Experiment 4 Molecular cloning DNA cloning of Recombinant

Experiment 4 Molecular cloning (DNA cloning) of Recombinant DNA

What is DNA cloning? The production of large numbers of small genetically identical (copies) of DNA plasmids or viruses bearing one or more genes that were previously present in single copy in a very large chromosome. Clone = Copy

What are requirements for DNA cloning? A typical DNA cloning procedure included the following steps: Ø Preparation of competent bacterial cells. Ø Isolation of two kinds of DNA: bacterial plasmids and foreign DNA (required to copy part of it). Ø Treatment of plasmid & foreign DNA with the same restriction enzyme. Ø Mixture of foreign DNA & plasmids then addition of a ligation enzyme to produce recombinant plasmid DNA.

What is requirements for DNA cloning? Ø Introduction (Transformation) of recombinant plasmid into competent cells. Ø Production of multiple gene copies by gene cloning & selection process for transformed cells. Ø Final screening for transformed cells.
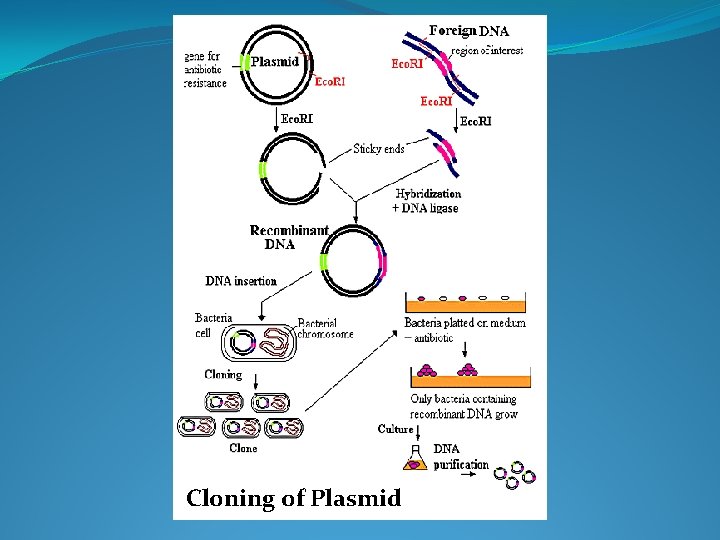
Cloning of Plasmid

Preparation of competent bacterial cells For DNA cloning the following hosts are used : E. coli, Bacillus subtilis, S. typhimurium, viruses, Saccharomices cerevisiae, mice, hamsters & human tissue culture cells.

Why we are using E. coli as a host? 1. Small chromosome (4000 kbp). 2. Fast growth rate (20 min doubling time). 3. Can harbour small cloning vectors (plasmids or viruses). 4. Plasmids can be amplified to large numbers overnight (>1000 copies per cell and 3 x 109 cells per ml).

I. Preparation of competent E. coli using Ca. Cl 2 Ø Transformation: The process of absorption (uptake) of plasmid DNA by competent bacterial cells. Ø Transformation is facilitated by divalent cations (Ca 2+ or Mg 2+). Cells pre-incubated in Ca. Cl 2 solution at 0° C are said to become competent to transformation by DNA. Even so, only 3 - 10% of the cells become truly competent to accept DNA. Competence is also transient, reaching a maximum at about 24 hrs, then falling away rapidly beyond two days for cells refrigerated in Ca. Cl 2 solution. Preservation of competence by freezing at (-70 to -80° C) is an appropriate alternative to repeatedly preparing fresh cells. Ø Optimum results are achieved with competent cells prepared from early or late log phase cells, but not mid-phase cells

I. Protocol of preparation of competent E. coli using Ca. Cl 2 A special strain of E. coli cell is usually used in DNA cloning, called (DH 5α strain): 1. A small overnight culture of DH 5α is used to inoculate one litre of fresh medium [LB broth + 10 m. M Mg. SO 4] at 1: 100. This shaken vigorously (aerated) at 37° C until it reaches early to mid-log phase [An A 600 nm of approx. 0. 4]. 2. Chill the cells on ice for 10 min. 3. Transfer the culture to culture bottles & spin at 5000 rpm for 5 min. 4. Decant the supernatant & resuspend the pellet in 100 ml of ice-cold 50 m. M Ca. Cl 2 solution.

I. Protocol of preparation of competent E. coli using Ca. Cl 2 5. Leave on ice for 15 min. 6. Repeat with another 100 ml of ice-cold Ca. Cl 2 solution. 7. Centrifuge. Resuspend the pellet in 10 ml of ice-cold 15% glycerol / 50 m. M Ca. Cl 2. 8. Dispense 1 ml lots of competent cells into pre-cold 1. 5 Eppendorf tubes & freeze immediately in a -80° C freezer until needed.

II. Isolation of two kinds of DNA: bacterial plasmids and foreign DNA In this experiment; p. UC 18 plasmid vector & bacteriophage lambda DNA will be used and both can be provided commercially.

What is Restriction Enzyme? Restriction enzymes ( also called Restriction endonuclease), occur in bacteria, where they act as protectors against foreign invasive DNA. These enzymes can recognise specific sequences on the DNA molecule & cleave (cut) it. Thus, cleaving of the DNA molecules produces restriction DNA fragments. The sites of action of restriction enzymes are called Restriction or recognition sites & usually short palindromic sequences.
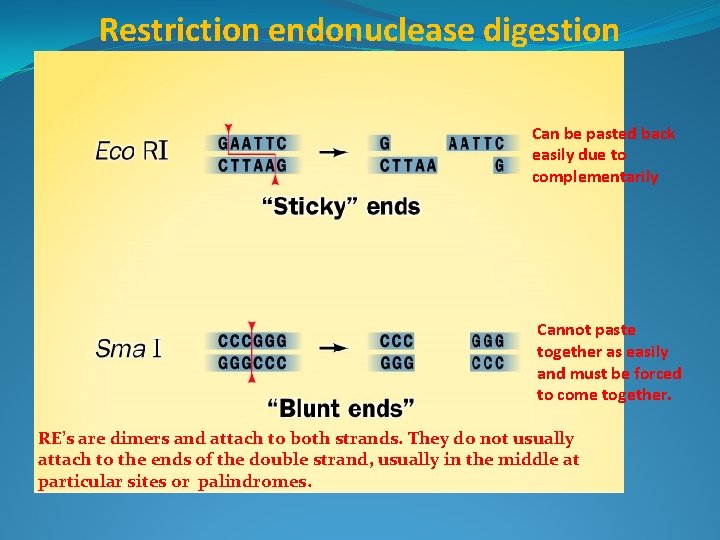
Restriction endonuclease digestion Can be pasted back easily due to complementarily Cannot paste together as easily and must be forced to come together. RE’s are dimers and attach to both strands. They do not usually attach to the ends of the double strand, usually in the middle at particular sites or palindromes.

Restriction endonuclease properities 1. Cut double-stranded DNA palindromes (reads the same, backward and forward sequences). 2. Four-base recognition enzymes (like Hae III) cut on average every 44 = 256 base pairs. 3. Six-base recognition enzymes (like Eco RI) cut on average every 46 = 4096 base pairs. 4. Sites are distributed in DNA on the basis of the frequency of the palindromes and not at random or average (that is, % content of AT or GC pairs). 5. Digestions generate new 5’P and 3’OH ends.

Ligation enzyme The joining of two molecules or pieces of DNA that have cut by the same restriction endonuclease, using the enzyme DNA ligase (usually T 4 DNA ligase) which reforms the phosphodiester bond between adjacent 5`-PO 4 & 3`-OH of the same or different DNA nucleotide bases. The enzyme has required for ATP and Mg 2+. Sticky end OH 5’ - p. A - p. C p. A - p. G - p. T - p. G - p. A - 3’ 3’ - p. T - p. G - p. T - p. C - p. A - p. C - p. T - 5’ OH OH 5’ - p. A - p. C - p. A - p. G p. T - p. G - p. A - 3’ 3’ - p. T - p. G - p. T - p. C p. A - p. C - p. T - 5’ Blunt end OH
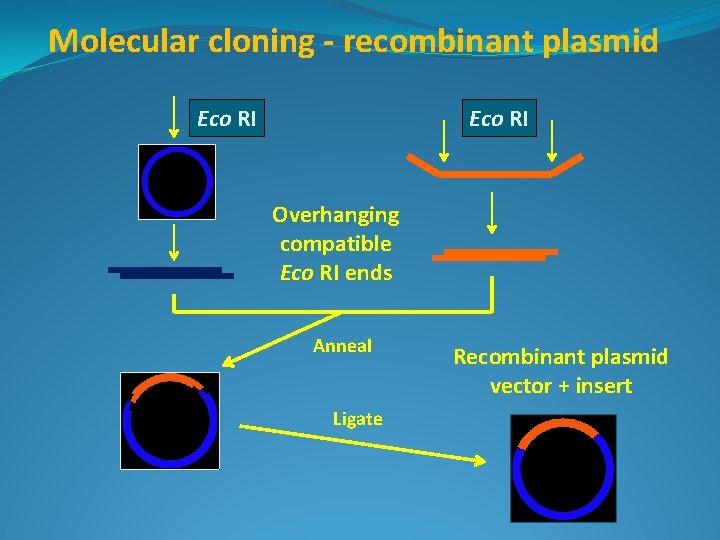
Molecular cloning - recombinant plasmid Eco RI Overhanging compatible Eco RI ends Anneal Ligate Recombinant plasmid vector + insert

Ligation - orientation of ends It’s a 50 -50 chance of joining. DNA ligase is not specific. It just takes one 3’ end attaches it to one 5’ end. OH to P end. A Insert A V B T 4 DNA ligase I B R B Vector Digest vector at site ‘R’ V I A
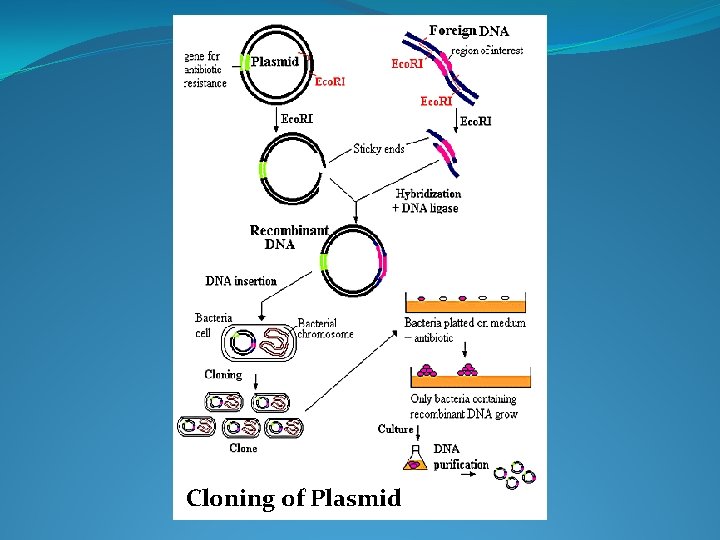
Cloning of Plasmid

V. Transformation of recombinant plasmids DNA into E. coli using Ca. Cl 2 Transformation: The process of absorption (uptake) of plasmid DNA by competent bacterial cells. Transformant: A cell (usually bacteria) that has stably inherited a transformed DNA plasmid. Properties of transformation are: Ø Transformation is only achieved if DNA and competent cells (already treated with Ca. Cl 2 solution) held on ice (or 0° C) for 30 min (Ca 2+ helps to precipitated plasmid on bacterial cell membrane), followed by a 2 min heat shock treatment at 42° C (increase cell membrane permeability). Ø The frequency of transformation is higher by several log for circular DNA than linear DNA. Ø Relaxed circular plasmids are transformed at the same frequency as supercoiled plasmids.

V. Transformation of recombinant plasmids DNA into E. coli using Ca. Cl 2 Ø The frequency of transformation has inverse functionality dependent upon the size of transforming DNA (2 kbp plasmid is transformed at 10 times the efficiency of a 4 kbp plasmid, 100 times more efficiently than 8 kbp, etc. up to 30 kbp). Ø The DNA concentration is critical for frequency of transformation. 10 – 50 ng of plasmid DNA per 100 µl of competent cells gives the best result.

Transformation using Ca. Cl 2 procedure 1. Pipette sample of transforming DNA (1 – 20 ng in 1 - 15 µl) into suspension of competent cells (200 µl) contains & swirl gently to mix. 2. Leave on ice for 30 min. 3. Heat shock cells at 42° C for 2 min then place on ice for 1 min. 4. Add 1 ml of sterile LB broth [+ 20 m. M glucose] to cell suspension & incubate cells at 37° C for 30 min in a shaker water bath with vigorous aeration. 5. Use 1 x LB plate [+ Ampecillin (50 µg/ml) + X-gal (20 µg/plate) + IPTG (0. 1 M) (use 50 µl/plate)]. 6. Spread 300 µl from the culture into the plate. 7. The plates will be incubated right way up at 37° C for 24 hrs. 8. Hold the plates at 4° C.

Lac. Zα/β-galactosidase complementation system (blue / white colony screening) Complementation is a phenomenon in which two separate parts of a normally contiguous protein can associate into a functional composite complex. Ø E. coli Lac. Z gene encodes β-galactosidase (enzyme allows cells to utilise lactose). Ø Many cloning vector plasmids contain a small 5` fragment of Lac. Z gene which encodes the N-terminal portion of lac. Z protein (the α-peptide or M 15 segment containing amino acids 11 to 41 of Lac. Z). Ø The remaining portion of the gene is present in the E. coli host cell chromosome.

Lac. Zα/β-galactosidase complementation system (blue / white colony screening) Ø Transformation containing such vector will supply both portions (a portion within vector + completed portion within bacterial chromosome = complementation) & produced a functional β-galactosidase. Ø IPTG inducer (Isopropyl-β-D-thiogalactopyranoside) allows expression of Lac. Zα-peptide from the vector plasmid. Ø X-Gal (5 -bromo-4 -chloro-3 -indolyl-β-D-galactoside) is colourless modified galactose sugar used as an indicator for hydrolysis of β-galactoside by β-galactosidase enzyme into monosaccharides & release an indole (rich blue colour) which can be easily detected.


Molecular cloning - Lac. Zα - β-Gal complementation Ø In the α-complementation scheme above, the chromosome (Chr) has uncompleted lac. Z gene (ΔM 15 (lac. Z’) that cannot encode the N-terminal α-peptide of β-galactosidase. If a victor (p. UC 18) is present in the same bacterial cell (left panel), an α-peptide can be produced & can combine with the reminder of C-terminal portion of lac. Z to generate a functional β-galactosidase (that can hydrolyse the X-Gal substance to produce a blue colour). Ø If a DNA insert cloned into the vector (right panel), the αpeptide cannot be encoded since its sequence is now interrupted by the insertion. In this case, complementation is impossible & the X-Gal substance cannot be hydrolysed, resulting in white colonies.

Lac. Zα/β-galactosidase complementation system The competent cells are grown in the presence of X-Gal. If the ligation (vector + insert = recombinant plasmid) was successful, the bacterial colony will be white (no X-Gal hydrolyse); if not, the colony will be blue due to hydrolyse XGal by β-galactosidase & release the blue coloured indole. This technique allows for the quick and easy detection of successful ligation of transformant.

Results �The inoculated plate (contains Amp + IPTG + X-Gal) with E. coli will produce one of the following: �If the bacterial cell contains no plasmid (untransformed), the cell will not grow due to Amp selection. �If the cell transformed with a plasmid but without inserted gene, the colonies turn blue. �If the cell transformed with a plasmid + inserted gene, the colonies turn white.

Results The inoculated plate (contains Amp + IPTG + X-Gal) with E. coli will produce one of the following: Ø If the bacterial cell contains no plasmid (untransformed), the cell will not grow due to Amp selection. Ø If the cell transformed with a plasmid but without inserted gene, the colonies turn blue. Ø If the cell transformed with a plasmid + inserted gene, the colonies turn white.
- Slides: 28