and number of time points • Random step position and step height of 5σ • Two-step signals require more time points than one-step signals • Matches increase on increasing P-value but at the cost of higher False Discovery Rate
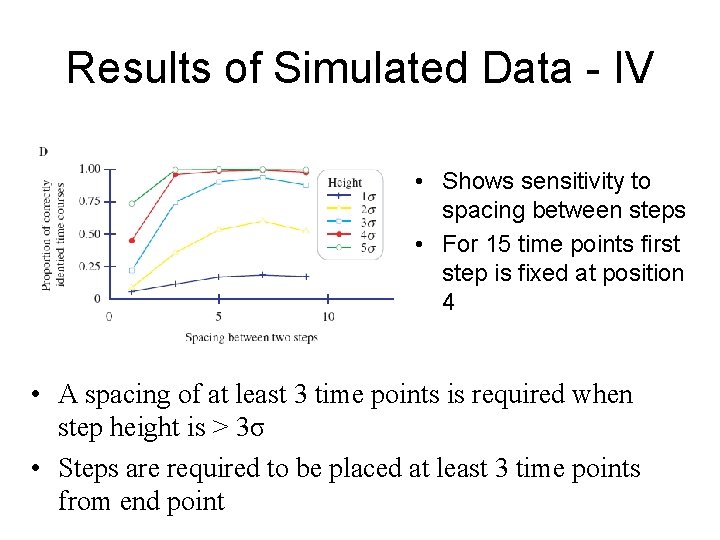
Results of Simulated Data - IV • Shows sensitivity to spacing between steps • For 15 time points first step is fixed at position 4 • A spacing of at least 3 time points is required when step height is > 3σ • Steps are required to be placed at least 3 time points from end point

Diauxic Shift • In the initial phases of a growing batch culture, yeast prefers to metabolize glucose and produce ethanol even when oxygen is abundant. • When the glucose is exhausted, cells undergo a “diauxic shift, ” in which they switch abruptly to an oxidative metabolism. This pathway allows the oxidation of the accumulated fermentation products and is highly efficient as a mechanism for generating ATP. Brauer et. al. , Mol Biol Cell. 2005 May; 16(5): 2503– 2517
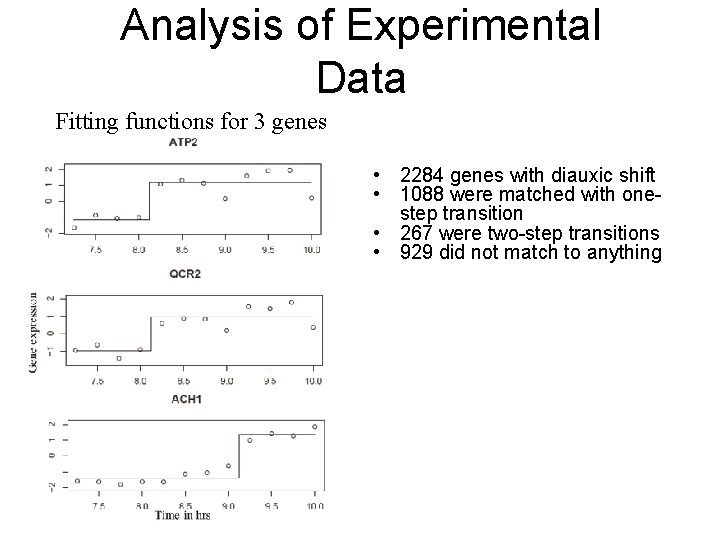
Analysis of Experimental Data Fitting functions for 3 genes • 2284 genes with diauxic shift • 1088 were matched with onestep transition • 267 were two-step transitions • 929 did not match to anything
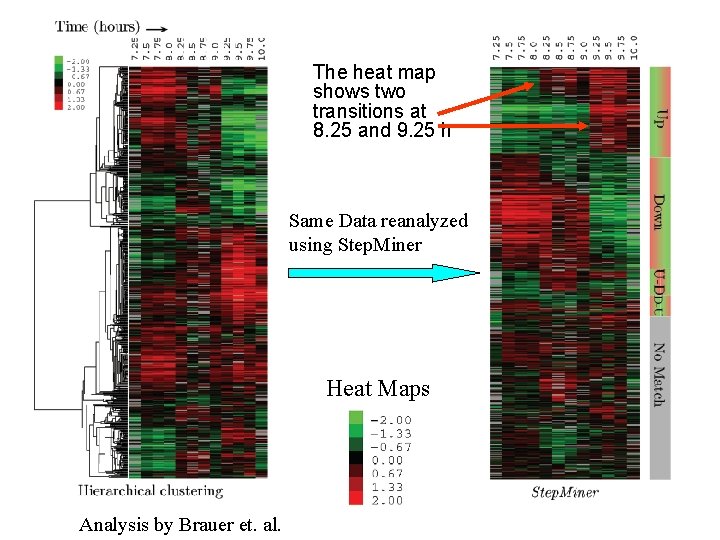
The heat map shows two transitions at 8. 25 and 9. 25 h Same Data reanalyzed using Step. Miner Heat Maps Analysis by Brauer et. al.

Comparison With Brauer et al’s Results • The GO annotations and FDR-corrected P-values for the clusters reported in Brauer et al. was recomputed with the latest yeast gene annotations from the Gene Ontology Consortium Website • Table shows the results of the p-values from GO- Term Finder as well as Step Miner.
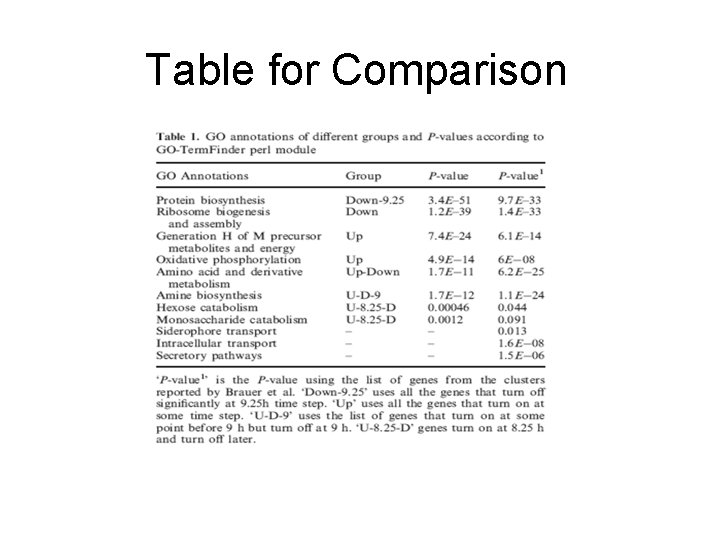
Table for Comparison

Results Of Comparison • The annotation that had the lowest P-values in Brauer et al. had even low P-values in the Step. Miner groups. • In most cases, the P-values in the reanalysis are lower than Brauer et al’s, implies that grouping by time-of-change is at least as effective as hierarchical clustering at identifying relevant genes. • GO annotations are obtained fully automatically using Step. Miner – it is not necessary to select interesting clusters manually. • Those clusters which has no P-values from Step. Miner were “less interpretable in terms of diauxic shift”, in the words of Brauer et al.

Comparison of Step. Miner to Other Tools • Hierarchical clustering: finds clusters that transition at same time point – Manual search required to find transitions • SAM: finds transitions by looking for significant differences in average expression before and after a specified time point. – However, many of the genes selected by this method do not, in fact, have a transition at the specified time point. • EDGE: identify genes whose expression systematically change over time and significantly different from the mean of the expressions over time. – Clearly, this method doesn’t provide the direction and position of significant change directly.
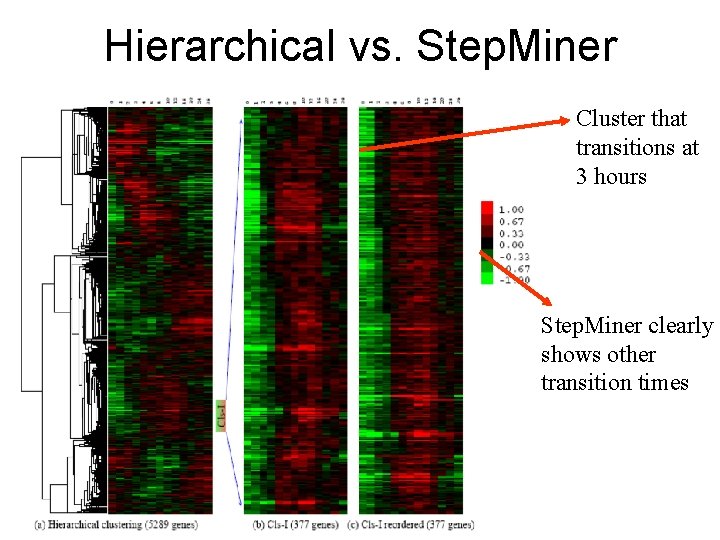
Hierarchical vs. Step. Miner Cluster that transitions at 3 hours Step. Miner clearly shows other transition times
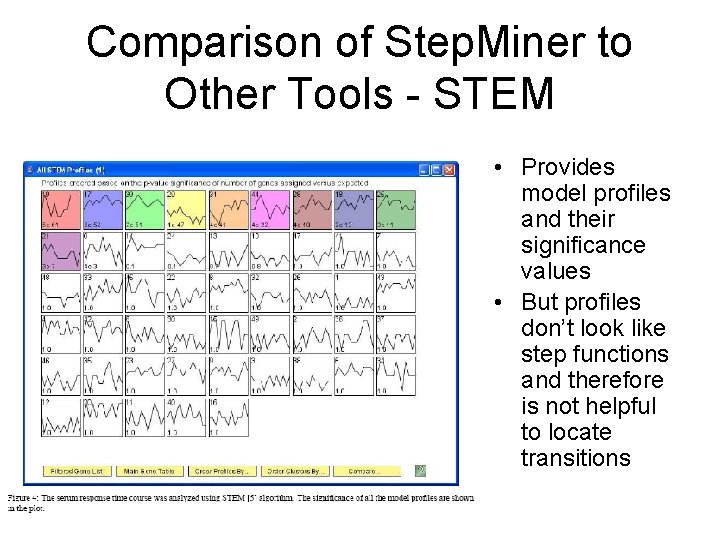
Comparison of Step. Miner to Other Tools - STEM • Provides model profiles and their significance values • But profiles don’t look like step functions and therefore is not helpful to locate transitions

Strengths and Limitations • Easy to understand • Few parameters • Biologically transitions can be more interesting • Very fast < 15 s for 15 microarrays of 40000 genes • Can deal with missing measurements • Provides statistical parameters like P-value, FDR etc. • Binary model • There can be other cases: eg, transition is not step • Short and long time courses are not good Most appropriate for 10 -30 Time measurements.

Post Step. Miner Analysis • Once Step. Miner is run genes undergoing binary transitions can easily be partitioned into sets based on the number, direction, and timing of transitions. • These sets can be merged at the user’s discretion (e. g. , the set of one-step genes that rise at time 3 could be merged with the twostep genes that rise at time 3), or can be further subdivided etc.

• BACK UP SLIDES

Replication vs. Resolution • For accuracy it is better to take more frequent measurements that to get replicates • It comes at a cost of correctly identifying the kind of step
- Slides: 25