Exploration Normalization and Summaries of High Density Oligonucleotide
















- Slides: 37

Exploration, Normalization, and Summaries of High Density Oligonucleotide Array Probe Level Data Rafael A. Irizarry Department of Biostatistics, JHU (joint work with Leslie Cope, Ben Bolstad, Francois Collin, Bridget Hobbs, and Terry Speed) http: //biosun 01. biostat. jhsph. edu/~ririzarr

Summary • • Review of technology Probe level summaries Normalization Assess technology and expression measures • Conclusion/future work

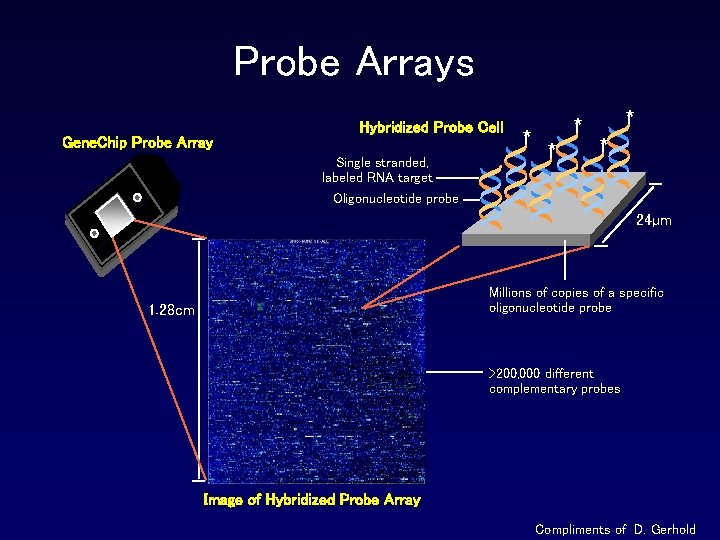
Probe Arrays Gene. Chip Probe Array Hybridized Probe Cell Single stranded, labeled RNA target Oligonucleotide probe * * * 24µm Millions of copies of a specific oligonucleotide probe 1. 28 cm >200, 000 different complementary probes Image of Hybridized Probe Array Compliments of D. Gerhold

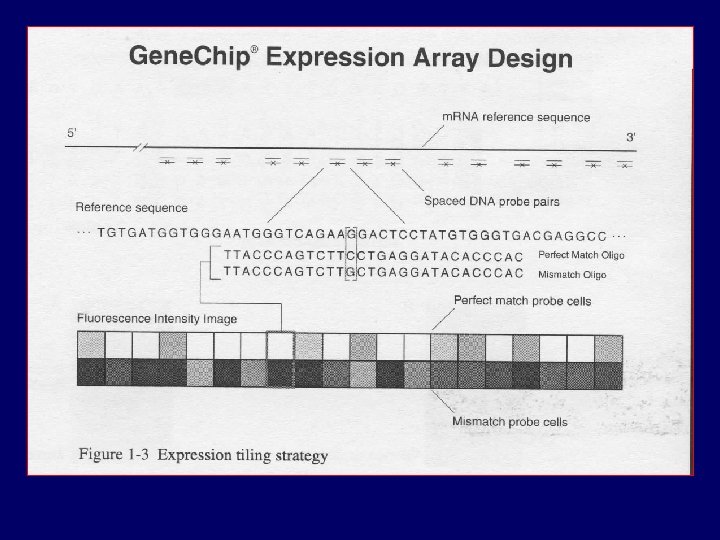
PM MM

Data and Notation PMijn , MMijn = Intensity for perfect/mis-match probe cell j, in chip i, in gene n i = 1, …, I (ranging from 1 to hundreds) j=1, …, J (usually 16 or 20) n = 1, …, N (between 8, 000 and 12, 000)

The Big Picture • Summarize 20 PM, MM pairs (probe level data) into one number for each gene • We call this number an expression measure • Affymetrix Gene. Chip’s Software has defaults. • Does it work? Can it be improved?

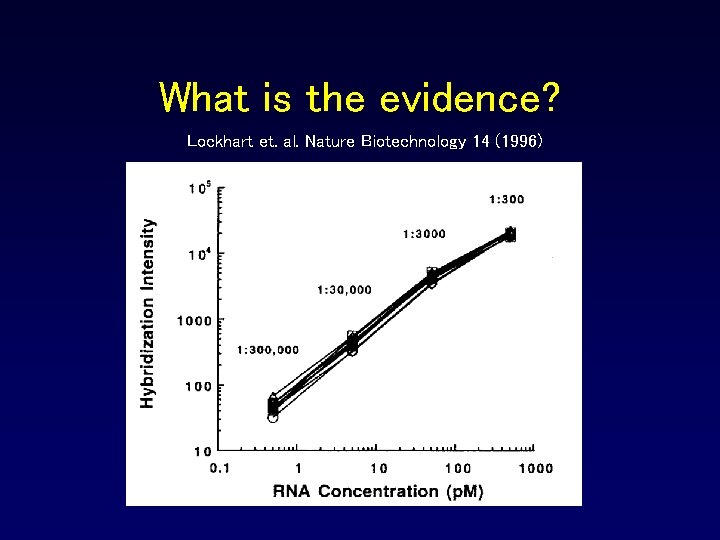
What is the evidence? Lockhart et. al. Nature Biotechnology 14 (1996)

Competing Measures of Expression • Gene. Chip® software uses Avg. diff with A a set of “suitable” pairs chosen by software. • Log ratio version is also used. • For differential expression Avg. diffs are compared between chips.

Competing Measures of Expression • Gene. Chip® new version uses something else with MM* a version of MM that is never bigger than PM.

Competing Measures of Expression • Li and Wong fit a model Consider expression in chip i • Efron et. al. consider log PM – 0. 5 log MM • Another is second largest PM

Competing Measures of Expression • Why not stick to what has worked for c. DNA? with A a set of “suitable” pairs.

Features of Probe Level Data

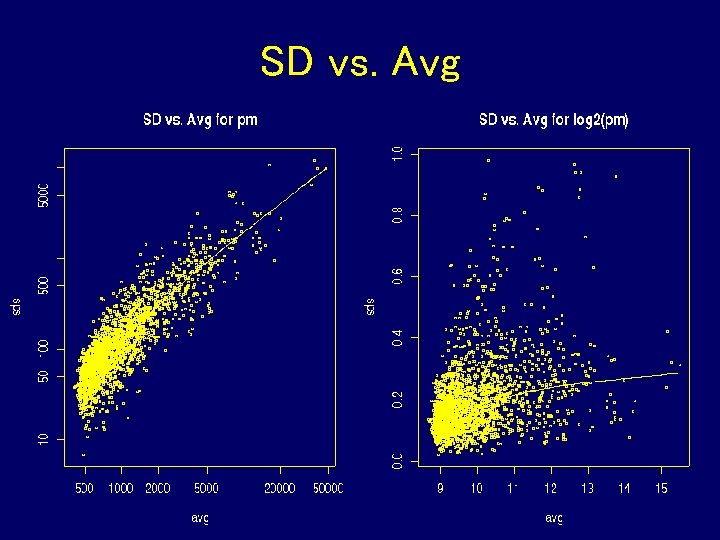
SD vs. Avg

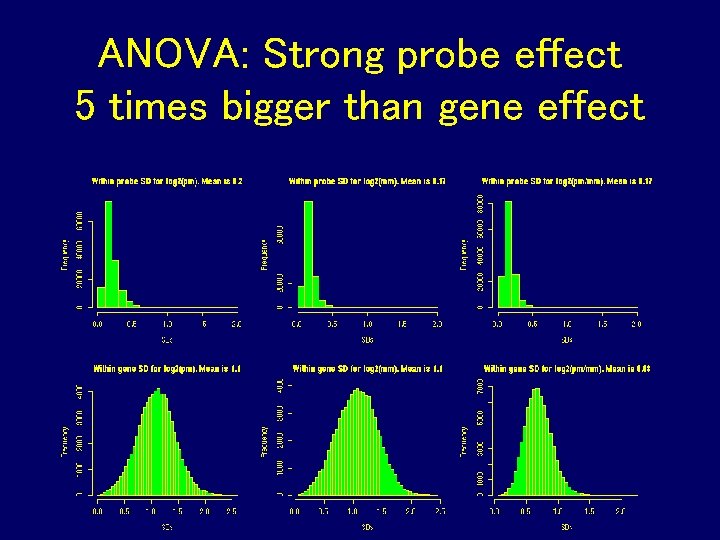
ANOVA: Strong probe effect 5 times bigger than gene effect

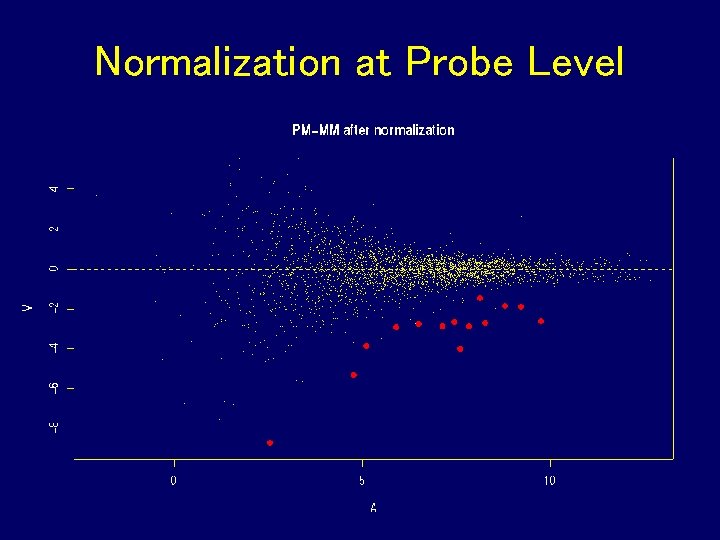
Normalization at Probe Level

Spike-In Experiments • Set A: 11 control c. RNAs were spiked in, all at the same concentration, which varied across chips. • Set B: 11 control c. RNAs were spiked in, all at different concentrations, which varied across chips. The concentrations were arranged in 12 x 12 cyclic Latin square (with 3 replicates)

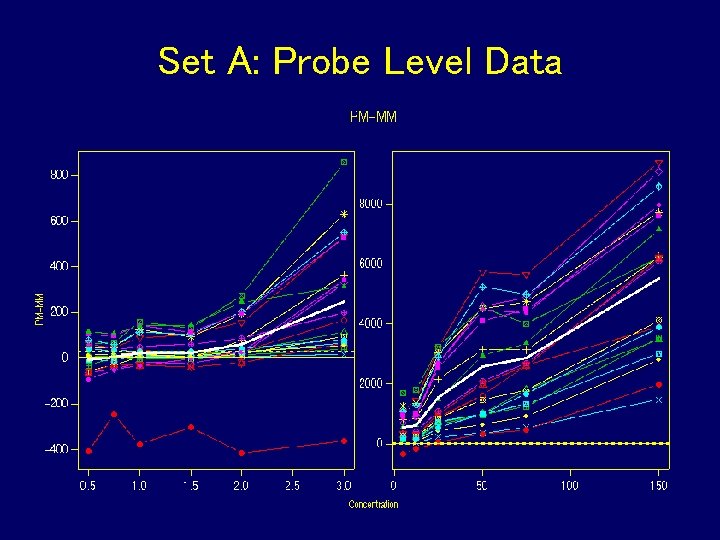
Set A: Probe Level Data

What Did We Learn? • Don’t subtract or divide by MM • Probe effect is additive on log scale • Take logs

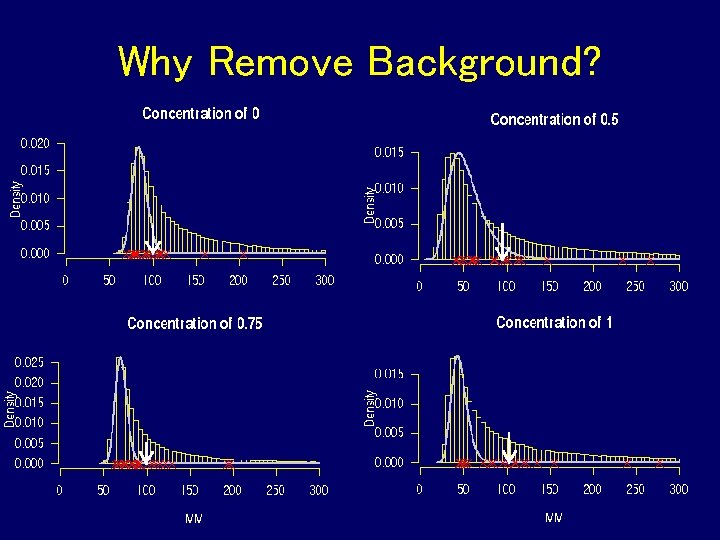
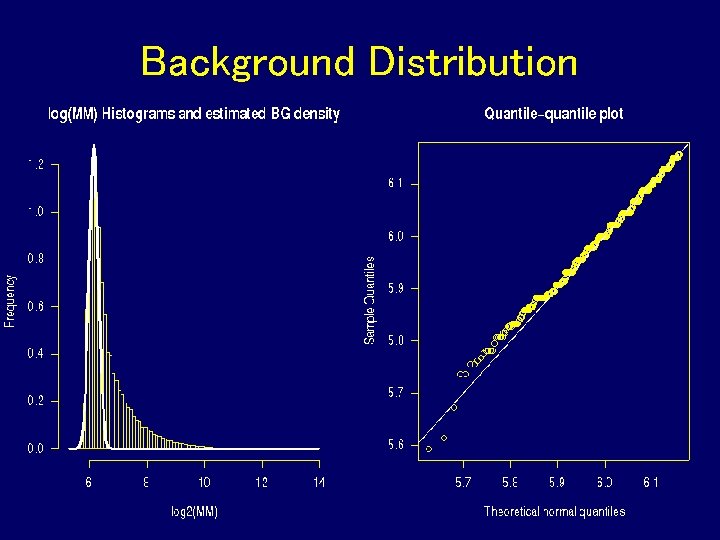
Why Remove Background?

Background Distribution

RMA • “Background correct” PM • Normalize (quantile normalization) • Assume additive model: • Estimate ai using robust method

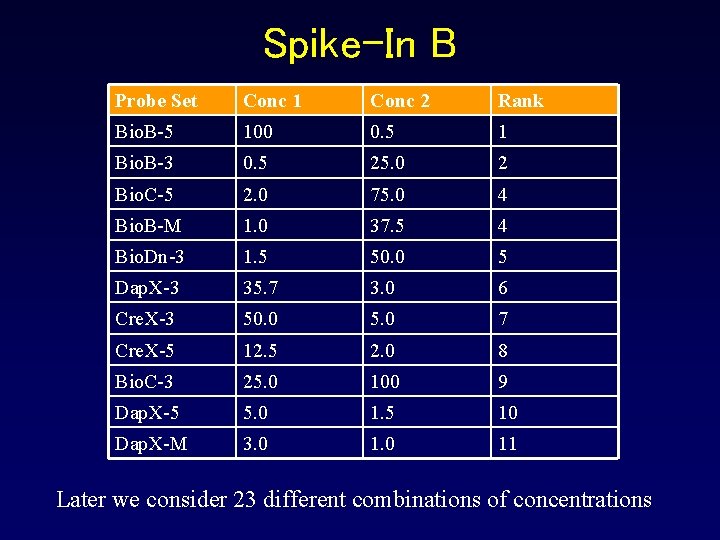
Spike-In B Probe Set Conc 1 Conc 2 Rank Bio. B-5 100 0. 5 1 Bio. B-3 0. 5 25. 0 2 Bio. C-5 2. 0 75. 0 4 Bio. B-M 1. 0 37. 5 4 Bio. Dn-3 1. 5 50. 0 5 Dap. X-3 35. 7 3. 0 6 Cre. X-3 50. 0 5. 0 7 Cre. X-5 12. 5 2. 0 8 Bio. C-3 25. 0 100 9 Dap. X-5 5. 0 1. 5 10 Dap. X-M 3. 0 11 Later we consider 23 different combinations of concentrations

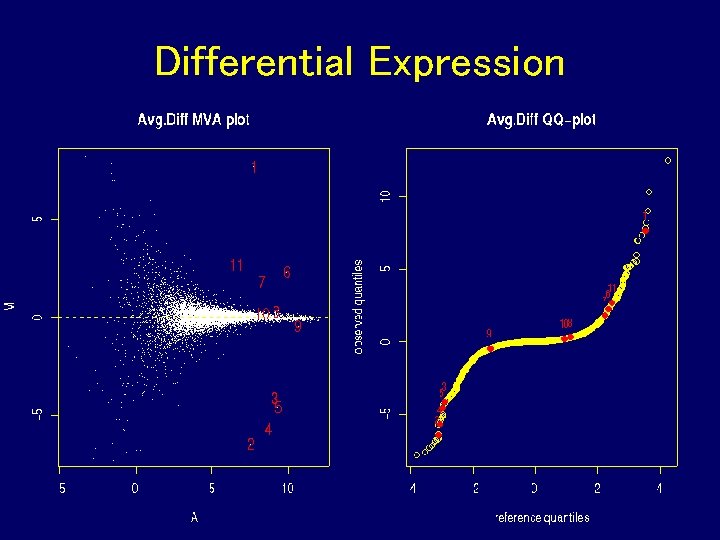
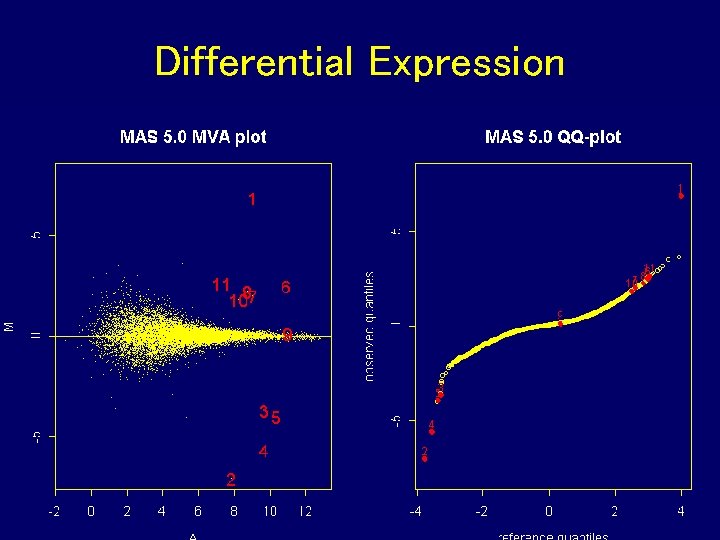
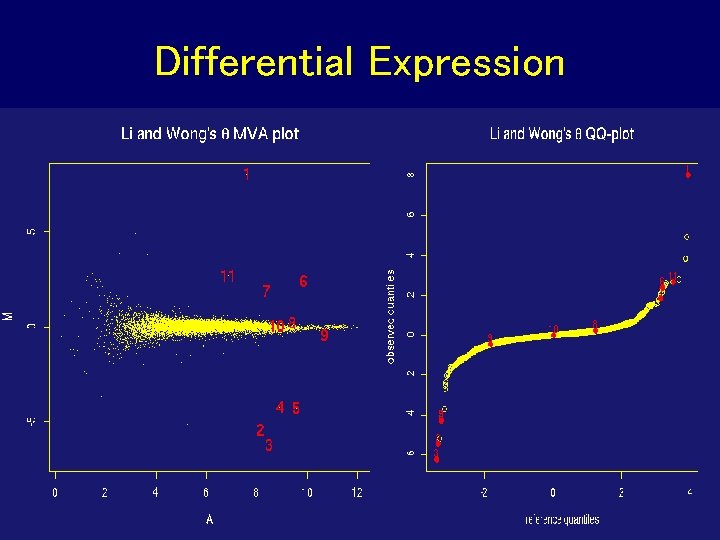
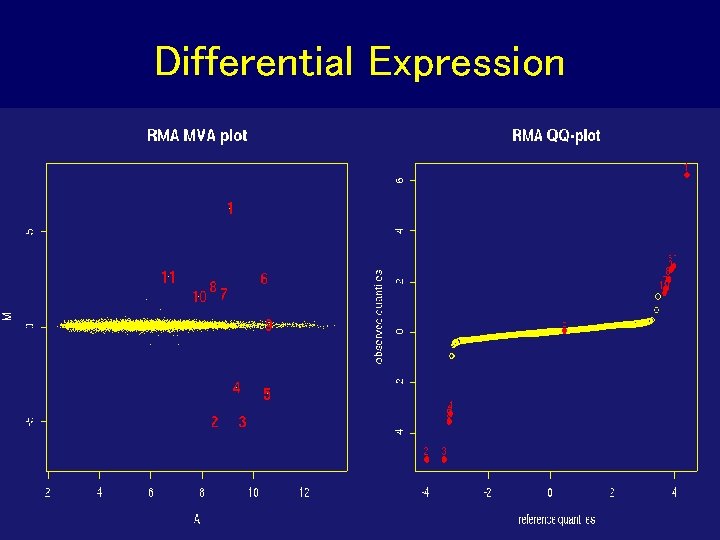
Differential Expression

Differential Expression

Differential Expression

Differential Expression

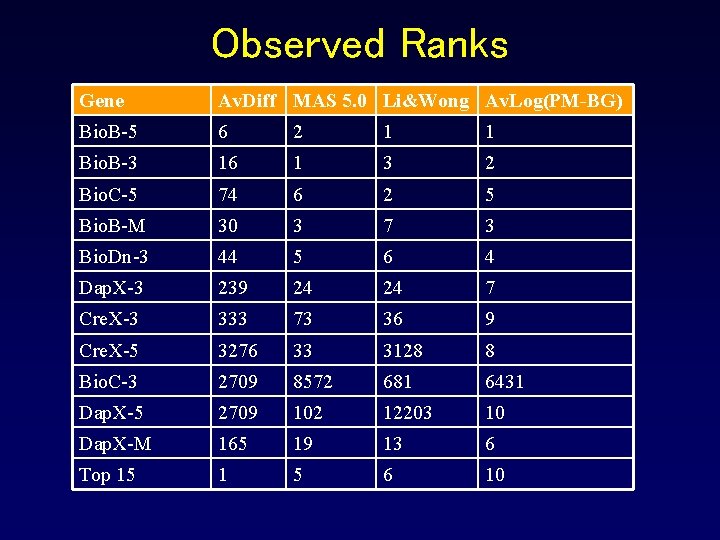
Observed Ranks Gene Av. Diff MAS 5. 0 Li&Wong Av. Log(PM-BG) Bio. B-5 6 2 1 1 Bio. B-3 16 1 3 2 Bio. C-5 74 6 2 5 Bio. B-M 30 3 7 3 Bio. Dn-3 44 5 6 4 Dap. X-3 239 24 24 7 Cre. X-3 333 73 36 9 Cre. X-5 3276 33 3128 8 Bio. C-3 2709 8572 681 6431 Dap. X-5 2709 102 12203 10 Dap. X-M 165 19 13 6 Top 15 1 5 6 10

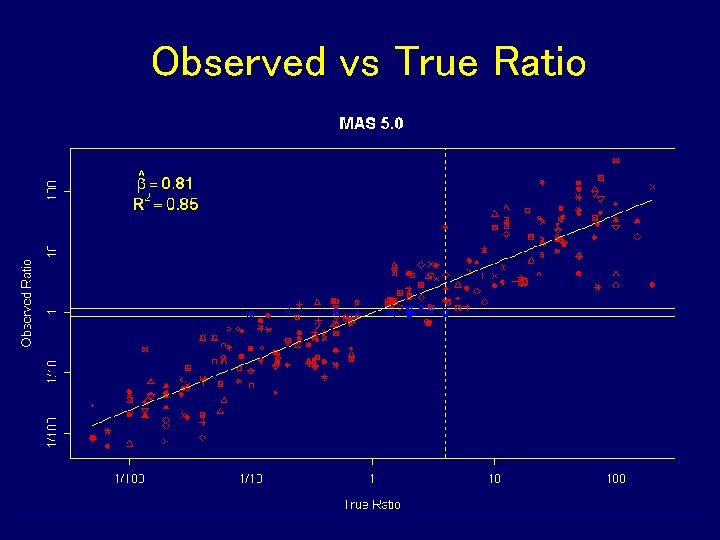
Observed vs True Ratio

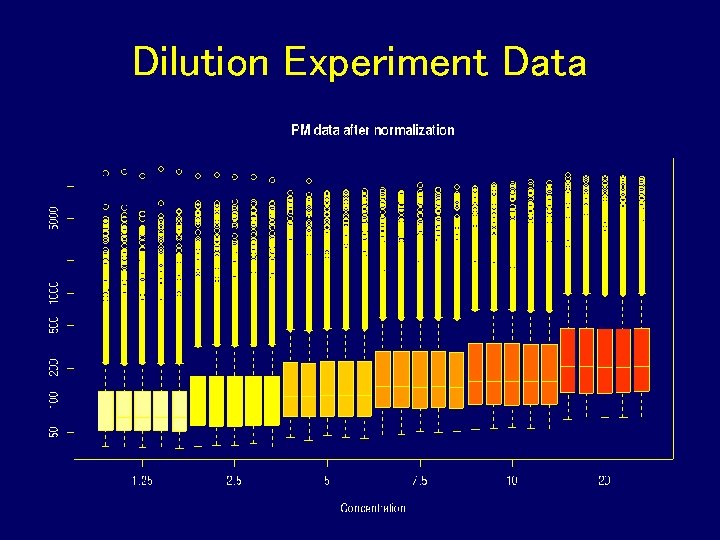
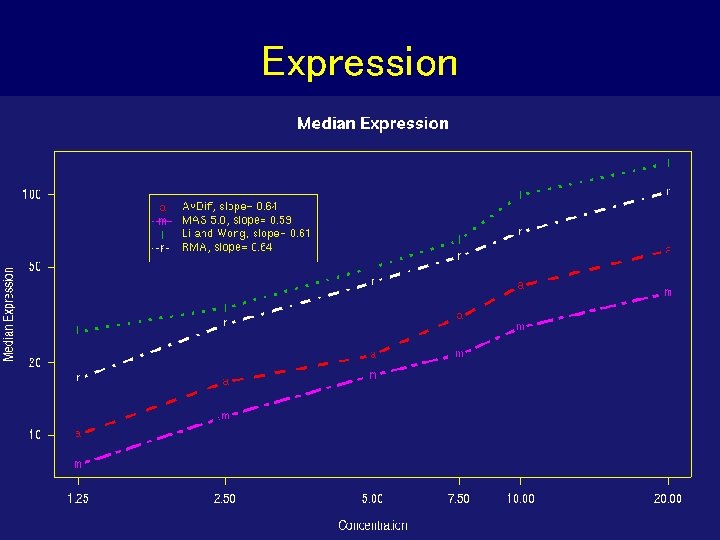
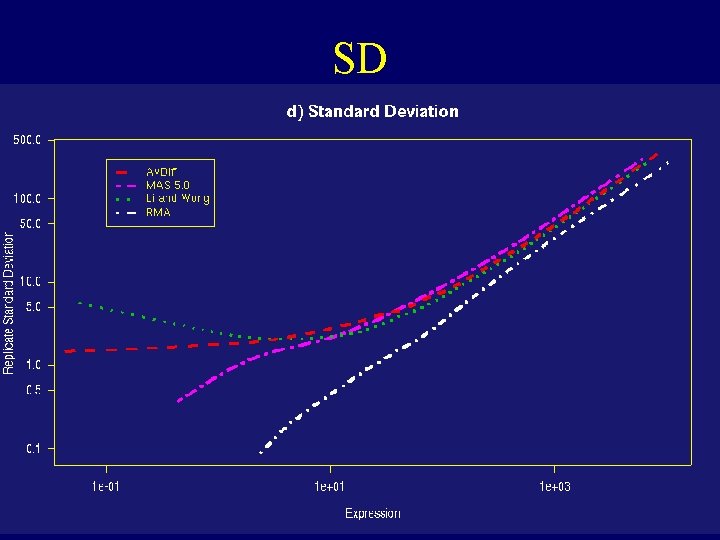
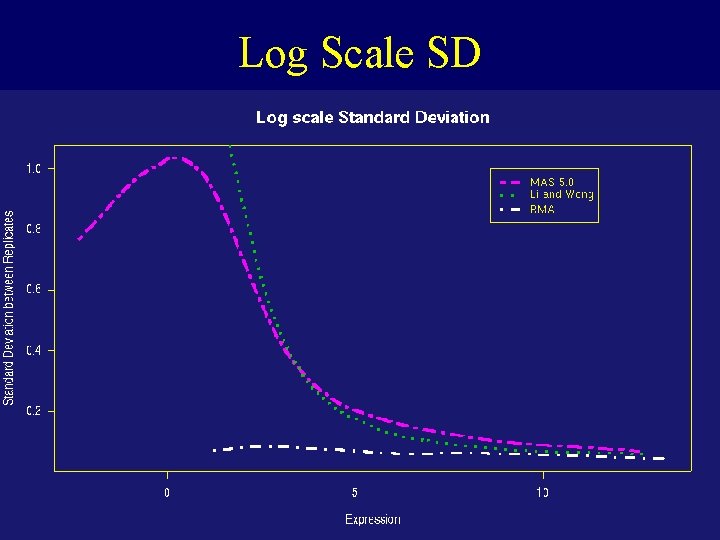
Dilution Experiment • c. RNA hybridized to human chip (HGU 95) in range of proportions and dilutions • Dilution series begins at 1. 25 g c. RNA per Gene. Chip array, and rises through 2. 5, 5. 0, 7. 5, 10. 0, to 20. 0 g per array. 5 replicate chips were used at each dilution • Normalize just within each set of 5 replicates • For each probe set compute expression, average and SD over replicates

Dilution Experiment Data

Expression

SD

Log Scale SD

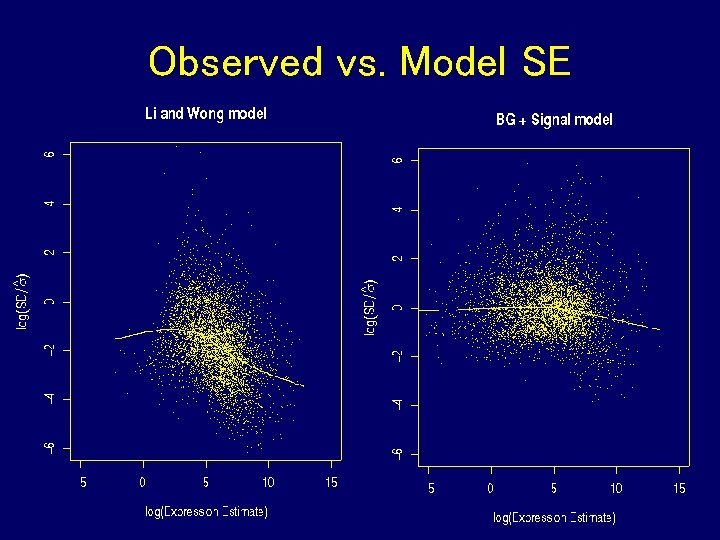
Model check • Compute observed SD of 5 replicate expression estimates • Compute RMS of 5 nominal SDs • Compare by taking the log ratio • Closeness of observed and nominal SD taken as a measure of goodness of fit of the model

Observed vs. Model SE

Conclusion • Take logs • PMs need to be normalized • Using global background improves on use of probe-specific MM • Gene Logic spike-in and dilution study show technology works well • RMA is arguably the best summary in terms of bias, variance and model fit • Future: What stastistic should we use to rank?

Acknowledgements • Gene Brown’s group at Wyeth/Genetics Institute, and Uwe Scherf’s Genomics Research & Development Group at Gene Logic, for generating the spike-in and dilution data • Gene Logic for permission to use these data • Magnus Åstrand (Astra Zeneca Mölndal) • Skip Garcia, Tom Cappola, and Joshua Hare (JHU)