Exploratory Data Analysis of High Density Oligonucleotide Array
























- Slides: 46

Exploratory Data Analysis of High Density Oligonucleotide Array Rafael A. Irizarry, Bridget Hobbs, Terry Speed http: //biosun 01. biostat. jhsph. edu/~ririzarr/Raffy

Outline • • • Review of technology Form of Data Description of Data Normalization Future/current work: Defining expression

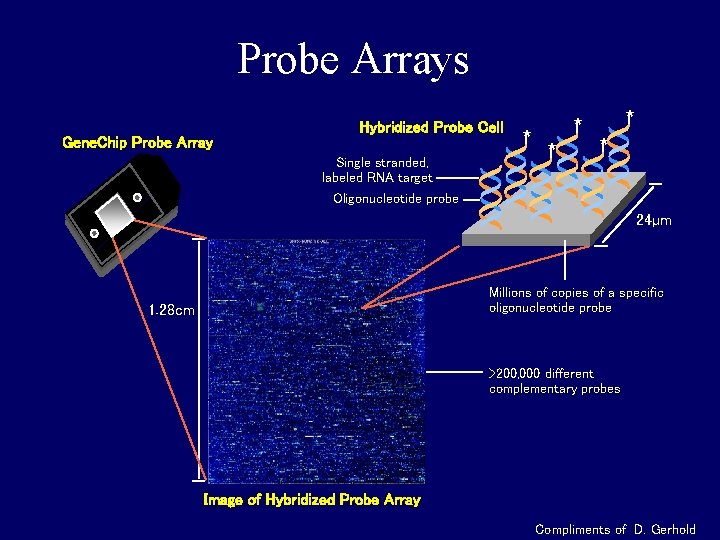
Probe Arrays Gene. Chip Probe Array Hybridized Probe Cell Single stranded, labeled RNA target Oligonucleotide probe * * * 24µm Millions of copies of a specific oligonucleotide probe 1. 28 cm >200, 000 different complementary probes Image of Hybridized Probe Array Compliments of D. Gerhold

Image analysis • About 100 pixels per probe cell • These intensities are combined to form one number representing expression for the probe cell oligo • What about genes?

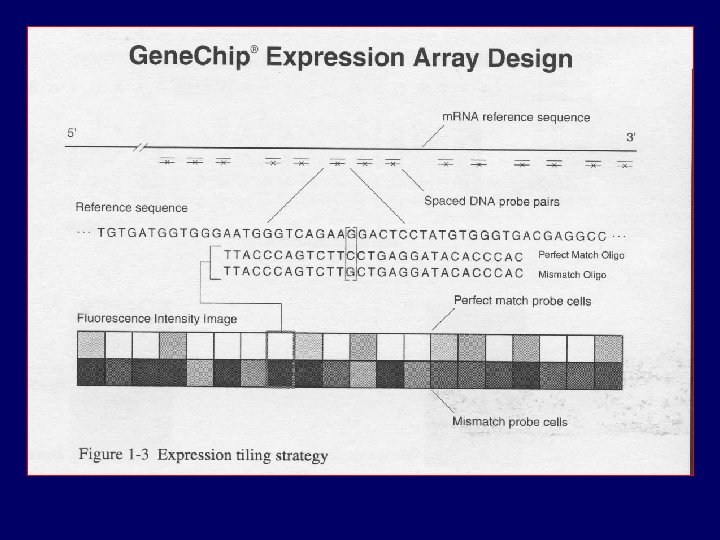
PM MM

Data and notation PMijn , MMijn = Intensity for perfect/mis-match probe cell j, in chip i, in gene n i = 1, …, I (ranging from 1 to hundreds) j=1, …, J (usually 16 or 20) n = 1, …, N (between 8, 000 and 12, 000)

$64 K Question • How do we define expression? or • What is the one number summary of the 20 PMs and 20 MMs that best quantifies expression? • How about differential expression?

Current default • Gene. Chip® software uses Avg. diff with A a set of “suitable” pairs chosen by software. • Log ratio version is also used. • For differential expression Avg. diffs are compared between chips.

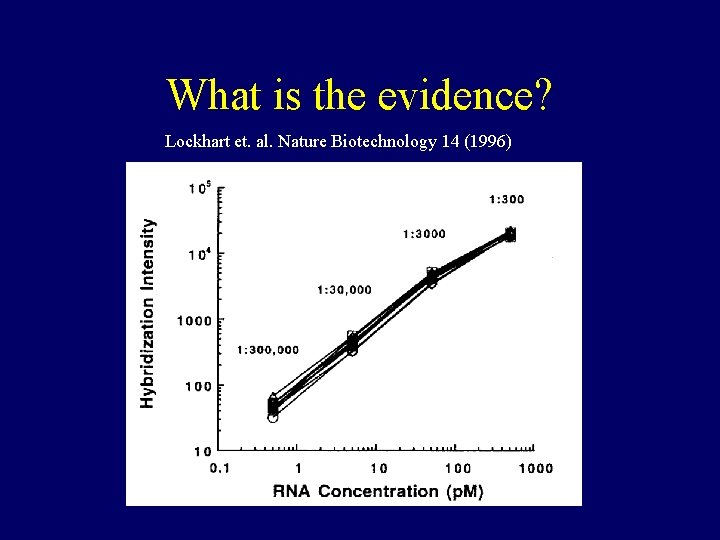
What is the evidence? Lockhart et. al. Nature Biotechnology 14 (1996)

• Chips used in Lockhart et. al. contained around 1000 probes per gene • Current chips contain 20 probes per gene • These are different situations • We haven’t seen a plot like the previous one, for current chips

Possible problems What if • a small number of the probe pairs hybridize much better than the rest? • removing the middle base does not make a difference for some probes? • some MM are PM for some other gene? • there is need for normalization? We explore these possibilities using data from 3 experiments

Experiment 1 • 8 Rats, under 4 experimental conditions – Control NV 21 – Ventilation V 21 – Oxygen NV 100 – Oxygen and Ventilation V 100 • 2 rats in each condition • RNA is pooled and divided to form 2 technical replicates for each condition

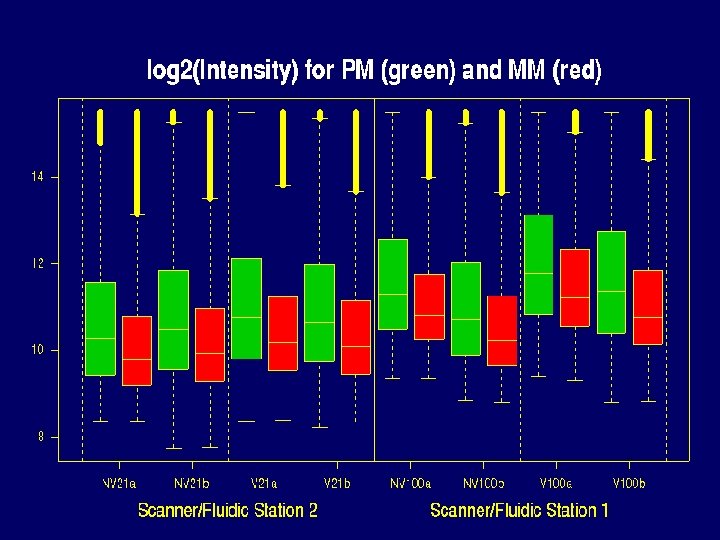
Notice • Experimental condition is confounded with couples: we can’t distinguish between biological variability and variability due to experimental condition • NV 21, V 21 and NV 100, V 100 processed in different scanners/fluidic stations: Oxygen effect confounded with scanner/fluidic station effect

Experiment 2 • 6 Rats, under 3 experimental conditions – Control – ENOS – NNOS • 2 rats in each condition • RNA is pooled and divided to form 2 technical replicates

Notice • One of the chips for NNOS did not “work” • Biological variability confounded with variability due to experimental condition • About 1/5 of the probes on chips used where defective.

Experiment 3 • Five mice with different characteristics: – 4 week old female NOD (J 4 FD, R 4 FD) – 4 week old female NOD (J 4 FD) – 4 week old male NOD (J 4 MD) – 4 week old female homozygous transgenic mouse which can't get diabetes (R 4 FN)

Notice • Each of the 5 chips were scanned twice • Two separate stains are used • This gives us 10 sets of results

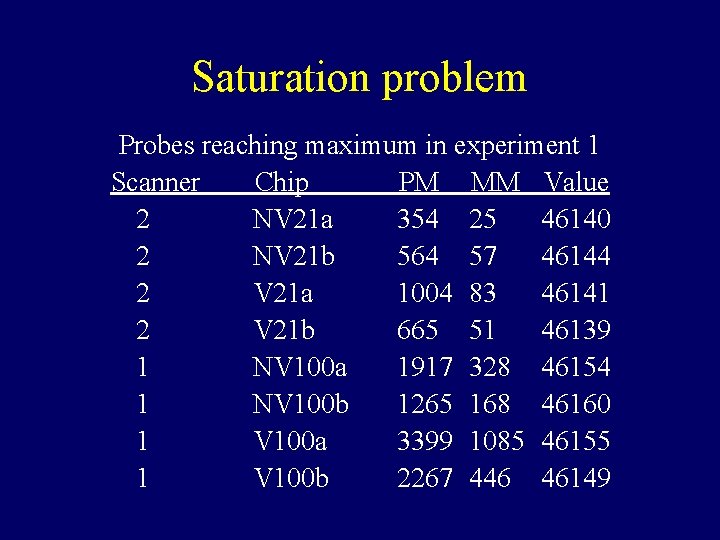
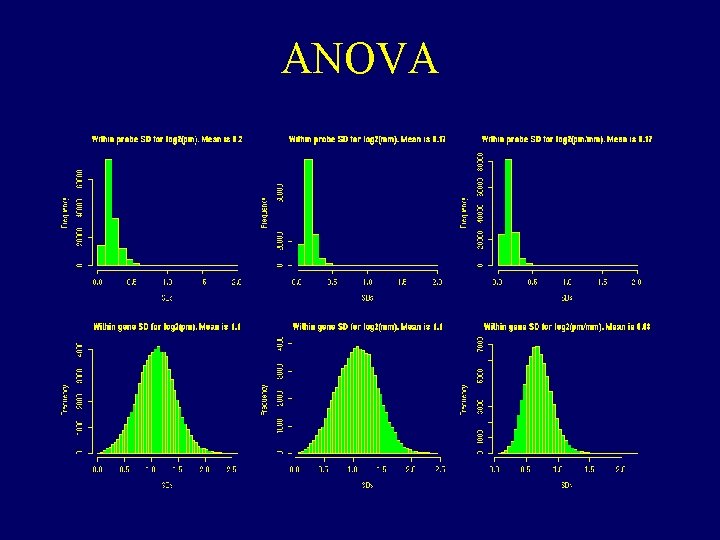
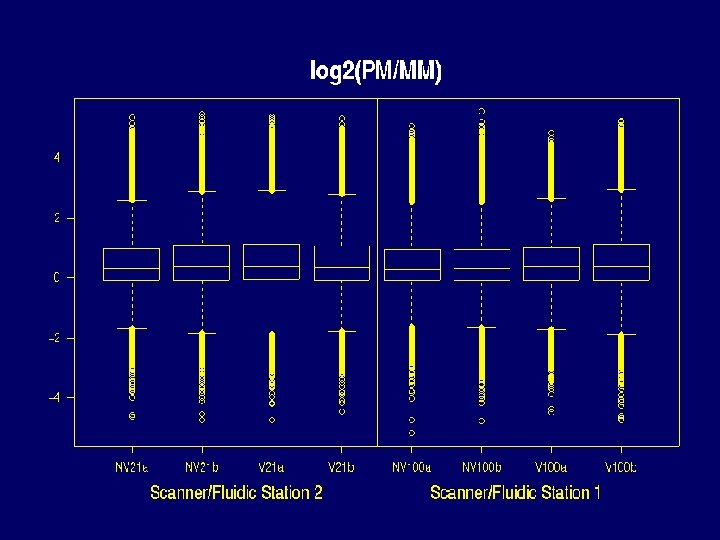
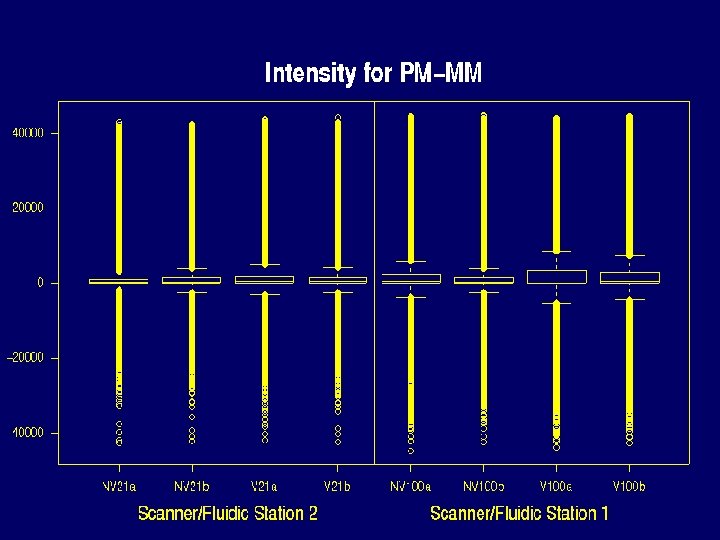
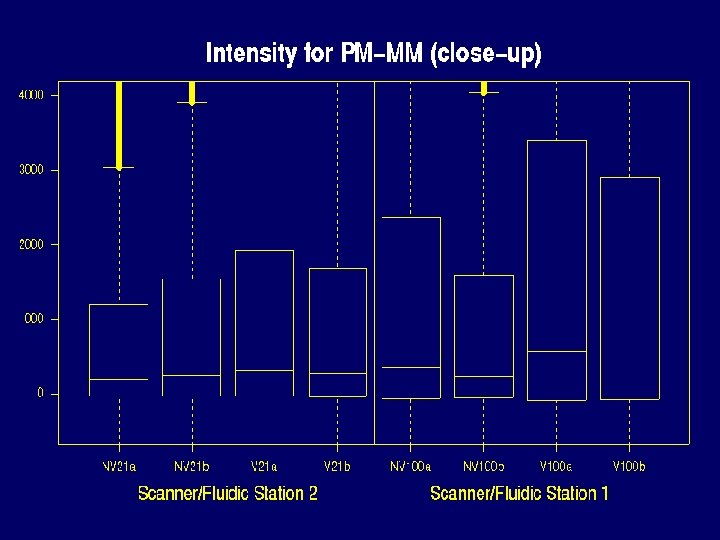
Properties of Data that make defining expression hard • • There can be saturation log 2(PM / MM) and PM-MM are noisy MM >> PM for many probes PMs of the same probe vary about 5 times less from chip to chip than from probe to probe within the same probe set.

Saturation problem Probes reaching maximum in experiment 1 Scanner Chip PM MM Value 2 NV 21 a 354 25 46140 2 NV 21 b 564 57 46144 2 V 21 a 1004 83 46141 2 V 21 b 665 51 46139 1 NV 100 a 1917 328 46154 1 NV 100 b 1265 168 46160 1 V 100 a 3399 1085 46155 1 V 100 b 2267 446 46149

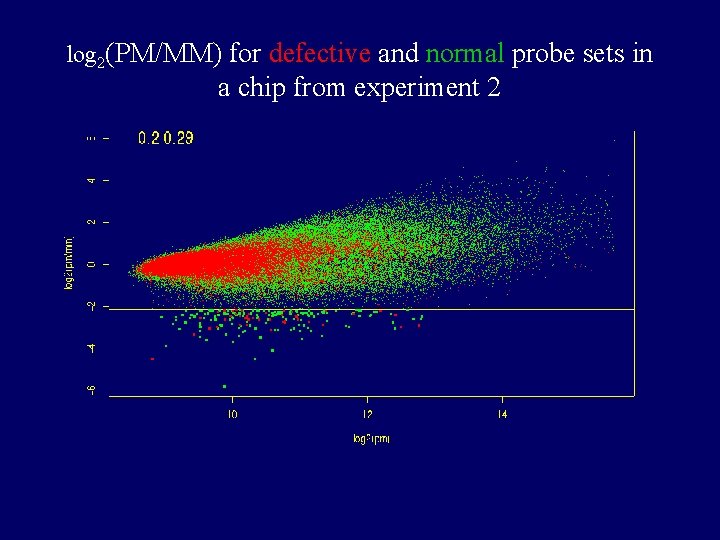
log 2(PM/MM) for defective and normal probe sets in a chip from experiment 2

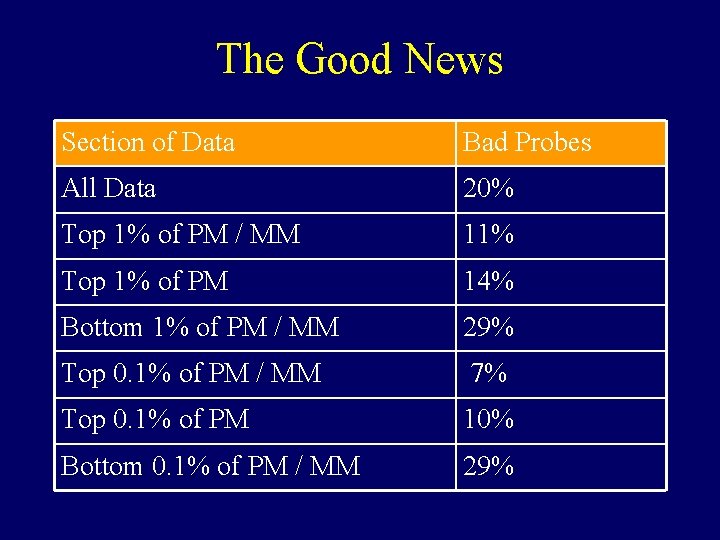
The Good News Section of Data Bad Probes All Data 20% Top 1% of PM / MM 11% Top 1% of PM 14% Bottom 1% of PM / MM 29% Top 0. 1% of PM / MM 7% Top 0. 1% of PM 10% Bottom 0. 1% of PM / MM 29%

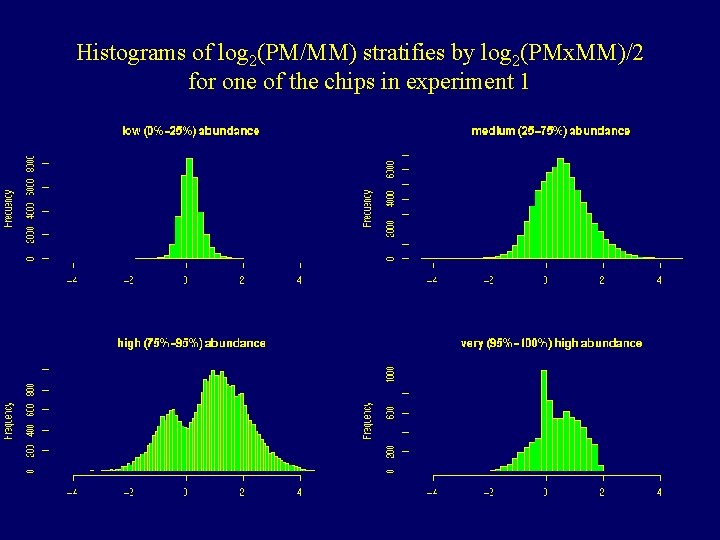
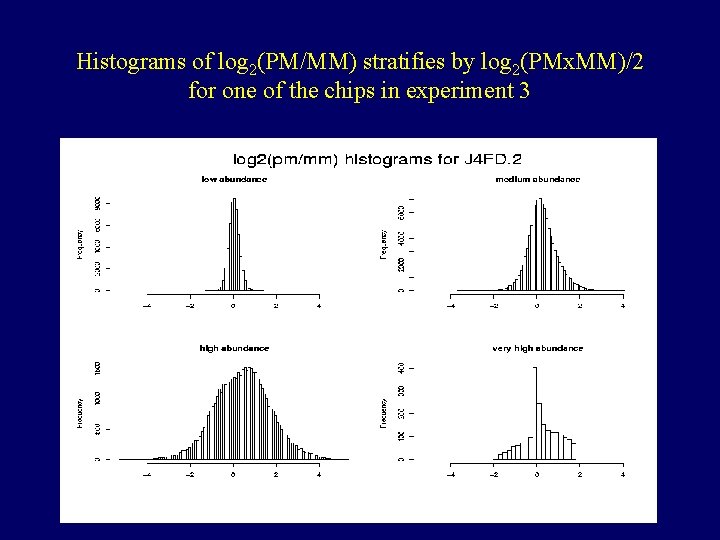
Histograms of log 2(PM/MM) stratifies by log 2(PMx. MM)/2 for one of the chips in experiment 1

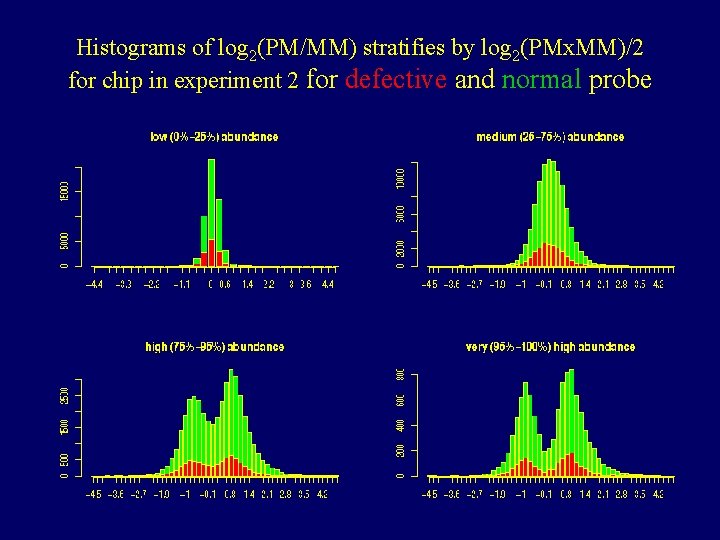
Histograms of log 2(PM/MM) stratifies by log 2(PMx. MM)/2 for chip in experiment 2 for defective and normal probe

Histograms of log 2(PM/MM) stratifies by log 2(PMx. MM)/2 for one of the chips in experiment 3

ANOVA

Normalization • There are many sources of experimental variation: – During preparation: e. g. m. RNA extraction, introduction of labeling – During manufacture of array: e. g. amount of oligos on cells – During hybridization: e. g. amount of sample applied, amount of target hybridized – After hybridization: e. g. optical measurements, label intensity, scanner • Proper normalization is need before intensities from different chips are compared





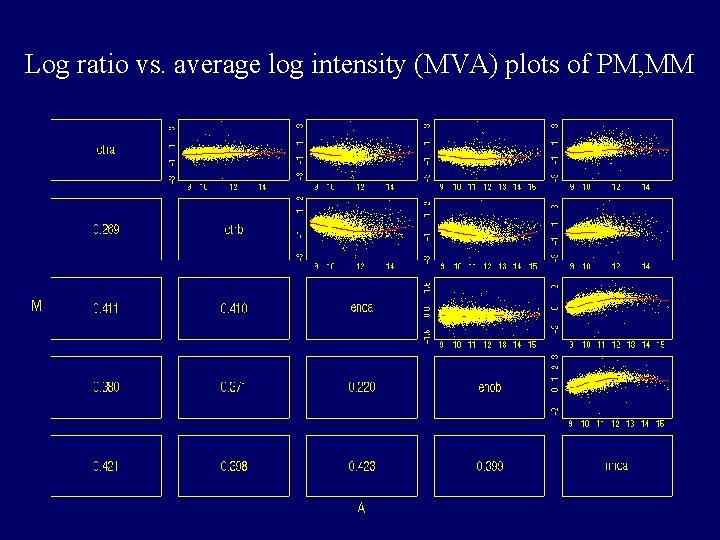
Log ratio vs. average log intensity (MVA) plots of PM, MM

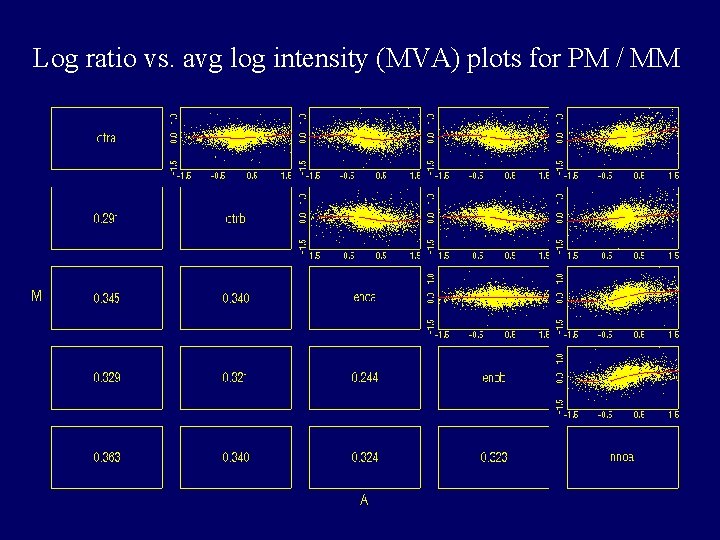
Log ratio vs. avg log intensity (MVA) plots for PM / MM

Normalization • Pair-wise normalization? • Which chips do we compare? • The following three plots show the 3 pairwise comparisons of chips Control A, ENOB, and NNOA

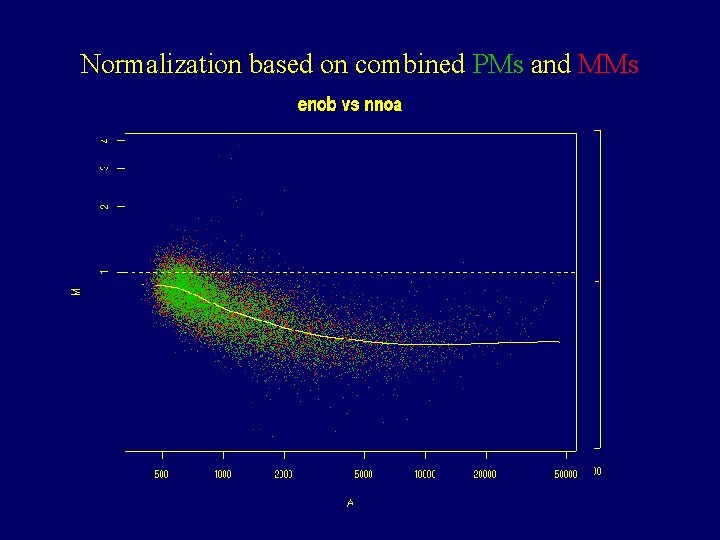
Normalization based on combined PMs and MMs

Cyclic algorithm (version 0. 1) • For chip j, with entries X 1 define the functions f 1, …, fj-1, fj+1, …, f. J to be the results of smoothing the scatter plot {Xj-Xk , (Xj+Xk)/2} • Define the normalized chip as Xj’= Xj- (f 1+…+fj-1+fj+1+…+f. J)/J • Chips X 1, …, XJ are normalized in the same way • We iterate until Xi’, Xi are very similar for all i.

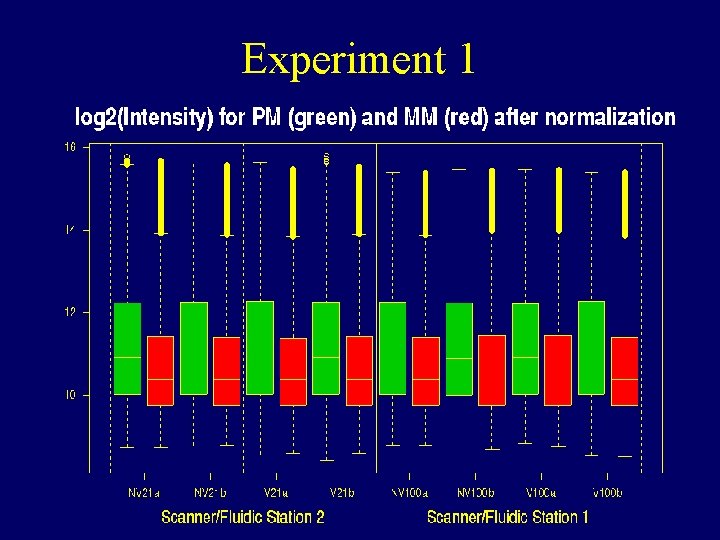
Before and after normalization

Experiment 1

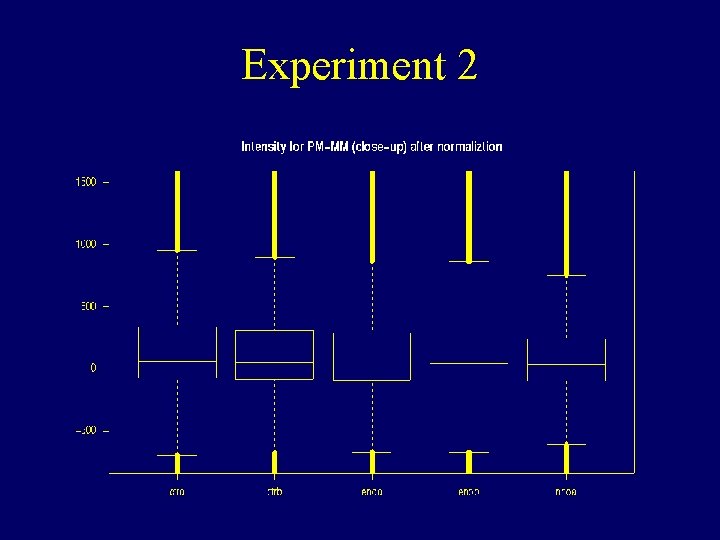
Experiment 2

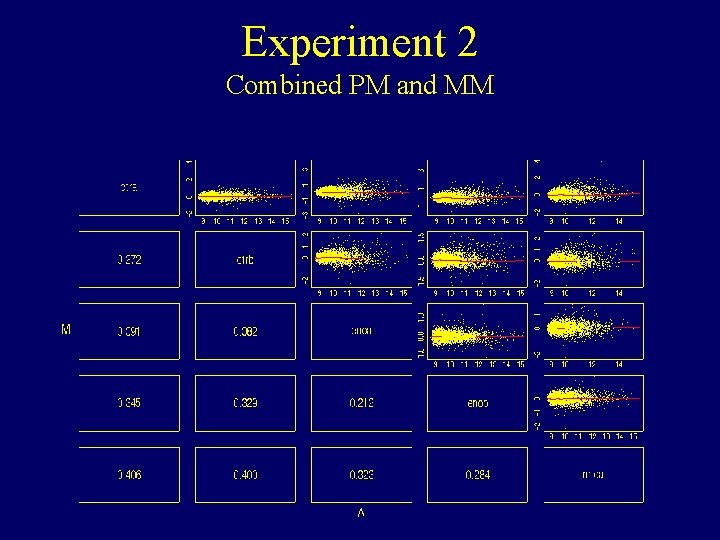
Experiment 2 Combined PM and MM

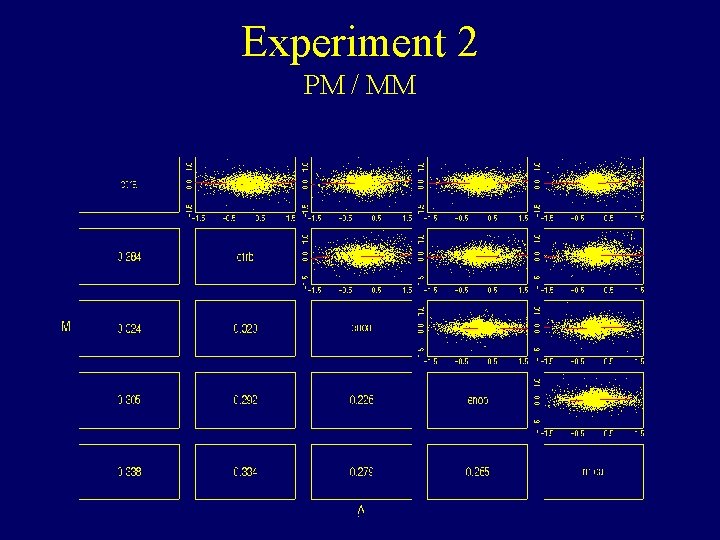
Experiment 2 PM / MM

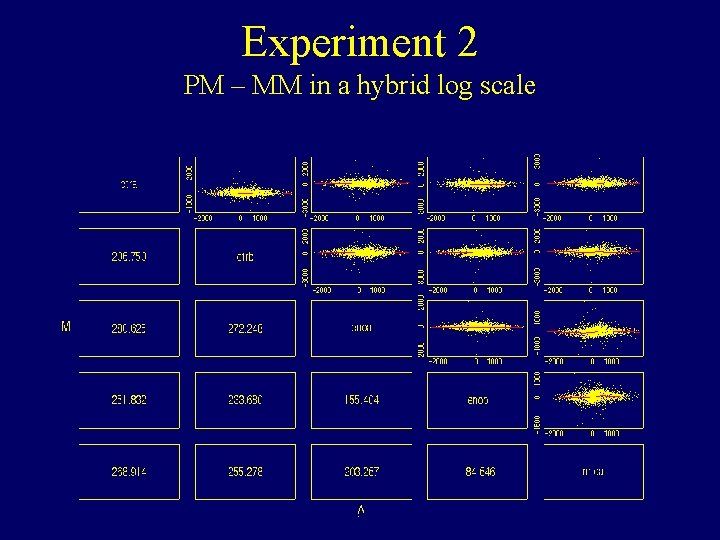
Experiment 2 PM – MM in a hybrid log scale

Experiment 3 Combined PM and MM

Competing definitions of expression • Li and Wong fit a model Consider expression in chip i • Efron et. al. consider log PM – 0. 5 log MM • Another is second largest PM

How do we compare? • We want small variance, small bias. • Up to now we don’t know truth in any of our data sets so hard to assess bias. • One possibility is to assume some gene is differentially expressed in the experiments we study, find it, and look at its probe profile.

Conclusion • Features of data suggest that avg. diff may be improved as a definition of expression • It seems that normalization is needed to remove experimental variation and make meaningful comparison of data from different chips fair

Acknowledgements • JHU: Leslie Cope, Tom Coppola, Shwu-Fan Ma, Skip Garcia • CNMC: Rehannah Borup, Josephine Chen, Eric Hoffman • UC Berkeley: Ben Bolstad • WEHI: Runa Daniel, Len Harrison