Extended sibships Danielle Posthuma Kate Morley Files danielleSibs

Extended sibships Danielle Posthuma Kate Morley Files: \danielleSibs TC 19 - Boulder 2006

Classic Twin Design Twins share the same womb at the same n ACE / ADE q q n time, thisassortative may not only make. DZ them Due to mating, twinmore similar, but sharing theleading womb to itself may correlations will rise, increased also lead to twin Cultural transmission mayspecific lead toto GE estimates ofcomplications C. births, rendering twins unrepresentative of covariance in the offspring the normal population heterogeneity multivariate Sibling interaction Developmental Issues q q q generalizibility >Additional Siblings Assortative mating >Parents/spouses Cultural transmission >Parents TC 19 - Boulder 2006

There is more than the classical twin design n Larger pedigrees n n n Parent-offspring (incl. cultural transmission, assortative mating) Grandparents-offspring Spouses of co-twins/siblings Larger sibships Adoption studies n n n MZA DZA MZT DZT Non-biological siblings Virtual twins (non-biological siblings of same age) TC 19 - Boulder 2006
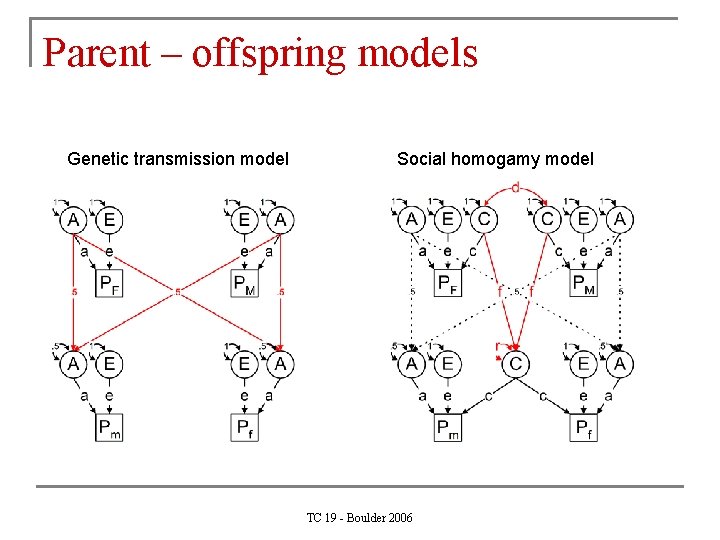
Parent – offspring models Genetic transmission model Social homogamy model TC 19 - Boulder 2006
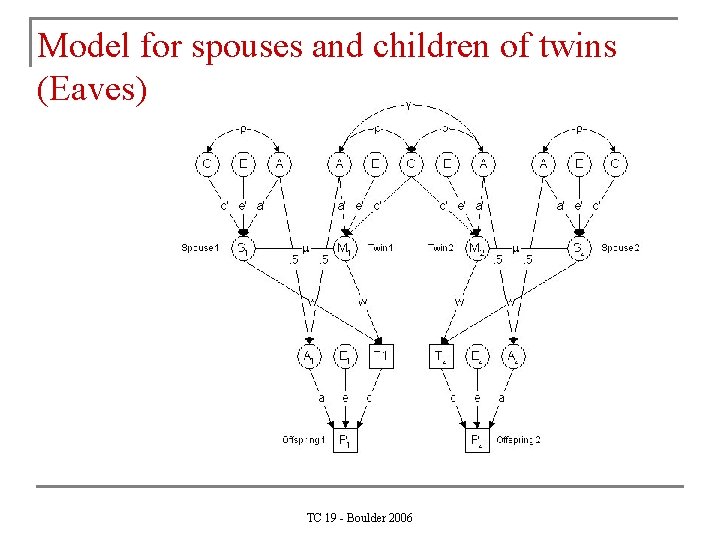
Model for spouses and children of twins (Eaves) TC 19 - Boulder 2006

Extended sibships TC 19 - Boulder 2006
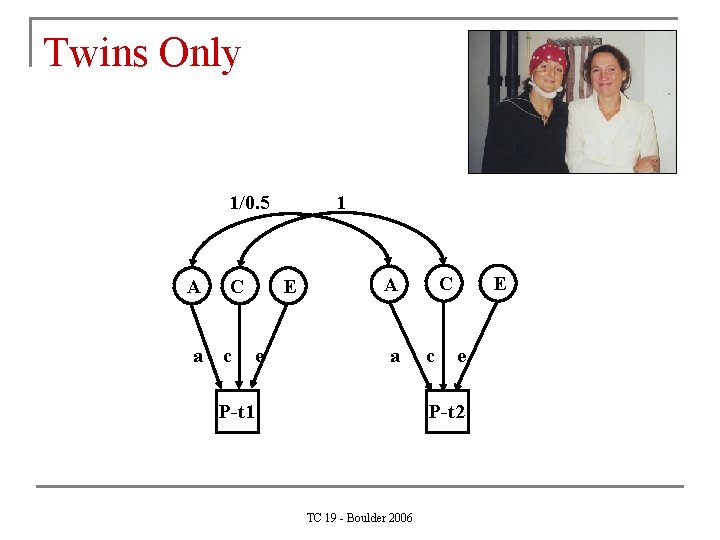
Twins Only 1/0. 5 A a C c 1 E e C A a P-t 1 c E e P-t 2 TC 19 - Boulder 2006

Twins Only: var-cov matrices MZ DZ t 1 t 2 t 1 a 2+c 2+e 2 a 2+c 2 t 2 a 2+c 2+e 2 t 1 t 2 t 1 a 2+c 2+e 2 0. 5 a 2+c 2 t 2 0. 5 a 2+c 2+e 2 In Mx: MZ group: DZ group: A+C+E | A+C_ A+C+E | H@A+C_ A+C H@A+C| A+C+E ; TC 19 - Boulder 2006

Adding siblings Is easy! TC 19 - Boulder 2006 But why should I?
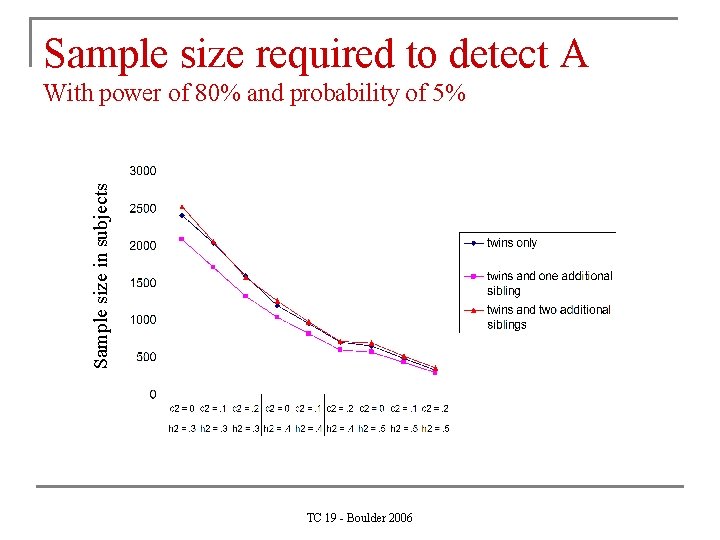
Sample size required to detect A Sample size in subjects With power of 80% and probability of 5% TC 19 - Boulder 2006
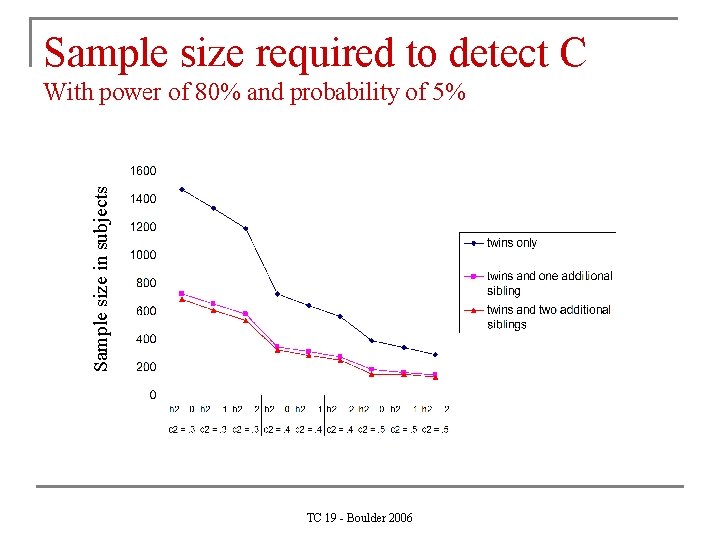
Sample size required to detect C Sample size in subjects With power of 80% and probability of 5% TC 19 - Boulder 2006

Larger sibships n. Provides a bit more power to detect A n. Provides a lot more power to detect C Since C is usually small (e. g. A =. 60, C =. 20, E =. 20), C is usually dropped from the model as it is not significant. As C is a familial source of variance, part of it will the end up in A, which will now be overestimated. Therefore, more power for C protects against overestimation of A. TC 19 - Boulder 2006

Larger sibships n Will also allow you to test certain assumptions such as: q q q Are twins different from singletons with respect to means? Are twins different from singletons with respect to variances? Do DZ twins correlate any different than non-twin sibpairs? TC 19 - Boulder 2006
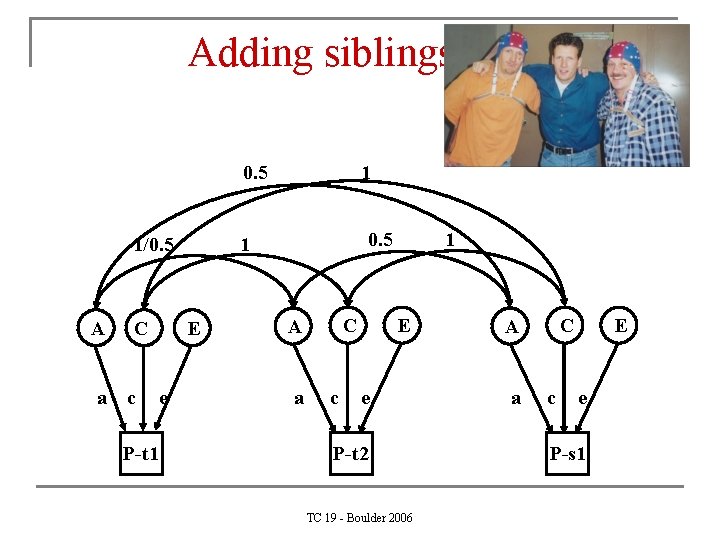
Adding siblings 0. 5 1/0. 5 A a C c P-t 1 0. 5 1 E e 1 C A a c 1 E e P-t 2 TC 19 - Boulder 2006 C A a c E e P-s 1

MZ and one additional sibling t 1 t 2 s 1 t 1 a 2+c 2+e 2 a 2+c 2 0. 5 a 2+c 2 t 2 a 2+c 2+e 2 0. 5 a 2+c 2 s 1 0. 5 a 2+c 2+e 2 DZ and one additional sibling t 1 t 2 s 1 t 1 a 2+c 2+e 2 0. 5 a 2 +c 2 0. 5 a 2+c 2 t 2 0. 5 a 2 +c 2 a 2+c 2+e 2 0. 5 a 2+c 2 s 1 0. 5 a 2+c 2+e 2 TC 19 - Boulder 2006

Exercise n n Copy Twins. Only. mx and mriiq. rec Open Mx script Twins. Only. mx Modify this script such that n Data from sib 1 is included n Data from sib 1 to sib 6 are included n Check – 2 ll, df, estimated pms, n observations for each model TC 19 - Boulder 2006 .

Twins +1 Twins +6 -2 ll 3878. 177 5025. 192 5388. 980 df 494 639 684 TC 19 - Boulder 2006 est 4 4 4 obs 498 643 688

Adding more siblings becomes tedious! (and errorprone. . ) MZ’s and 6 additional siblings A+C+E | A+C | H@A+C |H@A+C _ A+C | A+C+E | H@A+C |H@A+C _ H@A+C | A+C+E |H@A+C |H@A+C _ H@A+C | A+C+E | H@A+C | H@A+C __ H@A+C | A+C+E |H@A+C | H@A+C __ H@A+C | H@A+C |A+C+E | H@A+C __ H@A+C | H@A+C | H@A+C | H@A+C | A+C+E ; TC 19 - Boulder 2006

Adding more siblings 6 extra siblings MZ’s and 6 additional siblings A+C+E | A+C | H@A+C |H@A+C _ A+C | A+C+E | H@A+C |H@A+C _ H@A+C | A+C+E |H@A+C |H@A+C _ H@A+C | A+C+E | H@A+C | H@A+C __ H@A+C | A+C+E |H@A+C | H@A+C __ H@A+C | H@A+C |A+C+E | H@A+C __ H@A+C | H@A+C | H@A+C | H@A+C | A+C+E ; TC 19 - Boulder 2006
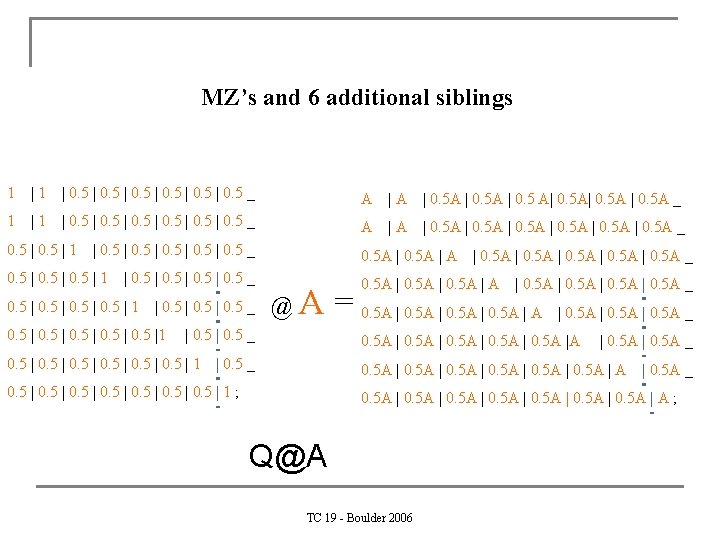
MZ’s and 6 additional siblings 1 | 0. 5 | 0. 5 _ A |A | 0. 5 A | 0. 5 A| 0. 5 A _ 1 | 0. 5 | 0. 5 _ A |A | 0. 5 A _ 0. 5 | 1 | 0. 5 _ 0. 5 A | A 0. 5 | 1 0. 5 A | 0. 5 _ 0. 5 | 1 | 0. 5 _ 0. 5 |1 @A | 0. 5 _ 0. 5 | 1 | 0. 5 A | 0. 5 A _ = 0. 5 A | A | 0. 5 A _ 0. 5 A |A | 0. 5 _ | 0. 5 A _ 0. 5 A | A 0. 5 | 0. 5 | 1 ; | 0. 5 A _ 0. 5 A | 0. 5 A | A ; Q@A TC 19 - Boulder 2006
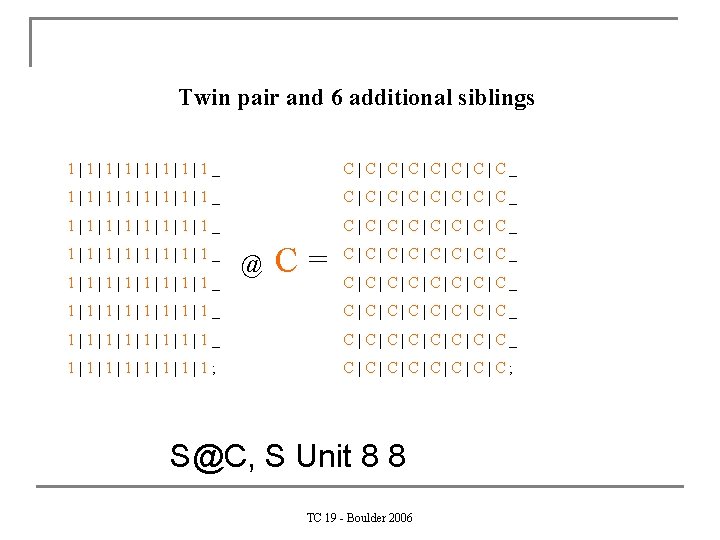
Twin pair and 6 additional siblings 1|1|1|1|1|1|1|1_ C|C|C|C|C|C|C|C_ 1|1|1|1|1_ C|C|C|C|C_ 1|1|1|1|1|1|1|1_ @ C= C|C|C|C|C|C|C|C_ 1|1|1|1|1|1|1|1_ C|C|C|C|C|C|C|C_ 1|1|1|1|1; C|C|C|C|C; S@C, S Unit 8 8 TC 19 - Boulder 2006
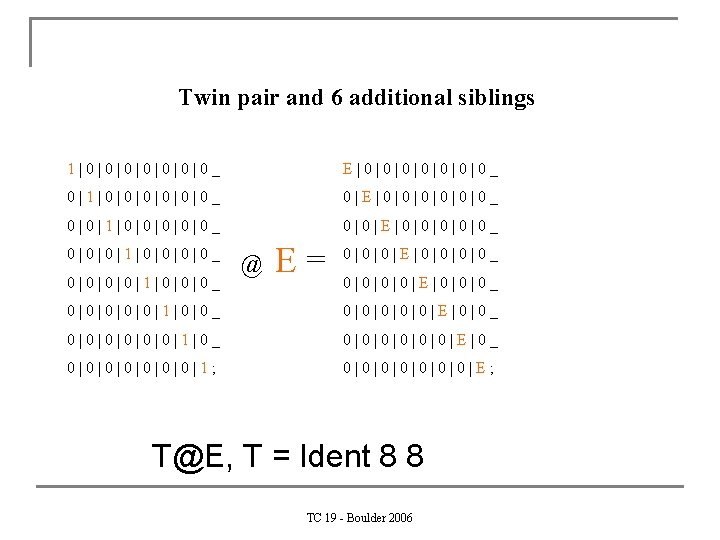
Twin pair and 6 additional siblings 1|0|0|0|0_ E|0|0|0|0_ 0|1|0|0|0_ 0|E|0|0|0_ 0|0|1|0|0|0_ 0|0|E|0|0|0_ 0|0|0|1|0|0_ 0|0|1|0|0|0_ @ E= 0|0|0|E|0|0_ 0|0|E|0|0|0_ 0|0|0|1|0|0_ 0|0|0|E|0|0_ 0|0|0|1|0_ 0|0|0|E|0_ 0|0|0|0|1; 0|0|0|0|E; T@E, T = Ident 8 8 TC 19 - Boulder 2006

Mx n n Copy Twins&6. mx Open Twins&6. mx TC 19 - Boulder 2006 .

Exercise n Modify this script for maximum nr of siblings =. 3, 4, or 5, write down – 2 ll, df, estimated pms, n observations for each model TC 19 - Boulder 2006
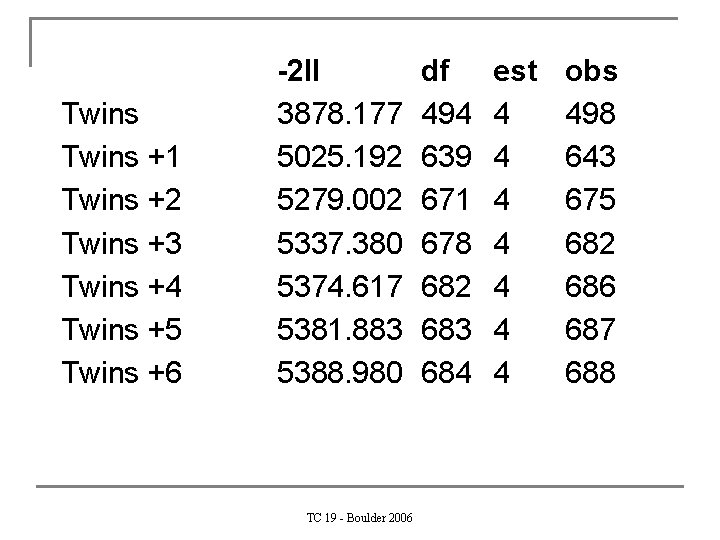
Twins +1 Twins +2 Twins +3 Twins +4 Twins +5 Twins +6 -2 ll 3878. 177 5025. 192 5279. 002 5337. 380 5374. 617 5381. 883 5388. 980 TC 19 - Boulder 2006 df 494 639 671 678 682 683 684 est 4 4 4 4 obs 498 643 675 682 686 687 688

Exercise n n Modify the script with 6 additional siblings (so 8 persons) to a bivariate script for wmem and greym. If correct: -2 ll=8083. 085, df = 935 You can start the mean for wmem at 60 and the mean for greym at 400. all variance components (SD) can be started at 30 Add standardization matrices for A and E TC 19 - Boulder 2006

In Summary n Adding additional persons: add expectations to Covariance statement n Adding additional phenotypes: change matrix dimensions TC 19 - Boulder 2006
- Slides: 27